Introduction Published in Part in Wikipedia Wikipedia ( Is a Web-Based, Free-Content Encyclopedia and Uses an Openly Editable Model
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Killer-Like Receptors and GPR56 Progressive Expression Defines Cytokine Production of Human CD4+ Memory T Cells
ARTICLE https://doi.org/10.1038/s41467-019-10018-1 OPEN Killer-like receptors and GPR56 progressive expression defines cytokine production of human CD4+ memory T cells Kim-Long Truong1,7, Stephan Schlickeiser1,2,7, Katrin Vogt1, David Boës1, Katarina Stanko1, Christine Appelt1, Mathias Streitz1, Gerald Grütz1,2, Nadja Stobutzki1, Christian Meisel1, Christina Iwert1, Stefan Tomiuk3, Julia K. Polansky2,4, Andreas Pascher5, Nina Babel2,6, Ulrik Stervbo 6, Igor Sauer 5, Undine Gerlach5 & Birgit Sawitzki1,2 1234567890():,; All memory T cells mount an accelerated response on antigen reencounter, but significant functional heterogeneity is present within the respective memory T-cell subsets as defined by CCR7 and CD45RA expression, thereby warranting further stratification. Here we show that several surface markers, including KLRB1, KLRG1, GPR56, and KLRF1, help define low, high, or exhausted cytokine producers within human peripheral and intrahepatic CD4+ memory T-cell populations. Highest simultaneous production of TNF and IFN-γ is observed in KLRB1+KLRG1+GPR56+ CD4 T cells. By contrast, KLRF1 expression is associated with T-cell exhaustion and reduced TNF/IFN-γ production. Lastly, TCRβ repertoire analysis and in vitro differentiation support a regulated, progressive expression for these markers during CD4+ memory T-cell differentiation. Our results thus help refine the classification of human memory T cells to provide insights on inflammatory disease progression and immunotherapy development. 1 Institute of Medical Immunology, Charité – Universitätsmedizin Berlin, Freie Universität Berlin, Humboldt-Universität zu Berlin and Berlin Institute of Health, 13353 Berlin, Germany. 2 Berlin-Brandenburg Center for Regenerative Therapies (BCRT), Charité – Universitätsmedizin Berlin, 13353 Berlin, Germany. 3 Milteny Biotec GmbH, 51429 Bergisch Gladbach, Germany. -

Strategies to Increase ß-Cell Mass Expansion
This electronic thesis or dissertation has been downloaded from the King’s Research Portal at https://kclpure.kcl.ac.uk/portal/ Strategies to increase -cell mass expansion Drynda, Robert Lech Awarding institution: King's College London The copyright of this thesis rests with the author and no quotation from it or information derived from it may be published without proper acknowledgement. END USER LICENCE AGREEMENT Unless another licence is stated on the immediately following page this work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International licence. https://creativecommons.org/licenses/by-nc-nd/4.0/ You are free to copy, distribute and transmit the work Under the following conditions: Attribution: You must attribute the work in the manner specified by the author (but not in any way that suggests that they endorse you or your use of the work). Non Commercial: You may not use this work for commercial purposes. No Derivative Works - You may not alter, transform, or build upon this work. Any of these conditions can be waived if you receive permission from the author. Your fair dealings and other rights are in no way affected by the above. Take down policy If you believe that this document breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 Strategies to increase β-cell mass expansion A thesis submitted by Robert Drynda For the degree of Doctor of Philosophy from King’s College London Diabetes Research Group Division of Diabetes & Nutritional Sciences Faculty of Life Sciences & Medicine King’s College London 2017 Table of contents Table of contents ................................................................................................. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Progastrin Production Transitions from Bmi1+/Prox1+ to Lgr5high Cells During Early Intestinal Tumorigenesis Julie Giraud, M
Progastrin production transitions from Bmi1+/Prox1+ to Lgr5high cells during early intestinal tumorigenesis Julie Giraud, M. Foroutan, Jihane Boubaker-Vitre, Fanny Grillet, Z. Homayed, U. Jadhav, Philippe Crespy, Cyril Breuker, J-F. Bourgaux, J. Hazerbroucq, et al. To cite this version: Julie Giraud, M. Foroutan, Jihane Boubaker-Vitre, Fanny Grillet, Z. Homayed, et al.. Progastrin production transitions from Bmi1+/Prox1+ to Lgr5high cells during early intestinal tumorigene- sis. Translational Oncology, Elsevier, 2020, 14 (2), pp.101001. 10.1016/j.tranon.2020.101001. hal- 03090781 HAL Id: hal-03090781 https://hal.archives-ouvertes.fr/hal-03090781 Submitted on 12 Jan 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution - NonCommercial - NoDerivatives| 4.0 International License Translational Oncology 14 (2021) 101001 Contents lists available at ScienceDirect Translational Oncology journal homepage: www.elsevier.com/locate/tranon Original Research + + high Progastrin production transitions from Bmi1 /Prox1 to Lgr5 cells during early intestinal tumorigenesis J. Giraud a, M. Foroutan b,c,1, J. Boubaker-Vitre a,1, F. Grillet a, Z. Homayed a, U. Jadhav d, P. Crespy a, C. Breuker a, J-F. -

Pancancer Progression Human Vjune2017
Gene Symbol Accession Alias/Prev Symbol Official Full Name AAMP NM_001087.3 - angio-associated, migratory cell protein ABI3BP NM_015429.3 NESHBP|TARSH ABI family, member 3 (NESH) binding protein ACHE NM_000665.3 ACEE|ARACHE|N-ACHE|YT acetylcholinesterase ACTG2 NM_001615.3 ACT|ACTA3|ACTE|ACTL3|ACTSG actin, gamma 2, smooth muscle, enteric ACVR1 NM_001105.2 ACTRI|ACVR1A|ACVRLK2|ALK2|FOP|SKR1|TSRI activin A receptor, type I ACVR1C NM_145259.2 ACVRLK7|ALK7 activin A receptor, type IC ACVRL1 NM_000020.1 ACVRLK1|ALK-1|ALK1|HHT|HHT2|ORW2|SKR3|TSR-I activin A receptor type II-like 1 ADAM15 NM_207195.1 MDC15 ADAM metallopeptidase domain 15 ADAM17 NM_003183.4 ADAM18|CD156B|CSVP|NISBD|TACE ADAM metallopeptidase domain 17 ADAM28 NM_014265.4 ADAM 28|ADAM23|MDC-L|MDC-Lm|MDC-Ls|MDCL|eMDC II|eMDCII ADAM metallopeptidase domain 28 ADAM8 NM_001109.4 CD156|MS2 ADAM metallopeptidase domain 8 ADAM9 NM_001005845.1 CORD9|MCMP|MDC9|Mltng ADAM metallopeptidase domain 9 ADAMTS1 NM_006988.3 C3-C5|METH1 ADAM metallopeptidase with thrombospondin type 1 motif, 1 ADAMTS12 NM_030955.2 PRO4389 ADAM metallopeptidase with thrombospondin type 1 motif, 12 ADAMTS8 NM_007037.4 ADAM-TS8|METH2 ADAM metallopeptidase with thrombospondin type 1 motif, 8 ADAP1 NM_006869.2 CENTA1|GCS1L|p42IP4 ArfGAP with dual PH domains 1 ADD1 NM_001119.4 ADDA adducin 1 (alpha) ADM2 NM_001253845.1 AM2|dJ579N16.4 adrenomedullin 2 ADRA2B NM_000682.4 ADRA2L1|ADRA2RL1|ADRARL1|ALPHA2BAR|alpha-2BAR adrenoceptor alpha 2B AEBP1 NM_001129.3 ACLP AE binding protein 1 AGGF1 NM_018046.3 GPATC7|GPATCH7|HSU84971|HUS84971|VG5Q -

GPR56 Regulates VEGF Production and Angiogenesis During Melanoma Progression
Published OnlineFirst July 1, 2011; DOI: 10.1158/0008-5472.CAN-10-4543 Cancer Tumor and Stem Cell Biology Research GPR56 Regulates VEGF Production and Angiogenesis during Melanoma Progression Liquan Yang1, Guangchun Chen1, Sonali Mohanty1, Glynis Scott2, Fabeha Fazal3, Arshad Rahman3, Shahinoor Begum4, Richard O. Hynes4, and Lei Xu1,2 Abstract Angiogenesis is a critical step during cancer progression. The VEGF is a major stimulator for angiogenesis and is predominantly contributed by cancer cells in tumors. Inhibition of the VEGF signaling pathway has shown promising therapeutic benefits for cancer patients, but adaptive tumor responses are often observed, indicating the need for further understanding of VEGF regulation. We report that a novel G protein–coupled receptor, GPR56, inhibits VEGF production from the melanoma cell lines and impedes melanoma angiogen- esis and growth, through the serine threonine proline-rich segment in its N-terminus and a signaling pathway involving protein kinase Ca. We also present evidence that the two fragments of GPR56, which are generated by autocatalyzed cleavage, played distinct roles in regulating VEGF production and melanoma progression. Finally, consistent with its suppressive roles in melanoma progression, the expression levels of GPR56 are inversely correlated with the malignancy of melanomas in human subjects. We propose that components of the GPR56-mediated signaling pathway may serve as new targets for antiangiogenic treatment of melanoma. Cancer Res; 71(16); 1–11. Ó2011 AACR. Introduction its occurrence strongly argues for combinations of antiangio- genic regimens to effectively treat cancer. Angiogenesis is a process of nascent blood vessel formation VEGF is a potent growth factor for angiogenesis and is a (1) and is critical for tumor growth and metastasis (2). -

The N-Terminus of the Saccharomyces Cerevisiae G Protein-Coupled Receptor Ste2p: Formation of Dimer Interfaces and Negative Regulation
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Doctoral Dissertations Graduate School 8-2013 The N-terminus of the Saccharomyces cerevisiae G protein- coupled receptor Ste2p: formation of dimer interfaces and negative regulation Mohammad Seraj Uddin [email protected] Follow this and additional works at: https://trace.tennessee.edu/utk_graddiss Part of the Biochemistry Commons, Molecular Biology Commons, and the Structural Biology Commons Recommended Citation Uddin, Mohammad Seraj, "The N-terminus of the Saccharomyces cerevisiae G protein-coupled receptor Ste2p: formation of dimer interfaces and negative regulation. " PhD diss., University of Tennessee, 2013. https://trace.tennessee.edu/utk_graddiss/2490 This Dissertation is brought to you for free and open access by the Graduate School at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Doctoral Dissertations by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. To the Graduate Council: I am submitting herewith a dissertation written by Mohammad Seraj Uddin entitled "The N- terminus of the Saccharomyces cerevisiae G protein-coupled receptor Ste2p: formation of dimer interfaces and negative regulation." I have examined the final electronic copy of this dissertation for form and content and recommend that it be accepted in partial fulfillment of the requirements for the degree of Doctor of Philosophy, with a major in Microbiology. Jeffrey -

An Overview on G Protein-Coupled Receptor-Induced Signal Transduction in Acute Myeloid Leukemia
An overview on G protein-coupled receptor-induced signal transduction in Acute Myeloid Leukemia 1* 1,3 4,5,6 2* Frode Selheim , Elise Aasebø , Catalina Ribas and Anna M. Aragay 1The Proteomics Unit at the University of Bergen, Department of Biomedicine, University of Bergen, Jonas Lies vei 91, 5020 Bergen, Norway; 3 Department of Clinical Science, University of Bergen, Jonas Lies vei 87, 5021 Bergen, Norway; [email protected]. 2Departamento de Biologia Celular. Instituto de Biología Molecular de Barcelona (IBMB-CSIC), Spanish National Research Council (CSIC), Baldiri i Reixac, 15, 08028 Barcelona, Spain; [email protected]. 4Departamento de Biología Molecular and Centro de Biología Molecular “Severo Ochoa” (UAM-CSIC), 28049 Madrid, Spain; 5Instituto de Investigación Sanitaria La Princesa, 28006 Madrid, Spain; 6CIBER de Enfermedades Cardiovasculares, ISCIII (CIBERCV), 28029 Madrid, Spain, [email protected] * Corresponding authors: Frode Selheim Adr: Jonas Lies vei 91, 5020 Bergen, Norway Email: [email protected], Tel:+4755586091 Anna M. Aragay Adr: Baldiri i Reixac, 15, 08028 Barcelona. Spain. E-mail: [email protected]; Tel.: +934098671 1 Abstract Background: Acute myeloid leukemia (AML) is a genetically heterogeneous disease characterized by uncontrolled proliferation of precursor myeloid-lineage cells in the bone marrow. AML is also characterized with patients with poor long-term survival outcomes due to relapse. Many efforts have been made to understand the biological heterogeneity of AML and the challenges to develop new therapies are therefore enormous. G protein-coupled receptors (GPCRs) are a large attractive drug targeted family of transmembrane proteins, and aberrant GPCR expression and GPCR-mediated signaling have been implicated in leukemogenesis of AML. -

The Adhesion G Protein-Coupled Receptor GPR56 Is a Cell-Autonomous Regulator of Oligodendrocyte Development
ARTICLE Received 27 May 2014 | Accepted 14 Dec 2014 | Published 21 Jan 2015 DOI: 10.1038/ncomms7121 OPEN The adhesion G protein-coupled receptor GPR56 is a cell-autonomous regulator of oligodendrocyte development Stefanie Giera1,*, Yiyu Deng1,*,w, Rong Luo1,*, Sarah D. Ackerman2, Amit Mogha2, Kelly R. Monk2,3, Yanqin Ying1, Sung-Jin Jeong1,w, Manabu Makinodan4,5, Allison R. Bialas4,5, Bernard S. Chang6, Beth Stevens4,5, Gabriel Corfas4,5,w & Xianhua Piao1 Mutations in GPR56, a member of the adhesion G protein-coupled receptor family, cause a human brain malformation called bilateral frontoparietal polymicrogyria (BFPP). Magnetic resonance imaging (MRI) of BFPP brains reveals myelination defects in addition to brain malformation. However, the cellular role of GPR56 in oligodendrocyte development remains unknown. Here, we demonstrate that loss of Gpr56 leads to hypomyelination of the central nervous system in mice. GPR56 levels are abundant throughout early stages of oligodendrocyte development, but are downregulated in myelinating oligodendrocytes. Gpr56-knockout mice manifest with decreased oligodendrocyte precursor cell (OPC) proliferation and diminished levels of active RhoA, leading to fewer mature oligodendrocytes and a reduced number of myelinated axons in the corpus callosum and optic nerves. Conditional ablation of Gpr56 in OPCs leads to a reduced number of mature oligodendrocytes as seen in constitutive knockout of Gpr56. Together, our data define GPR56 as a cell-autonomous regulator of oligodendrocyte development. 1 Division of Newborn Medicine, Department of Medicine, Boston Children’s Hospital and Harvard Medical School, Boston, Massachusetts 02115, USA. 2 Department of Developmental Biology, Washington University School of Medicine, St Louis, Missouri 63110, USA. -

Multi-Functionality of Proteins Involved in GPCR and G Protein Signaling: Making Sense of Structure–Function Continuum with In
Cellular and Molecular Life Sciences (2019) 76:4461–4492 https://doi.org/10.1007/s00018-019-03276-1 Cellular andMolecular Life Sciences REVIEW Multi‑functionality of proteins involved in GPCR and G protein signaling: making sense of structure–function continuum with intrinsic disorder‑based proteoforms Alexander V. Fonin1 · April L. Darling2 · Irina M. Kuznetsova1 · Konstantin K. Turoverov1,3 · Vladimir N. Uversky2,4 Received: 5 August 2019 / Revised: 5 August 2019 / Accepted: 12 August 2019 / Published online: 19 August 2019 © Springer Nature Switzerland AG 2019 Abstract GPCR–G protein signaling system recognizes a multitude of extracellular ligands and triggers a variety of intracellular signal- ing cascades in response. In humans, this system includes more than 800 various GPCRs and a large set of heterotrimeric G proteins. Complexity of this system goes far beyond a multitude of pair-wise ligand–GPCR and GPCR–G protein interactions. In fact, one GPCR can recognize more than one extracellular signal and interact with more than one G protein. Furthermore, one ligand can activate more than one GPCR, and multiple GPCRs can couple to the same G protein. This defnes an intricate multifunctionality of this important signaling system. Here, we show that the multifunctionality of GPCR–G protein system represents an illustrative example of the protein structure–function continuum, where structures of the involved proteins represent a complex mosaic of diferently folded regions (foldons, non-foldons, unfoldons, semi-foldons, and inducible foldons). The functionality of resulting highly dynamic conformational ensembles is fne-tuned by various post-translational modifcations and alternative splicing, and such ensembles can undergo dramatic changes at interaction with their specifc partners. -
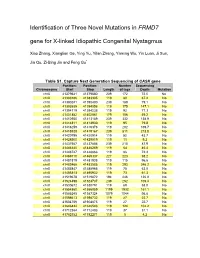
Identification of Three Novel Mutations in FRMD7 Gene for X-Linked Idiopathic Congenital Nystagmus
Identification of Three Novel Mutations in FRMD7 gene for X-linked Idiopathic Congenital Nystagmus Xiao Zhang, Xianglian Ge, Ying Yu, Yilan Zhang, Yaming Wu, Yin Luan, Ji Sun, Jia Qu, Zi-Bing Jin and Feng Gu* Table S1. Capture Next Generation Sequencing of CASK gene Position: Position: Number Sequencing Chromosome Start Stop Length of tags Depth Mutation chrX 41379641 41379880 239 172 72.0 No chrX 41383186 41383305 119 80 67.2 No chrX 41390241 41390480 239 189 79.1 No chrX 41393939 41394058 119 175 147.1 No chrX 41394119 41394238 119 92 77.3 No chrX 41401882 41402061 179 106 59.2 No chrX 41412950 41413189 239 332 138.9 No chrX 41414811 41414930 119 95 79.8 No chrX 41416259 41416378 119 202 169.7 No chrX 41418928 41419167 239 511 213.8 No chrX 41420795 41420914 119 52 43.7 No chrX 41428900 41429019 119 11 9.2 No chrX 41437567 41437806 239 210 87.9 No chrX 41446140 41446259 119 54 45.4 No chrX 41448747 41448866 119 86 72.3 No chrX 41469110 41469337 227 223 98.2 No chrX 41481819 41481938 119 115 96.6 No chrX 41483466 41483585 119 293 246.2 No chrX 41485847 41485966 119 75 63.0 No chrX 41495813 41495932 119 73 61.3 No chrX 41519678 41519872 194 246 126.8 No chrX 41524498 41524737 239 252 105.4 No chrX 41530672 41530791 119 69 58.0 No chrX 41554860 41556059 1199 1932 161.1 No chrX 41586245 41587324 1079 1044 96.8 No chrX 41598613 41598732 119 27 22.7 No chrX 41604756 41604875 119 27 22.7 No chrX 41646424 41646543 119 124 104.2 No chrX 41712364 41712483 119 37 31.1 No chrX 41782152 41782271 119 5 4.2 No Table S2.