Genotypes for Aldehyde Dehydrogenase Deficiency and Alcohol Sensitivity. the Inactive ALDH2(2) Allele Is Dominant
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Aldehyde Dehydrogenase ALDH2*2 Allele Exhibits Dominance Over ALDH2*1 in Transduced Hela Cells
The aldehyde dehydrogenase ALDH2*2 allele exhibits dominance over ALDH2*1 in transduced HeLa cells. Q Xiao, … , T Johnston, D W Crabb J Clin Invest. 1995;96(5):2180-2186. https://doi.org/10.1172/JCI118272. Research Article Individuals heterozygous or homozygous for the variant aldehyde dehydrogenase (ALDH2) allele (ALDH2*2), which encodes a protein differing only at residue 487 from the normal protein, have decreased ALDH2 activity in liver extracts and experience cutaneous flushing when they drink alcohol. The mechanisms by which this allele exerts its dominant effect is unknown. To study this effect, the human ALDH2*1 cDNA was cloned and the ALDH2*2 allele was generated by site-directed mutagenesis. These cDNAs were transduced using retroviral vectors into HeLa and CV1 cells, which do not express ALDH2. The normal allele directed synthesis of immunoreactive ALDH2 protein (ALDH2E) with the expected isoelectric point. Extracts of these cells contained increased aldehyde dehydrogenase activity with low Km for the aldehyde substrate. The ALDH2*2 allele directed synthesis of mRNA and immunoreactive protein (ALDH2K), but the protein lacked enzymatic activity. When ALDH2*1-expressing cells were transduced with ALDH2*2 vectors, both mRNAs were expressed and immunoreactive proteins with isoelectric points ranging between those of ALDH2E and ALDH2K were present, indicating that the subunits formed heteromers. ALDH2 activity in these cells was reduced below that of the parental ALDH2*1-expressing cells. Thus, the ALDH2*2 allele is sufficient to cause ALDH2 deficiency in vitro. Find the latest version: https://jci.me/118272/pdf The Aldehyde Dehydrogenase ALDH2*2 Allele Exhibits Dominance over ALDH2*1 in Transduced HeLa Cells Qing Xiao, * Henry Weiner,* Timothy Johnston,* and David W. -

Expression and Characterization of Pantoea CO Dehydrogenase To
www.nature.com/scientificreports OPEN Expression and characterization of Pantoea CO dehydrogenase to utilize CO-containing industrial Received: 31 October 2016 Accepted: 06 February 2017 waste gas for expanding the Published: 14 March 2017 versatility of CO dehydrogenase Eun Sil Choi1,2,*, Kyoungseon Min3,*, Geun-Joong Kim2, Inchan Kwon1 & Yong Hwan Kim3 Although aerobic CO dehydrogenases (CODHs) might be applicable in various fields, their practical applications have been hampered by low activity and no heterologous expression. We, for the first time, could functionally express recombinant PsCODH in E. coli and obtained a highly concentrated recombinant enzyme using an easy and convenient method. Its electron acceptor spectra, optimum −1 conditions (pH 6.5 and 30 °C), and kinetic parameters (kcat of 12.97 s , Km of 0.065 mM, and specific activity of 0.86 Umg−1) were examined. Blast furnace gas (BFG) containing 20% CO, which is a waste gas from the steel-making process, was tested as a substrate for PsCODH. Even with BFG, the recombinant PsCODH retained 88.2% and 108.4% activity compared with those of pure CO and 20% CO, respectively. The results provide not only a promising strategy to utilize CO-containing industrial waste gases as cheap, abundant, and renewable resources but also significant information for further studies about cascade reactions producing value-added chemicals via CO2 as an intermediate produced by a CODH- based CO-utilization system, which would ultimately expand the versatility of CODH. Carbon monoxide (CO), which is a pollutant in the atmosphere, is massively emitted through both natural (e.g., production by plants and volcanic activity) and artificial processes (e.g., incomplete combustion of fuels and industrial processes)1–3. -

The Evolution and Population Genetics of the ALDH2 Locus: Random Genetic Drift, Selection, and Low Levels of Recombination
doi: 10.1046/j.1529-8817.2003.00060.x The evolution and population genetics of the ALDH2 locus: random genetic drift, selection, and low levels of recombination Hiroki Oota1, Andrew J. Pakstis1, Batsheva Bonne-Tamir2, David Goldman3, Elena Grigorenko4, Sylvester L. B. Kajuna5, Nganyirwa J. Karoma5, Selemani Kungulilo6, Ru-Band Lu7, Kunle Odunsi8, Friday Okonofua9, Olga V. Zhukova10, Judith R. Kidd1 and Kenneth K. Kidd1,∗ 1Department of Genetics, Yale University School of Medicine, 333 Cedar Street, P.O. Box 208005, New Haven, CT 06520-8005, USA 2Department of Human Genetics, Sackler School of Medicine, Tel Aviv University, Tel Aviv, Israel 3Laboratory of Neurogenetics, National Institute of Alcohol Abuse and Alcoholism, Rockville, MD 20852, USA 4Department of Psychology, Yale University, New Haven, CT 06520, USA 5The Hubert Kairuki Memorial University, Dar es Salaam, Tanzania 6Muhimbili University College of Health Sciences, Dar es Salaam, Tanzania 7Department of Psychiatry, Tri-Service General hospital, National Defense Medical Center, Taipei, Taiwan, R.O.C. 8Department of Gynecological Oncology, Roswell Park Cancer Institute, Buffalo, NY 14263, USA 9Department of Obstetrics and Gynecology, Faculty of Medicine, University of Benin, Benin City, Nigeria 10N.I. Vavilov Institute of General Genetics RAS, Moscow, Russia Summary The catalytic deficiency of human aldehyde dehydrogenase 2 (ALDH2) is caused by a nucleotide substitution (G1510A; Glu487Lys) in exon 12 of the ALDH2 locus. This SNP,and four non-coding SNPs, including one in the promoter, span 40 kb of ALDH2; these and one downstream STRP have been tested in 37 worldwide populations. Only four major SNP-defined haplotypes account for almost all chromosomes in all populations. -

How Is Alcohol Metabolized by the Body?
Overview: How Is Alcohol Metabolized by the Body? Samir Zakhari, Ph.D. Alcohol is eliminated from the body by various metabolic mechanisms. The primary enzymes involved are aldehyde dehydrogenase (ALDH), alcohol dehydrogenase (ADH), cytochrome P450 (CYP2E1), and catalase. Variations in the genes for these enzymes have been found to influence alcohol consumption, alcohol-related tissue damage, and alcohol dependence. The consequences of alcohol metabolism include oxygen deficits (i.e., hypoxia) in the liver; interaction between alcohol metabolism byproducts and other cell components, resulting in the formation of harmful compounds (i.e., adducts); formation of highly reactive oxygen-containing molecules (i.e., reactive oxygen species [ROS]) that can damage other cell components; changes in the ratio of NADH to NAD+ (i.e., the cell’s redox state); tissue damage; fetal damage; impairment of other metabolic processes; cancer; and medication interactions. Several issues related to alcohol metabolism require further research. KEY WORDS: Ethanol-to acetaldehyde metabolism; alcohol dehydrogenase (ADH); aldehyde dehydrogenase (ALDH); acetaldehyde; acetate; cytochrome P450 2E1 (CYP2E1); catalase; reactive oxygen species (ROS); blood alcohol concentration (BAC); liver; stomach; brain; fetal alcohol effects; genetics and heredity; ethnic group; hypoxia The alcohol elimination rate varies state of liver cells. Chronic alcohol con- he effects of alcohol (i.e., ethanol) widely (i.e., three-fold) among individ- sumption and alcohol metabolism are on various tissues depend on its uals and is influenced by factors such as strongly linked to several pathological concentration in the blood T chronic alcohol consumption, diet, age, consequences and tissue damage. (blood alcohol concentration [BAC]) smoking, and time of day (Bennion and Understanding the balance of alcohol’s over time. -

Sudan University of Science and Technology College of Graduate
Sudan University of Science and Technology College of Graduate Studies Detection of CDX2 Tumor Marker and Human Papilloma Virus in Esophageal Tumors اﻟﻜﺸﻒ ﻋﻦ اﻟﻮاﺳﻤﺔ اﻟﻮرﻣﯿﺔ CDX2 و ﻓﯿﺮوس اﻟﻮرم اﻟﺤﻠﯿﻤﻲ اﻟﺒﺸﺮي ﻓﻰ اورام اﻟﻤﺮيء A dissertation submitted for partial fulfillment for the requirement for M.Sc degree in Medical laboratory Science (Histopathology and cytology) Submitted by: Eslam Mohamed khoghali Ahmed B.SC (honors) in chemical pathology and histopathology – National Ribat University (2010) Supervised by: Dr. Mohammed Siddig Abdelaziz 2014 ﻗﺎل ﷲ ﺗﻌﺎﻟﻰ ۖ◌ ٰ◌ ﺻدق ﷲ اﻟﻌظﯾم ﺳورة اﻟﺗوﺑﺔ اﻵﯾﺔ 105 Dedication To my father…. My mother…. My brothers…. My friends… Eslam, 2014 Acknowledgement All great thanks are firstly to Allah. I would like to send thanks to my supervisor Dr. Mohamed Siddige Abdalaziz for his guidance and continuous assistance. Thanks are also extend to staff of histopathology and cytology in Ibn-Seina hospital and to members of histopathology and cytology department in Sudan University for science and technology for providing me materials and equipment, finally thanks to every one helped me. Eslam, 2014 Abstract This is a descriptive retrospective study conducted at Ibn Seina hospital and Sudan University of Science and Technology during the period from April 2014 to October 2014. The study aimed at detecting CDX2 and association of Human Papilloma Virus (HPV) in esophageal tumor among Sudanese patients using immune-histochemistry and polymerase chain reaction. Samples from 30 patients previously diagnosed with esophageal tumor (20 with esophageal cancer and 10 with benign). Their ages ranging between 8 to 82 years with mean age 37. From each block two slides were cut one for CDX2 and other for HPV detection, also 10µ was cut from each block in eppendorf tube for detection of HPV type 18 gene using PCR.SPSS version 16 computer program was used to analyze the data. -

Bioalcohol Production by a New Synthetic Route in a Hyperthermophilic Archaeon
COMMENTARY COMMENTARY Bioalcohol production by a new synthetic route in a hyperthermophilic archaeon Volker Müller1 Department of Molecular Microbiology & Bioenergetics, Institute of Molecular Biosciences, Johann Wolfgang Goethe University Frankfurt am Main, 60438 Frankfurt, Germany The still growing demand for bioethanol is establishing an electrochemical ion gradient currently met by two microbiological pro- across the cytoplasmic membrane that then cesses. The by far most important process is drives ATP synthesis by a membrane-bound the use of first- (sugar) or second- (lignocel- ATP synthase (5). One widespread, ferre- lulosic biomass) generation raw materials and doxin-fueled ion-translocating enyzme is + yeast as catalysts. Yeasts convert the sugars the ferredoxin:NAD oxidoreductase found via glycolysis to pyruvate, which is decar- in many bacteria and few archaea (6). In P. furiosus boxylated, and the resulting acetaldehyde is , the reduced ferredoxin is oxidized Fig. 1. Pathways for ethanol formation from glucose in then reduced to ethanol by a monofunctional by a different enzyme, a membrane-bound, yeasts (A) and bacteria (B). Adh1, alcoholdehydrogenase ethanol dehydrogenase. The second and in- multisubunit hydrogenase (Mbh) with evolu- 1; AdhE, bifunctional CoA-dependent ethanol/aldehyde dustrially less important is the bacterial pro- tionarysimilaritytothecomplexIofbacterial dehydrogenase. cess, in which pyruvate is oxidized to acetyl- and mitochondrial electron transport chains CoA, which is then reduced by a bifunctional (7). The Mbh is a proton-reducing (hydrogen aldehyde dehydrogenase/ethanol dehydroge- evolving) and sodium ion-extruding mem- used to drive acetate reduction and not the nase (AdhE) to ethanol. This pathway is brane protein complex (8, 9). generation of a chemiosmotic ion gradient, found in many fermenting bacteria (Fig. -

Health-Related Effects of Genetic Variations Of
FOCUS ON SPECIAL POPULATIONS HEALTH-RELATED EFFECTS OF GENETIC effects deter people from alcohol consumption and therefore VARIATIONS OF ALCOHOL-METABOLIZING have a protective effect against alcoholism. ENZYMES IN AFRICAN AMERICANS Because alcohol metabolism can significantly influence drinking behavior and the risk for alcoholism (Yin and Agarwal 2001), it is important to understand how genetic Denise M. Scott, Ph.D., and Robert E. Taylor, factors influence metabolism. The different ADH and M.D., Ph.D. ALDH isoforms are encoded by different genes, each of which again may be present in different variations (i.e., Alcohol metabolism involves two key enzymes—alcohol alleles). The occurrence of alternate alleles in a population dehydrogenase (ADH) and aldehyde dehydrogenase is termed polymorphism. The activity and distribution of (ALDH). There are several types of ADH and ALDH, each of the ADH and ALDH alleles and of the proteins they encode which may exist in several variants (i.e., isoforms) that have been intensively studied. For example, researchers differ in their ability to break down alcohol and its toxic have identified differences across ethnic groups in the fre metabolite acetaldehyde. The isoforms are encoded by quencies with which individual genes or gene variants different gene variants (i.e., alleles) whose distribution occur (Osier et al. 2002; Chou et al. 1999). This brief among ethnic groups differs. One variant of ADH is article examines the prevalence and effects of genetic vari ADH1B, which is encoded by several alleles. An allele ants of ADH and ALDH genes in African Americans. called ADH1B*3 is unique to people of African descent and certain Native American tribes. -

Biochemical and Physiological Research on the Disposition and Fate of Ethanol in the Body
Garriott's Medicolegal Aspects of Alcohol, 5th edition, Edited by James Garriott PhD Lawyers & Judges Publishing Co., Tuscon, AZ, 2008 Chapter 3 Biochemical and Physiological Research on the Disposition and Fate of Ethanol in the Body A.W. Jones, Ph.D., D.Sc. Synopsis . Repetitive F Drinking 3.1 Introduction G. Effect of Age on Widmark Parameters 3.2 Fate of Drugs in the Body H. Blood-Alcohol Profiles after Drinking Beer 3.3 Forensic Science Aspects of Alcohol I. Retrograde Extrapolation 3.4 Ethyl Alcohol J. Massive Ingestion of Alcohol under Real-World Conditions A. Chemistry K. Effects of Drugs on Metabolism of Ethanol . B Amounts of Alcohol Consumed L. Elimination Rates Ethanol in Alcoholics During Detoxification C. Alcoholic Beverages M. Ethanol Metabolism in Pathological States . D Analysis of Ethanol in Body Fluids N. Short-Term Fluctuations in Blood-Alcohol Profiles E. Reporting Blood Alcohol Concentration . Intravenous O vs. Oral Route of Ethanol Administration . F Water Content of Biofluids 3.8 Ethanol in Body Fluids and Tissues 3.5 Alcohol in the Body A. Water Content of Specimens A. Endogenous Ethanol . Urine B . B Absorption 1. Diuresis 1. Uptake from the gut 2. Urine-blood ratios 2. Importance of gastric emptying 3. Concentration-time profiles 3. Inhalation of ethanol vapors C. Breath 4. Absorption through skin 1. Breath alcohol physiology 5. Concentration of ethanol in the beverage consumed 2. Blood-breath ratios C. Distribution 3. Concentration-time profiles 1. Arterial-venous differences . Saliva D 2. Plasma/blood and serum/blood ratios 1. Saliva production 3. Volume of distribution 2. -
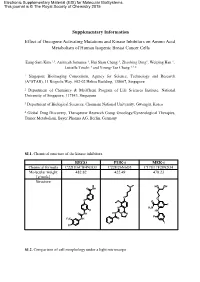
Supplementary Information Effect of Oncogene Activating Mutations And
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is © The Royal Society of Chemistry 2015 Supplementary Information Effect of Oncogene Activating Mutations and Kinase Inhibitors on Amino Acid Metabolism of Human Isogenic Breast Cancer Cells Eung-Sam Kim 1,3, Animesh Samanta 1, Hui Shan Cheng 1, Zhaobing Ding1, Weiping Han 1, Luisella Toschi 4 and Young-Tae Chang 1,2,* 1 Singapore Bioimaging Consortium, Agency for Science, Technology and Research (A*STAR), 11 Biopolis Way, #02-02 Helios Building, 138667, Singapore 2 Department of Chemistry & MedChem Program of Life Sciences Institute, National University of Singapore, 117543, Singapore. 3 Department of Biological Sciences, Chonnam National University, Gwangju, Korea 4 Global Drug Discovery, Therapeutic Research Group Oncology/Gynecological Therapies, Tumor Metabolism, Bayer Pharma AG, Berlin, Germany SI.1. Chemical structure of the kinase inhibitors. REGO PI3K-i MEK-i Chemical formula C22H16ClF4N3O3 C22H26N6O3 C17H17F2IN2O4 Molecular weight 482.82 422.49 478.23 [g/mole] Structure N HO OH O NH O O O O O F N H2N F HN O NH HN N N F F3C NH O N I Cl SI.2. Comparison of cell morphology under a light microscope Scale bar: 100 µm 10x objective lens at Ti microscope (Nikon) SI.3. Scheme of the experimental method for metabolite extraction SI.4. Gradient profile of HPLC run. A : Buffer solution B: Mobile phase H O Time 2 (40 mM NaHPO4, (MeOH/ACN/H2O, Flow rate pH7.8 at RT) 45:45:10, v/v/v) (ml/min) (min) (%) (%) (%) 0 100 0 0 1.0 3 90 10 0 1.0 15 87 13 0 1.0 25 70 30 0 1.0 37 65 35 0 1.0 43 58 42 0 1.0 52 43 57 0 1.0 55 0 100 0 1.0 60 0 0 100 1.0 65 to 80 100 0 0 1.0 SI.5. -

Associations of Alcohol Consumption and Alcohol Flush Reaction with Leukocyte Telomere Length in Korean Adults
Nutrition Research and Practice 2017;11(4):334-339 ⓒ2017 The Korean Nutrition Society and the Korean Society of Community Nutrition http://e-nrp.org Associations of alcohol consumption and alcohol flush reaction with leukocyte telomere length in Korean adults Hyewon Wang, Hyungjo Kim and Inkyung Baik§ Department of Foods and Nutrition, College of Science and Technology, Kookmin University, 77, Jeongnung-ro, Seongbuk-gu, Seoul 02707, Korea BACKGROUND/OBJECTIVES: Telomere length is a useful biomarker for determining general aging status. Some studies have reported an association between alcohol consumption and telomere length in a general population; however, it is unclear whether the alcohol flush reaction, which is an alcohol-related trait predominantly due to acetaldehyde dehydrogenase deficiency, is associated with telomere length. This cross-sectional study aimed to evaluate the associations of alcohol consumption and alcohol flush reaction with leukocyte telomere length (LTL). SUBJECTS/METHODS: The study included 1,803 Korean adults. Participants provided blood specimens for LTL measurement assay and reported their alcohol drinking status and the presence of an alcohol flush reaction via a questionnaire-based interview. Relative LTL was determined by using a quantitative polymerase chain reaction. Statistical analysis used multiple linear regression models stratified by sex and age groups, and potential confounding factors were considered. RESULTS: Age-specific analyses showed that heavy alcohol consumption (> 30 g/day) was strongly associated with a reduced LTL in participants aged ≥ 65 years (P < 0.001) but not in younger participants. Similarly, the alcohol flush reaction was associated with a reduced LTL only in older participants who consumed > 15 g/day of alcohol (P < 0.01). -

A Novel Bifunctional Aldehyde/Alcohol Dehydrogenase Mediating Ethanol Formation from Acetyl-Coa in Hyperthermophiles
A novel bifunctional aldehyde/alcohol dehydrogenase mediating ethanol formation from acetyl-CoA in hyperthermophiles Qiang Wang Jiangsu University Chong Sha Jiangsu University Hongcheng Wang Jiangsu University Kesen Ma University of Waterloo Juergen Wiegel University of Georgia Abd El-Fatah Abomohra Tanta University Weilan Shao ( [email protected] ) Jiangsu University Research Keywords: Acetyl-CoA reduction, Bifunctional aldehyde/alcohol dehydrogenase, Ethanol fermentation, Hyperthermophiles, Thermotoga neapolitana Posted Date: March 15th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-17223/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/15 Abstract Background: Hyperthermophilic fermentation at temperatures above 80 °C allows in situ product removal to mitigate the ethanol toxicity, and reduces microbial contamination without autoclaving/cooling of feedstock. Many species of Thermotoga grow at temperatures up to 90 °C, and have enzymes to degrade and utilize lignocelluloses, which provide advantages for achieving consolidated processes of cellulosic ethanol production. However, no CoA-dependent aldehyde dehydrogenase (CoA-Aldh) from any hyperthermophiles has been documented in literature so far. The pyruvate ferredoxin oxidoreductases from hyperthermophiles have pyruvate decarboxylase activity, which convert about 2% and 98% of pyruvate to acetaldehyde and acetyl-CoA (ac-CoA), respectively. Acetyl-CoA can be converted to acetic acid, if there is no CoA-Aldh -

Effect of the Allelic Variants of Aldehyde
REVIEW Effect of the allelic variants of aldehyde dehydrogenase ALDH2*2 and alcohol dehydrogenase ADH1B*2 on blood acetaldehyde concentrations Giia-Sheun Peng1 and Shih-Jiun Yin2* 1Department of Neurology, Tri-Service General Hospital, 325 Chenggong Road Section 2, Taipei 114, Taiwan 2Department of Biochemistry, National Defense Medical Center, 161 Minchuan East Road Section 6, Taipei 114, Taiwan *Correspondence to: Tel: þ886 2 8792 3100 ext 18800; Fax: þ886 2 8792 4818; E-mail: [email protected] Date received (in revised form): 30th July 2008 Abstract Alcoholism is a complex behavioural disorder. Molecular genetics studies have identified numerous candidate genes associated with alcoholism. It is crucial to verify the disease susceptibility genes by correlating the pinpointed allelic variations to the causal phenotypes. Alcohol dehydrogenase (ADH) and aldehyde dehydrogenase (ALDH) are the principal enzymes responsible for ethanol metabolism in humans. Both ADH and ALDH exhibit functional polymorphisms among racial populations; these polymorphisms have been shown to be the important genetic determinants in ethanol metabolism and alcoholism. Here, we briefly review recent advances in genomic studies of human ADH/ALDH families and alcoholism, with an emphasis on the pharmacogenetic consequences of venous blood acetaldehyde in the different ALDH2 genotypes following the intake of various doses of ethanol. This paper illustrates a paradigmatic example of phenotypic verifications in a protective disease gene for substance abuse. Keywords: alcohol dehydrogenase, aldehyde dehydrogenase, single nucleotide polymorphism, alcoholism, ethanol metab- olism, blood acetaldehyde Introduction Alcohol dehydrogenase (ADH) and aldehyde dehy- exposure and the concentrations of ethanol and its drogenase (ALDH) are the principal enzymes respon- metabolite acetaldehyde attained in body fluids and sible for hepatic metabolism of ethanol.1 Human tissue within that period.