Escherichia Coli Β-Clamp Slows Down DNA Polymerase I Dependent Nick Translation While Accelerating Ligation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Copyright by Young-Sam Lee 2010
Copyright by Young-Sam Lee 2010 The Dissertation Committee for Young-Sam Lee Certifies that this is the approved version of the following dissertation: Structural and Functional Studies of the Human Mitochondrial DNA Polymerase Committee: Whitney Yin, Supervisor Ian Molineux Kenneth Johnson Tanya Paull Jon Robertus Structural and Functional Studies of the Human Mitochondrial DNA Polymerase by Young-Sam Lee, B.S, M.S. Dissertation Presented to the Faculty of the Graduate School of The University of Texas at Austin in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy The University of Texas at Austin August, 2010 Dedication For my wife, In-Sook Jung. Acknowledgements I would like to appreciate Dr. Whitney Yin for giving me chance to working in her lab and mentoring me through my graduate program. Not only the scientific insights, also the warmness that she gave me and my family encouraged me to pursue my Ph. D. degree in the foreign country. I also would like to thank “a guru of molecular biology” Dr. Ian Molineux and “a guru of enzyme kinetics” Dr. Kenneth Johnson. Without their critical advice, I would not be accomplished my publication. I hope to be a respectable expert in my research field like them. I also should remember friendship and generosity given by many current and former Yin lab members: Hey-Ryung Chang, Qingchao “Eric” Meng, Xu Yang, Jeff Knight, Dr. Michio Matsunaga, Dr. He “River” Quan, Taewung Lee, Xin “Ella” Wang, Jamila Momand, and Max Shay. Most of all, I really appreciate my parents for their endless love and support, and my wife, In-Sook Jung, and my son, Jason Seung-Hyeon Lee who always stand by me with patients during my graduate carrier. -
DNA REPLICATION Objectives: 1
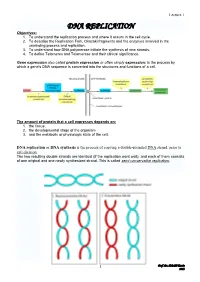
Lecture 1 DNA REPLICATION Objectives: 1. To understand the replication process and where it occurs in the cell cycle. 2. To describe the Replication Fork, Okazaki fragments and the enzymes involved in the unwinding process and replication. 3. To understand how DNA polymerase initiate the synthesis of new strands. 4. To define Telomeres and Telomerase and their clinical significance. Gene expression also called protein expression or often simply expression: is the process by which a gene's DNA sequence is converted into the structures and functions of a cell. The amount of protein that a cell expresses depends on: 1. the tissue, 2. the developmental stage of the organism 3. and the metabolic or physiologic state of the cell. DNA replication or DNA synthesis is the process of copying a double-stranded DNA strand, prior to cell division. The two resulting double strands are identical (if the replication went well), and each of them consists of one original and one newly synthesized strand. This is called semi conservative replication. 1 Prof. Dr. H.D.El-Yassin 2013 Lecture 1 The process of replication consists of three steps, initiation, replication and termination. 1. Prokaryotic replication Basic Requirement for DNA Synthesis 1. Substrates: the four deoxy nucleosides triphosphates are needed as substrates for DNA synthesis. Cleavage of the high-energy phosphate bond between the α and β phosphates provides the energy for the addition of the nucleotide. 2. Template: DNA replication cannot occur without a template. A template is required to direct the addition of the appropriate complementary deoxynucleotide to the newly synthesized DNA strand. -
DNA Polymerase Exchange and Lesion Bypass in Escherichia Coli
DNA Polymerase Exchange and Lesion Bypass in Escherichia Coli The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Kath, James Evon. 2016. DNA Polymerase Exchange and Lesion Bypass in Escherichia Coli. Doctoral dissertation, Harvard University, Graduate School of Arts & Sciences. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:26718716 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA ! ! ! ! ! ! ! DNA!polymerase!exchange!and!lesion!bypass!in!Escherichia)coli! ! A!dissertation!presented! by! James!Evon!Kath! to! The!Committee!on!Higher!Degrees!in!Biophysics! ! in!partial!fulfillment!of!the!requirements! for!the!degree!of! Doctor!of!Philosophy! in!the!subject!of! Biophysics! ! Harvard!University! Cambridge,!Massachusetts! October!2015! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ©!2015!L!James!E.!Kath.!Some!Rights!Reserved.! ! This!work!is!licensed!under!the!Creative!Commons!Attribution!3.0!United!States!License.!To! view!a!copy!of!this!license,!visit:!http://creativecommons.org/licenses/By/3.0/us! ! ! Dissertation!Advisor:!Professor!Joseph!J.!Loparo! ! ! !!!!!!!!James!Evon!Kath! ! DNA$polymerase$exchange$and$lesion$bypass$in$Escherichia)coli$ $ Abstract$ ! Translesion! synthesis! (TLS)! alleviates! -

PURIFIED THERMOSTABLE NUCLEIC ACID POLYMERASE ENZYME from $I(TERMOTOGA MARITIMA)
Europäisches Patentamt *EP000544789B1* (19) European Patent Office Office européen des brevets (11) EP 0 544 789 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.7: C12N 15/54, C12N 9/12 of the grant of the patent: 05.03.2003 Bulletin 2003/10 (86) International application number: PCT/US91/05753 (21) Application number: 91915802.2 (87) International publication number: (22) Date of filing: 13.08.1991 WO 92/003556 (05.03.1992 Gazette 1992/06) (54) PURIFIED THERMOSTABLE NUCLEIC ACID POLYMERASE ENZYME FROM $i(TERMOTOGA MARITIMA) GEREINIGTES THERMOSTABILES NUKLEINSÄURE-POLYMERASEENZYM AUS THERMOTOGA MARITIMA ENZYME D’ACIDE NUCLEIQUE THERMOSTABLE PURIFIEE PROVENANT DE L’EUBACTERIE $i(THERMOTOGA MARITIMA) (84) Designated Contracting States: (74) Representative: Poredda, Andreas et al AT BE CH DE DK ES FR GB GR IT LI LU NL SE Roche Diagnostics GmbH, Patentabteilung, (30) Priority: 13.08.1990 US 567244 Sandhofer Strasse 116 68305 Mannheim (DE) (43) Date of publication of application: 09.06.1993 Bulletin 1993/23 (56) References cited: • CHEMICAL ABSTRACTS, vol. 105, no. 5, 04 (73) Proprietor: F. HOFFMANN-LA ROCHE AG August 1986, Columbus, OH (US); R. HUBER et 4002 Basel (CH) al., p. 386, AN 38901u • JOURNAL OF BIOLOGICAL CHEMISTRY, vol. (72) Inventors: 264, no. 11, 15 April 1989, American Society for • GELFAND, David, H. Biochemistry & Molecular Biology Inc., Oakland, CA 94611 (US) Baltimore, MD (US); F.C. LAWYER et al., pp. • LAWYER, Frances, C. 6427-6437 Oakland, CA 94611 (US) • STOFFEL, Susanne El Cerrito, CA 94530 (US) Note: Within nine months from the publication of the mention of the grant of the European patent, any person may give notice to the European Patent Office of opposition to the European patent granted. -

Glucokinase Regulatory Protein Is Essential for the Proper Subcellular Localisation of Liver Glucokinase
FEBS Letters 456 (1999) 332^338 FEBS 22420 Glucokinase regulatory protein is essential for the proper subcellular localisation of liver glucokinase Nu¨ria de la Iglesiaa, Maria Veiga-da-Cunhab, Emile Van Schaftingenb, Joan J. Guinovarta, Juan C. Ferrera;* aDepartament de Bioqu|¨mica i Biologia Molecular, Universitat de Barcelona, Mart|¨ i Franque©s, 1, E-08028 Barcelona, Spain bLaboratory of Physiological Chemistry, Christian de Duve Institute of Cellular Pathology and Universite¨ Catholique de Louvain, B-1200 Brussels, Belgium Received in revised form 24 June 1999 pressed by fructose 1-phosphate, both of which bind to Abstract Glucokinase (GK), a key enzyme in the glucose homeostatic responses of the liver, changes its intracellular GKRP and modify its a¤nity for GK [4]. This 68 kDa protein localisation depending on the metabolic status of the cell. Rat is only found in the livers of species that express GK and liver GK and Xenopus laevis GK, fused to the green fluorescent although there is some evidence for its presence in pancreatic protein (GFP), concentrated in the nucleus of cultured rat tissue [5,6], a direct demonstration is still not available. hepatocytes at low glucose and translocated to the cytoplasm at In in vitro assays, rat GKRP can inhibit human pancreatic high glucose. Three mutant forms of Xenopus GK with reduced GK, which shows a high degree of identity to the rat liver affinity for GK regulatory protein (GKRP) did not concentrate isoform [7], as well as Xenopus laevis liver GK [8], a more in the hepatocyte nuclei, even at low glucose. In COS-1 and distantly related protein. -

Mutations That Separate the Functions of the Proofreading Subunit of the Escherichia Coli Replicase
G3: Genes|Genomes|Genetics Early Online, published on April 15, 2015 as doi:10.1534/g3.115.017285 Mutations that separate the functions of the proofreading subunit of the Escherichia coli replicase Zakiya Whatley*,1 and Kenneth N Kreuzer*§ *University Program in Genetics & Genomics, Duke University, Durham, NC 27705 §Department of Biochemistry, Duke University Medical Center, Durham, NC 27710 1 © The Author(s) 2013. Published by the Genetics Society of America. Running title: E. coli dnaQ separation of function mutants Keywords: DNA polymerase, epsilon subunit, linker‐scanning mutagenesis, mutation rate, SOS response Corresponding author: Kenneth N Kreuzer, Department of Biochemistry, Box 3711, Nanaline Duke Building, Research Drive, Duke University Medical Center, Durham, NC 27710 Phone: 919 684 6466 FAX: 919 684 6525 Email: [email protected] 1 Present address: Department of Biology, 300 N Washington Street, McCreary Hall, Campus Box 392, Gettysburg College, Gettysburg, PA 17325 Phone: 717 337 6160 Fax: 7171 337 6157 Email: [email protected] 2 ABSTRACT The dnaQ gene of Escherichia coli encodes the ε subunit of DNA polymerase III, which provides the 3’ 5’ exonuclease proofreading activity of the replicative polymerase. Prior studies have shown that loss of ε leads to high mutation frequency, partially constitutive SOS, and poor growth. In addition, a previous study from our lab identified dnaQ knockout mutants in a screen for mutants specifically defective in the SOS response following quinolone (nalidixic acid) treatment. To explain these results, we propose a model whereby in addition to proofreading, ε plays a distinct role in replisome disassembly and/or processing of stalled replication forks. -

1 Mechanistic Studies of DNA Replication, Lesion Bypass, And
Mechanistic Studies of DNA Replication, Lesion Bypass, and Editing Dissertation Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Austin Tyler Raper, B.S. The Ohio State University Biochemistry Graduate Program The Ohio State University 2018 Dissertation Committee Dr. Zucai Suo, Advisor Dr. Ross Dalbey Dr. Michael Poirier Dr. Richard Swenson 1 Copyrighted by Austin Tyler Raper 2018 2 Abstract DNA acts as a molecular blueprint for life. Adenosine, cytidine, guanosine, and thymidine nucleotides serve as the building blocks of DNA and can be arranged in near- endless combinations. These unique sequences of DNA may encode genes that when expressed produce RNA, proteins, and enzymes responsible for executing diverse tasks necessary for biological existence. Accordingly, careful maintenance of the molecular integrity of DNA is paramount for the growth, development, and functioning of organisms. However, DNA is damaged upon reaction with pervasive chemicals generated by normal cellular metabolism or encountered through the environment. The resulting DNA lesions act as roadblocks to high-fidelity A- and B-family DNA polymerases responsible for replicating DNA in preparation for cell division which may lead to programmed cell death. Additionally, these lesions may fool the polymerase into making errors during DNA replication, leading to genetic mutations and cancer. Fortunately, the cell has evolved DNA damage tolerance as an emergency response to such lesions. During DNA damage tolerance, a damage-stalled high-fidelity polymerase is substituted for a specialized Y-family polymerase, capable of bypassing the offending DNA lesion, for replication to continue. -

Analysis of a Multicomponent Thermostable DNA Polymerase III Replicase from an Extreme Thermophile*
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 277, No. 19, Issue of May 10, pp. 17334–17348, 2002 © 2002 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Analysis of a Multicomponent Thermostable DNA Polymerase III Replicase from an Extreme Thermophile* Received for publication, October 23, 2001, and in revised form, February 18, 2002 Published, JBC Papers in Press, February 21, 2002, DOI 10.1074/jbc.M110198200 Irina Bruck‡, Alexander Yuzhakov§¶, Olga Yurieva§, David Jeruzalmi§, Maija Skangalis‡§, John Kuriyan‡§, and Mike O’Donnell‡§ʈ From §The Rockefeller University and ‡Howard Hughes Medical Institute, New York, New York 10021 This report takes a proteomic/genomic approach to polymerase III (pol III) structure and function has been ob- characterize the DNA polymerase III replication appa- tained from studies of the Escherichia coli replicase, DNA ratus of the extreme thermophile, Aquifex aeolicus. polymerase III holoenzyme (reviewed in Ref. 6). Therefore, a Genes (dnaX, holA, and holB) encoding the subunits re- brief overview of its structure and function is instructive for the ␦ ␦ quired for clamp loading activity ( , , and ) were iden- comparisons to be made in this report. In E. coli, the catalytic Downloaded from tified. The dnaX gene produces only the full-length subunit of DNA polymerase III is the ␣ subunit (129.9 kDa) product, , and therefore differs from Escherichia coli encoded by dnaE; it lacks a proofreading exonuclease (7). The dnaX that produces two proteins (␥ and ). Nonetheless, Ј Ј ⑀ ␦␦ proofreading 3 –5 -exonuclease activity is contained in the the A. aeolicus proteins form a complex. The dnaN ␣ ,␦␦ (27.5 kDa) subunit (dnaQ) that forms a 1:1 complex with (8  gene encoding the clamp was identified, and the ␣Ϫ⑀  9). -

Structure of the Human Clamp Loader Reveals an Autoinhibited Conformation of a Substrate-Bound AAA+ Switch
Structure of the human clamp loader reveals an autoinhibited conformation of a substrate-bound AAA+ switch Christl Gaubitza,1, Xingchen Liua,b,1, Joseph Magrinoa,b, Nicholas P. Stonea, Jacob Landecka,b, Mark Hedglinc, and Brian A. Kelcha,2 aDepartment of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester MA 01605; bGraduate School of Biomedical Sciences, University of Massachusetts Medical School, Worcester MA 01605; and cDepartment of Chemistry, The Pennsylvania State University, University Park, PA 16802 Edited by Michael E. O’Donnell, HHMI and Rockefeller University, New York, NY, and approved July 27, 2020 (received for review April 20, 2020) DNA replication requires the sliding clamp, a ring-shaped protein areflexia syndrome (15), Hutchinson–Gilford progeria syn- complex that encircles DNA, where it acts as an essential cofactor drome (16), and in the replication of some viruses (17–19). It for DNA polymerases and other proteins. The sliding clamp needs is unknown whether loading by RFC contributes to PARD to be opened and installed onto DNA by a clamp loader ATPase of disease. the AAA+ family. The human clamp loader replication factor C Clamp loaders are members of the AAA+ family of ATPases (RFC) and sliding clamp proliferating cell nuclear antigen (PCNA) (ATPases associated with various cellular activities), a large are both essential and play critical roles in several diseases. De- protein family that uses the chemical energy of adenosine 5′- spite decades of study, no structure of human RFC has been re- triphosphate (ATP) to generate mechanical force (20). Most solved. Here, we report the structure of human RFC bound to AAA+ proteins form hexameric motors that use an undulating PCNA by cryogenic electron microscopy to an overall resolution ∼ spiral staircase mechanism to processively translocate a substrate of 3.4 Å. -

Arthur Kornberg Discovered (The First) DNA Polymerase Four
Arthur Kornberg discovered (the first) DNA polymerase Using an “in vitro” system for DNA polymerase activity: 1. Grow E. coli 2. Break open cells 3. Prepare soluble extract 4. Fractionate extract to resolve different proteins from each other; repeat; repeat 5. Search for DNA polymerase activity using an biochemical assay: incorporate radioactive building blocks into DNA chains Four requirements of DNA-templated (DNA-dependent) DNA polymerases • single-stranded template • deoxyribonucleotides with 5’ triphosphate (dNTPs) • magnesium ions • annealed primer with 3’ OH Synthesis ONLY occurs in the 5’-3’ direction Fig 4-1 E. coli DNA polymerase I 5’-3’ polymerase activity Primer has a 3’-OH Incoming dNTP has a 5’ triphosphate Pyrophosphate (PP) is lost when dNMP adds to the chain E. coli DNA polymerase I: 3 separable enzyme activities in 3 protein domains 5’-3’ polymerase + 3’-5’ exonuclease = Klenow fragment N C 5’-3’ exonuclease Fig 4-3 E. coli DNA polymerase I 3’-5’ exonuclease Opposite polarity compared to polymerase: polymerase activity must stop to allow 3’-5’ exonuclease activity No dNTP can be re-made in reversed 3’-5’ direction: dNMP released by hydrolysis of phosphodiester backboneFig 4-4 Proof-reading (editing) of misincorporated 3’ dNMP by the 3’-5’ exonuclease Fidelity is accuracy of template-cognate dNTP selection. It depends on the polymerase active site structure and the balance of competing polymerase and exonuclease activities. A mismatch disfavors extension and favors the exonuclease.Fig 4-5 Superimposed structure of the Klenow fragment of DNA pol I with two different DNAs “Fingers” “Thumb” “Palm” red/orange helix: 3’ in red is elongating blue/cyan helix: 3’ in blue is getting edited Fig 4-6 E. -

DNA POLYMERASE III HOLOENZYME: Structure and Function of a Chromosomal Replicating Machine
Annu. Rev. Biochem. 1995.64:171-200 Copyright Ii) 1995 byAnnual Reviews Inc. All rights reserved DNA POLYMERASE III HOLOENZYME: Structure and Function of a Chromosomal Replicating Machine Zvi Kelman and Mike O'Donnell} Microbiology Department and Hearst Research Foundation. Cornell University Medical College. 1300York Avenue. New York. NY }0021 KEY WORDS: DNA replication. multis ubuni t complexes. protein-DNA interaction. DNA-de penden t ATPase . DNA sliding clamps CONTENTS INTRODUCTION........................................................ 172 THE HOLO EN ZYM E PARTICL E. .......................................... 173 THE CORE POLYMERASE ............................................... 175 THE � DNA SLIDING CLAM P............... ... ......... .................. 176 THE yC OMPLEX MATCHMAKER......................................... 179 Role of ATP . .... .............. ...... ......... ..... ............ ... 179 Interaction of y Complex with SSB Protein .................. ............... 181 Meclwnism of the yComplex Clamp Loader ................................ 181 Access provided by Rockefeller University on 08/07/15. For personal use only. THE 't SUBUNIT . .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. .. 182 Annu. Rev. Biochem. 1995.64:171-200. Downloaded from www.annualreviews.org AS YMMETRIC STRUC TURE OF HOLO EN ZYM E . 182 DNA PO LYM ER AS E III HOLO ENZ YME AS A REPLIC ATING MACHINE ....... 186 Exclwnge of � from yComplex to Core .................................... 186 Cycling of Holoenzyme on the LaggingStrand -

Distinct Co-Evolution Patterns of Genes Associated to DNA Polymerase III Dnae and Polc Stefan Engelen1,2, David Vallenet2, Claudine Médigue2 and Antoine Danchin1,3*
Engelen et al. BMC Genomics 2012, 13:69 http://www.biomedcentral.com/1471-2164/13/69 RESEARCHARTICLE Open Access Distinct co-evolution patterns of genes associated to DNA polymerase III DnaE and PolC Stefan Engelen1,2, David Vallenet2, Claudine Médigue2 and Antoine Danchin1,3* Abstract Background: Bacterial genomes displaying a strong bias between the leading and the lagging strand of DNA replication encode two DNA polymerases III, DnaE and PolC, rather than a single one. Replication is a highly unsymmetrical process, and the presence of two polymerases is therefore not unexpected. Using comparative genomics, we explored whether other processes have evolved in parallel with each polymerase. Results: Extending previous in silico heuristics for the analysis of gene co-evolution, we analyzed the function of genes clustering with dnaE and polC. Clusters were highly informative. DnaE co-evolves with the ribosome, the transcription machinery, the core of intermediary metabolism enzymes. It is also connected to the energy-saving enzyme necessary for RNA degradation, polynucleotide phosphorylase. Most of the proteins of this co-evolving set belong to the persistent set in bacterial proteomes, that is fairly ubiquitously distributed. In contrast, PolC co- evolves with RNA degradation enzymes that are present only in the A+T-rich Firmicutes clade, suggesting at least two origins for the degradosome. Conclusion: DNA replication involves two machineries, DnaE and PolC. DnaE co-evolves with the core functions of bacterial life. In contrast PolC co-evolves with a set of RNA degradation enzymes that does not derive from the degradosome identified in gamma-Proteobacteria. This suggests that at least two independent RNA degradation pathways existed in the progenote community at the end of the RNA genome world.