Locating Protein-Coding Sequences Under Selection for Additional, Overlapping Functions in 29 Mammalian Genomes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Investigating the Genetic Basis of Cisplatin-Induced Ototoxicity in Adult South African Patients
--------------------------------------------------------------------------- Investigating the genetic basis of cisplatin-induced ototoxicity in adult South African patients --------------------------------------------------------------------------- by Timothy Francis Spracklen SPRTIM002 SUBMITTED TO THE UNIVERSITY OF CAPE TOWN In fulfilment of the requirements for the degree MSc(Med) Faculty of Health Sciences UNIVERSITY OF CAPE TOWN University18 December of Cape 2015 Town Supervisor: Prof. Rajkumar S Ramesar Co-supervisor: Ms A Alvera Vorster Division of Human Genetics, Department of Pathology, University of Cape Town 1 The copyright of this thesis vests in the author. No quotation from it or information derived from it is to be published without full acknowledgement of the source. The thesis is to be used for private study or non- commercial research purposes only. Published by the University of Cape Town (UCT) in terms of the non-exclusive license granted to UCT by the author. University of Cape Town Declaration I, Timothy Spracklen, hereby declare that the work on which this dissertation/thesis is based is my original work (except where acknowledgements indicate otherwise) and that neither the whole work nor any part of it has been, is being, or is to be submitted for another degree in this or any other university. I empower the university to reproduce for the purpose of research either the whole or any portion of the contents in any manner whatsoever. Signature: Date: 18 December 2015 ' 2 Contents Abbreviations ………………………………………………………………………………….. 1 List of figures …………………………………………………………………………………... 6 List of tables ………………………………………………………………………………….... 7 Abstract ………………………………………………………………………………………… 10 1. Introduction …………………………………………………………………………………. 11 1.1 Cancer …………………………………………………………………………….. 11 1.2 Adverse drug reactions ………………………………………………………….. 12 1.3 Cisplatin …………………………………………………………………………… 12 1.3.1 Cisplatin’s mechanism of action ……………………………………………… 13 1.3.2 Adverse reactions to cisplatin therapy ………………………………………. -

A 1Bp Deletion in the Dual Reading Frame Deafness Gene LRTOMT Causes a Frameshift from the First Into the Second Reading Frame
RESEARCH LETTER A 1 bp Deletion in the Dual Reading Frame Deafness Gene LRTOMT Causes a Frameshift From the First Into the Second Reading Frame Maarten Vanwesemael,1 Isabelle Schrauwen,1 Ruben Ceuppens,1 Fatemeh Alasti,1 Ellen Jorssen,1 Effat Farrokhi,2 Morteza Hashemzadeh Chaleshtori,2 and Guy Van Camp1* 1Department of Medical Genetics, University of Antwerp, 2610, Wilrijk, Belgium 2Cellular and Molecular Research Center, School of Medicine, Shahrekord University of Medical Sciences, Shahrekord, Iran Received 29 November 2010; Accepted 12 April 2011 TO THE EDITOR: How to Cite this Article: Congenital hearing impairment (HI) affects one in 650 newborns Vanwesemael M, Schrauwen I, Ceuppens R, and has a genetic cause in 50% of the cases. Autosomal recessive Alasti F, Jorssen E, Farrokhi E, Chaleshtori non-syndromic hearing impairment (ARNSHI) is responsible for MH, Van Camp G. 2011. A 1 bp deletion in the 70–80% of all hereditary cases of HI. ARNSHI is genetically highly dual reading frame deafness gene LRTOMT heterogeneous and until today mutations in 37 genes have been causes a frameshift from the first into the identified [Van Camp and Smith, 2011]. One of the latest genes for second reading frame. ARNSHI is LRTOMT (at locus DFNB63 on 11q13.3-q13.4). The Am J Med Genet Part A 155:2021–2023. structure of human LRTOMT has only recently been elucidated [Ahmed et al., 2008]. LRTOMT is an evolutionary recent fusion gene with alternative reading frames that exclusively exists in analysis was performed on an ABI 3130XL DNA sequence primates and is a fusion of non-primate Lrrc51 (Leucine rich repeat analyzer (Applied Biosystems Inc., Foster City, CA), using standard containing 51) and Tomt (Transmembrane O-methyltransferase) procedures. -

Sharmin Supple Legend 150706
Supplemental data Supplementary Figure 1 Generation of NPHS1-GFP iPS cells (A) TALEN activity tested in HEK 293 cells. The targeted region was PCR-amplified and cloned. Deletions in the NPHS1 locus were detected in four clones out of 10 that were sequenced. (B) PCR screening of human iPS cell homologous recombinants (C) Southern blot screening of human iPS cell homologous recombinants Supplementary Figure 2 Human glomeruli generated from NPHS1-GFP iPS cells (A) Morphological changes of GFP-positive glomeruli during differentiation in vitro. A different aggregate from the one shown in Figure 2 is presented. Lower panels: higher magnification of the areas marked by rectangles in the upper panels. Note the shape changes of the glomerulus (arrowheads). Scale bars: 500 µm. (B) Some, but not all, of the Bowman’s capsule cells were positive for nephrin (48E11 antibody: magenta) and GFP (green). Scale bars: 10 µm. Supplementary Figure 3 Histology of human podocytes generated in vitro (A) Transmission electron microscopy of the foot processes. Lower magnification of Figure 4H. Scale bars: 500 nm. (B) (C) The slit diaphragm between the foot processes. Higher magnification of the 1 regions marked by rectangles in panel A. Scale bar: 100 nm. (D) Absence of mesangial or vascular endothelial cells in the induced glomeruli. Anti-PDGFRβ and CD31 antibodies were used to detect the two lineages, respectively, and no positive signals were observed in the glomeruli. Podocytes are positive for WT1. Nuclei are counterstained with Nuclear Fast Red. Scale bars: 20 µm. Supplementary Figure 4 Cluster analysis of gene expression in various human tissues (A) Unbiased cluster analysis across various human tissues using the top 300 genes enriched in GFP-positive podocytes. -

Human Leucine-Rich Repeat Proteins: a Genome-Wide Bioinformatic Categorization and Functional Analysis in Innate Immunity
Human leucine-rich repeat proteins: a genome-wide bioinformatic categorization and functional analysis in innate immunity Aylwin C. Y. Nga,b,1, Jason M. Eisenberga,b,1, Robert J. W. Heatha, Alan Huetta, Cory M. Robinsonc, Gerard J. Nauc, and Ramnik J. Xaviera,b,2 aCenter for Computational and Integrative Biology, and Gastrointestinal Unit, Massachusetts General Hospital and Harvard Medical School, Boston, MA 02114; bThe Broad Institute of Massachusetts Institute of Technology and Harvard, Cambridge, MA 02142; and cMicrobiology and Molecular Genetics, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261 Edited by Jeffrey I. Gordon, Washington University School of Medicine, St. Louis, MO, and approved June 11, 2010 (received for review February 17, 2010) In innate immune sensing, the detection of pathogen-associated proteins have been implicated in human diseases to date, notably molecular patterns by recognition receptors typically involve polymorphisms in NOD2 in Crohn disease (8, 9), CIITA in leucine-rich repeats (LRRs). We provide a categorization of 375 rheumatoid arthritis and multiple sclerosis (10), and TLR5 in human LRR-containing proteins, almost half of which lack other Legionnaire disease (11). identifiable functional domains. We clustered human LRR proteins Most LRR domains consist of a chain of between 2 and 45 by first assigning LRRs to LRR classes and then grouping the proteins LRRs (12). Each repeat in turn is typically 20 to 30 residues long based on these class assignments, revealing several of the resulting and can be divided into a highly conserved segment (HCS) fol- protein groups containing a large number of proteins with certain lowed by a variable segment (VS). -

Integration of Tmc1/2 Into the Mechanotransduction Complex in Zebrafish Hair Cells Is Regulated by Transmembrane O-Methyltransferase (Tomt)
This is a repository copy of Integration of Tmc1/2 into the mechanotransduction complex in zebrafish hair cells is regulated by Transmembrane O-methyltransferase (Tomt).. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/117038/ Version: Accepted Version Article: Erickson, T. orcid.org/0000-0002-0910-2535, Morgan, C.P, Olt, J. et al. (9 more authors) (2017) Integration of Tmc1/2 into the mechanotransduction complex in zebrafish hair cells is regulated by Transmembrane O-methyltransferase (Tomt). Elife, 6. https://doi.org/10.7554/eLife.28474 Reuse Unless indicated otherwise, fulltext items are protected by copyright with all rights reserved. The copyright exception in section 29 of the Copyright, Designs and Patents Act 1988 allows the making of a single copy solely for the purpose of non-commercial research or private study within the limits of fair dealing. The publisher or other rights-holder may allow further reproduction and re-use of this version - refer to the White Rose Research Online record for this item. Where records identify the publisher as the copyright holder, users can verify any specific terms of use on the publisher’s website. Takedown If you consider content in White Rose Research Online to be in breach of UK law, please notify us by emailing [email protected] including the URL of the record and the reason for the withdrawal request. [email protected] https://eprints.whiterose.ac.uk/ Title Integration of Tmc1/2 into the mechanotransduction complex in zebrafish hair cells is regulated by Transmembrane O-methyltransferase (Tomt) Running title Tomt is required for mechanotransduction Authors Timothy Erickson1, Clive P. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

The Hypothalamus As a Hub for SARS-Cov-2 Brain Infection and Pathogenesis
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The hypothalamus as a hub for SARS-CoV-2 brain infection and pathogenesis Sreekala Nampoothiri1,2#, Florent Sauve1,2#, Gaëtan Ternier1,2ƒ, Daniela Fernandois1,2 ƒ, Caio Coelho1,2, Monica ImBernon1,2, Eleonora Deligia1,2, Romain PerBet1, Vincent Florent1,2,3, Marc Baroncini1,2, Florence Pasquier1,4, François Trottein5, Claude-Alain Maurage1,2, Virginie Mattot1,2‡, Paolo GiacoBini1,2‡, S. Rasika1,2‡*, Vincent Prevot1,2‡* 1 Univ. Lille, Inserm, CHU Lille, Lille Neuroscience & Cognition, DistAlz, UMR-S 1172, Lille, France 2 LaBoratorY of Development and PlasticitY of the Neuroendocrine Brain, FHU 1000 daYs for health, EGID, School of Medicine, Lille, France 3 Nutrition, Arras General Hospital, Arras, France 4 Centre mémoire ressources et recherche, CHU Lille, LiCEND, Lille, France 5 Univ. Lille, CNRS, INSERM, CHU Lille, Institut Pasteur de Lille, U1019 - UMR 8204 - CIIL - Center for Infection and ImmunitY of Lille (CIIL), Lille, France. # and ƒ These authors contriButed equallY to this work. ‡ These authors directed this work *Correspondence to: [email protected] and [email protected] Short title: Covid-19: the hypothalamic hypothesis 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Delineation of Homozygous Variants Associated with Prelingual Sensorineural Hearing Loss in Pakistani Families
G C A T T A C G G C A T genes Communication Delineation of Homozygous Variants Associated with Prelingual Sensorineural Hearing Loss in Pakistani Families 1, 2,3, 1 1,2, Muhammad Noman y, Rafaqat Ishaq y, Shazia A. Bukhari , Zubair M. Ahmed * and Saima Riazuddin 2,* 1 Department of Biochemistry, Government College University, Faisalabad 38000, Pakistan 2 Department of Otorhinolaryngology Head and Neck Surgery, University of Maryland School of Medicine, Baltimore, MD 21201, USA 3 University Institute of Biochemistry & Biotechnology, PMAS-Arid Agriculture University, Rawalpindi 46000, Pakistan * Correspondence: [email protected] (Z.M.A); [email protected] (S.R.) These authors contribute equally to this work. y Received: 6 November 2019; Accepted: 4 December 2019; Published: 10 December 2019 Abstract: Hearing loss is a genetically heterogeneous disorder affecting approximately 360 million people worldwide and is among the most common sensorineural disorders. Here, we report a genetic analysis of seven large consanguineous families segregating prelingual sensorineural hearing loss. Whole-exome sequencing (WES) revealed seven different pathogenic variants segregating with hearing loss in these families, three novel variants (c.1204G>A, c.322G>T, and c.5587C>T) in TMPRSS3, ESRRB, and OTOF, and four previously reported variants (c.208C>T, c.6371G>A, c.226G>A, and c.494C>T) in LRTOMT, MYO15A, KCNE1, and LHFPL5, respectively. All identified variants had very low frequencies in the control databases and were predicted to have pathogenic effects on the encoded proteins. In addition to being familial, we also found intersibship locus heterogeneity in the evaluated families. The known pathogenic c.226C>T variant identified in KCNE1 only segregates with the hearing loss phenotype in a subset of affected members of the family GCNF21. -

An Epigenetic Reprogramming Strategy to Resensitize
Published OnlineFirst March 16, 2016; DOI: 10.1158/0008-5472.CAN-15-2116 Cancer Molecular and Cellular Pathobiology Research An Epigenetic Reprogramming Strategy to Resensitize Radioresistant Prostate Cancer Cells Claudia Peitzsch1,MonicaCojoc1, Linda Hein1,InaKurth1,KatrinMabert€ 1, Franziska Trautmann1, Barbara Klink2,3,4,EvelinSchrock€ 2,3,4, Manfred P. Wirth5, Mechthild Krause1,3,4,6,7, Eduard A. Stakhovsky8,GennadyD.Telegeev9,VladimirNovotny5, Marieta Toma10, Michael Muders10, Gustavo B. Baretton10, Fiona M. Frame11, Norman J. Maitland11, Michael Baumann1,3,4,6,7, and Anna Dubrovska1,3,4,7 Abstract Radiotherapy is a mainstay of curative prostate cancer treat- switch occurred during a course of radiotherapy that was ment, but risks of recurrence after treatment remain significant associated with stable genetic and epigenetic changes. Specif- in locally advanced disease. Given that tumor relapse can be ically, we found that irradiation triggered histone H3 meth- attributed to a population of cancer stem cells (CSC) that ylation at the promoter of the CSC marker aldehyde dehydro- survives radiotherapy, analysis of this cell population might genase 1A1 (ALDH1A1), stimulating its gene transcription. illuminate tactics to personalize treatment. However, this Inhibiting this methylation event triggered apoptosis, promot- direction remains challenging given the plastic nature of pros- ed radiosensitization, and hindered tumorigenicity of radio- tate cancers following treatment. We show here that irradiating resistant prostate cancer cells. Overall, our results suggest that prostate cancer cells stimulates a durable upregulation of stem epigenetic therapies may restore the cytotoxic effects of irra- cell markers that epigenetically reprogram these cells. In both diation in radioresistant CSC populations. Cancer Res; 76(9); tumorigenic and radioresistant cell populations, a phenotypic 2637–51. -
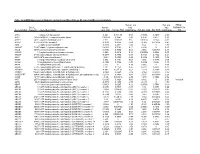
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -

Mutations of the Gene Encoding Otogelin Are a Cause of Autosomal-Recessive Nonsyndromic Moderate Hearing Impairment
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector REPORT Mutations of the Gene Encoding Otogelin Are a Cause of Autosomal-Recessive Nonsyndromic Moderate Hearing Impairment Margit Schraders,1,2,3 Laura Ruiz-Palmero,5,6 Ersan Kalay,7 Jaap Oostrik,1,2,3 Francisco J. del Castillo,5,6 Orhan Sezgin,8 Andy J. Beynon,1,3 Tim M. Strom,9 Ronald J.E. Pennings,1,3 Celia Zazo Seco,1,2,3 Anne M.M. Oonk,1,3 Henricus P.M. Kunst,1,3 Marı´a Domı´nguez-Ruiz,5,6 Ana M. Garcı´a-Arumi,10 Miguel del Campo,11 Manuela Villamar,5,6 Lies H. Hoefsloot,4 Felipe Moreno,5,6 Ronald J.C. Admiraal,1,3 Ignacio del Castillo,5,6 and Hannie Kremer1,2,3,4,* Already 40 genes have been identified for autosomal-recessive nonsyndromic hearing impairment (arNSHI); however, many more genes are still to be identified. In a Dutch family segregating arNSHI, homozygosity mapping revealed a 2.4 Mb homozygous region on chro- mosome 11 in p15.1-15.2, which partially overlapped with the previously described DFNB18 locus. However, no putative pathogenic variants were found in USH1C, the gene mutated in DFNB18 hearing impairment. The homozygous region contained 12 additional annotated genes including OTOG, the gene encoding otogelin, a component of the tectorial membrane. It is thought that otogelin contributes to the stability and strength of this membrane through interaction or stabilization of its constituent fibers. The murine orthologous gene was already known to cause hearing loss when defective. -

Human Leucine-Rich Repeat Proteins: a Genome-Wide Bioinformatic Categorization and Functional Analysis in Innate Immunity
Human leucine-rich repeat proteins: a genome-wide bioinformatic categorization and functional analysis in innate immunity Aylwin C. Y. Nga,b,1, Jason M. Eisenberga,b,1, Robert J. W. Heatha, Alan Huetta, Cory M. Robinsonc, Gerard J. Nauc, and Ramnik J. Xaviera,b,2 aCenter for Computational and Integrative Biology, and Gastrointestinal Unit, Massachusetts General Hospital and Harvard Medical School, Boston, MA 02114; bThe Broad Institute of Massachusetts Institute of Technology and Harvard, Cambridge, MA 02142; and cMicrobiology and Molecular Genetics, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261 Edited by Jeffrey I. Gordon, Washington University School of Medicine, St. Louis, MO, and approved June 11, 2010 (received for review February 17, 2010) In innate immune sensing, the detection of pathogen-associated proteins have been implicated in human diseases to date, notably molecular patterns by recognition receptors typically involve polymorphisms in NOD2 in Crohn disease (8, 9), CIITA in leucine-rich repeats (LRRs). We provide a categorization of 375 rheumatoid arthritis and multiple sclerosis (10), and TLR5 in human LRR-containing proteins, almost half of which lack other Legionnaire disease (11). identifiable functional domains. We clustered human LRR proteins Most LRR domains consist of a chain of between 2 and 45 by first assigning LRRs to LRR classes and then grouping the proteins LRRs (12). Each repeat in turn is typically 20 to 30 residues long based on these class assignments, revealing several of the resulting and can be divided into a highly conserved segment (HCS) fol- protein groups containing a large number of proteins with certain lowed by a variable segment (VS).