Phylogeny and Species Delimitation of Near Eastern Neurergus Newts (Salamandridae) Based on Genome-Wide Radseq Data Analysis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Review of Southern Iraq Herpetofauna
Vol. 3 (1): 61-71, 2019 A Review of Southern Iraq Herpetofauna Nadir A. Salman Mazaya University College, Dhi Qar, Iraq *Corresponding author: [email protected] Abstract: The present review discussed the species diversity of herpetofauna in southern Iraq due to their scientific and national interests. The review includes a historical record for the herpetofaunal studies in Iraq since the earlier investigations of the 1920s and 1950s along with the more recent taxonomic trials in the following years. It appeared that, little is known about Iraqi herpetofauna, and no comprehensive checklist has been done for these species. So far, 96 species of reptiles and amphibians have been recorded from Iraq, but only a relatively small proportion of them occur in the southern marshes. The marshes act as key habitat for globally endangered species and as a potential for as yet unexplored amphibian and reptile diversity. Despite the lack of precise localities, the tree frog Hyla savignyi, the marsh frog Pelophylax ridibunda and the green toad Bufo viridis are found in the marshes. Common reptiles in the marshes include the Caspian terrapin (Clemmys caspia), the soft-shell turtle (Trionyx euphraticus), the Euphrates softshell turtle (Rafetus euphraticus), geckos of the genus Hemidactylus, two species of skinks (Trachylepis aurata and Mabuya vittata) and a variety of snakes of the genus Coluber, the spotted sand boa (Eryx jaculus), tessellated water snake (Natrix tessellata) and Gray's desert racer (Coluber ventromaculatus). More recently, a new record for the keeled gecko, Cyrtopodion scabrum and the saw-scaled viper (Echis carinatus sochureki) was reported. The IUCN Red List includes six terrestrial and six aquatic amphibian species. -

Effects of Emerging Infectious Diseases on Amphibians: a Review of Experimental Studies
diversity Review Effects of Emerging Infectious Diseases on Amphibians: A Review of Experimental Studies Andrew R. Blaustein 1,*, Jenny Urbina 2 ID , Paul W. Snyder 1, Emily Reynolds 2 ID , Trang Dang 1 ID , Jason T. Hoverman 3 ID , Barbara Han 4 ID , Deanna H. Olson 5 ID , Catherine Searle 6 ID and Natalie M. Hambalek 1 1 Department of Integrative Biology, Oregon State University, Corvallis, OR 97331, USA; [email protected] (P.W.S.); [email protected] (T.D.); [email protected] (N.M.H.) 2 Environmental Sciences Graduate Program, Oregon State University, Corvallis, OR 97331, USA; [email protected] (J.U.); [email protected] (E.R.) 3 Department of Forestry and Natural Resources, Purdue University, West Lafayette, IN 47907, USA; [email protected] 4 Cary Institute of Ecosystem Studies, Millbrook, New York, NY 12545, USA; [email protected] 5 US Forest Service, Pacific Northwest Research Station, Corvallis, OR 97331, USA; [email protected] 6 Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA; [email protected] * Correspondence [email protected]; Tel.: +1-541-737-5356 Received: 25 May 2018; Accepted: 27 July 2018; Published: 4 August 2018 Abstract: Numerous factors are contributing to the loss of biodiversity. These include complex effects of multiple abiotic and biotic stressors that may drive population losses. These losses are especially illustrated by amphibians, whose populations are declining worldwide. The causes of amphibian population declines are multifaceted and context-dependent. One major factor affecting amphibian populations is emerging infectious disease. Several pathogens and their associated diseases are especially significant contributors to amphibian population declines. -

50 CFR Ch. I (10–1–20 Edition) § 16.14
§ 15.41 50 CFR Ch. I (10–1–20 Edition) Species Common name Serinus canaria ............................................................. Common Canary. 1 Note: Permits are still required for this species under part 17 of this chapter. (b) Non-captive-bred species. The list 16.14 Importation of live or dead amphib- in this paragraph includes species of ians or their eggs. non-captive-bred exotic birds and coun- 16.15 Importation of live reptiles or their tries for which importation into the eggs. United States is not prohibited by sec- Subpart C—Permits tion 15.11. The species are grouped tax- onomically by order, and may only be 16.22 Injurious wildlife permits. imported from the approved country, except as provided under a permit Subpart D—Additional Exemptions issued pursuant to subpart C of this 16.32 Importation by Federal agencies. part. 16.33 Importation of natural-history speci- [59 FR 62262, Dec. 2, 1994, as amended at 61 mens. FR 2093, Jan. 24, 1996; 82 FR 16540, Apr. 5, AUTHORITY: 18 U.S.C. 42. 2017] SOURCE: 39 FR 1169, Jan. 4, 1974, unless oth- erwise noted. Subpart E—Qualifying Facilities Breeding Exotic Birds in Captivity Subpart A—Introduction § 15.41 Criteria for including facilities as qualifying for imports. [Re- § 16.1 Purpose of regulations. served] The regulations contained in this part implement the Lacey Act (18 § 15.42 List of foreign qualifying breed- U.S.C. 42). ing facilities. [Reserved] § 16.2 Scope of regulations. Subpart F—List of Prohibited Spe- The provisions of this part are in ad- cies Not Listed in the Appen- dition to, and are not in lieu of, other dices to the Convention regulations of this subchapter B which may require a permit or prescribe addi- § 15.51 Criteria for including species tional restrictions or conditions for the and countries in the prohibited list. -

Taxonomic Checklist of Amphibian Species Listed in the CITES
CoP17 Doc. 81.1 Annex 5 (English only / Únicamente en inglés / Seulement en anglais) Taxonomic Checklist of Amphibian Species listed in the CITES Appendices and the Annexes of EC Regulation 338/97 Species information extracted from FROST, D. R. (2015) "Amphibian Species of the World, an online Reference" V. 6.0 (as of May 2015) Copyright © 1998-2015, Darrel Frost and TheAmericanMuseum of Natural History. All Rights Reserved. Additional comments included by the Nomenclature Specialist of the CITES Animals Committee (indicated by "NC comment") Reproduction for commercial purposes prohibited. CoP17 Doc. 81.1 Annex 5 - p. 1 Amphibian Species covered by this Checklist listed by listed by CITES EC- as well as Family Species Regulation EC 338/97 Regulation only 338/97 ANURA Aromobatidae Allobates femoralis X Aromobatidae Allobates hodli X Aromobatidae Allobates myersi X Aromobatidae Allobates zaparo X Aromobatidae Anomaloglossus rufulus X Bufonidae Altiphrynoides malcolmi X Bufonidae Altiphrynoides osgoodi X Bufonidae Amietophrynus channingi X Bufonidae Amietophrynus superciliaris X Bufonidae Atelopus zeteki X Bufonidae Incilius periglenes X Bufonidae Nectophrynoides asperginis X Bufonidae Nectophrynoides cryptus X Bufonidae Nectophrynoides frontierei X Bufonidae Nectophrynoides laevis X Bufonidae Nectophrynoides laticeps X Bufonidae Nectophrynoides minutus X Bufonidae Nectophrynoides paulae X Bufonidae Nectophrynoides poyntoni X Bufonidae Nectophrynoides pseudotornieri X Bufonidae Nectophrynoides tornieri X Bufonidae Nectophrynoides vestergaardi -
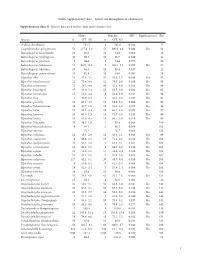
I Online Supplementary Data – Sexual Size Dimorphism in Salamanders
Online Supplementary data – Sexual size dimorphism in salamanders Supplementary data S1. Species data used in this study and references list. Males Females SSD Significant test Ref Species n SVL±SD n SVL±SD Andrias davidianus 2 532.5 8 383.0 -0.280 12 Cryptobranchus alleganiensis 53 277.4±5.2 52 300.9±3.4 0.084 Yes 61 Batrachuperus karlschmidti 10 80.0 10 84.8 0.060 26 Batrachuperus londongensis 20 98.6 10 96.7 -0.019 12 Batrachuperus pinchonii 5 69.6 5 74.6 0.070 26 Batrachuperus taibaiensis 11 92.9±12.1 9 102.1±7.1 0.099 Yes 27 Batrachuperus tibetanus 10 94.5 10 92.8 -0.017 12 Batrachuperus yenyuadensis 10 82.8 10 74.8 -0.096 26 Hynobius abei 24 57.8±2.1 34 55.0±1.2 -0.048 Yes 92 Hynobius amakusaensis 22 75.4±4.8 12 76.5±3.6 0.014 No 93 Hynobius arisanensis 72 54.3±4.8 40 55.2±4.8 0.016 No 94 Hynobius boulengeri 37 83.0±5.4 15 91.5±3.8 0.102 Yes 95 Hynobius formosanus 15 53.0±4.4 8 52.4±3.9 -0.011 No 94 Hynobius fuca 4 50.9±2.8 3 52.8±2.0 0.037 No 94 Hynobius glacialis 12 63.1±4.7 11 58.9±5.2 -0.066 No 94 Hynobius hidamontanus 39 47.7±1.0 15 51.3±1.2 0.075 Yes 96 Hynobius katoi 12 58.4±3.3 10 62.7±1.6 0.073 Yes 97 Hynobius kimurae 20 63.0±1.5 15 72.7±2.0 0.153 Yes 98 Hynobius leechii 70 61.6±4.5 18 66.5±5.9 0.079 Yes 99 Hynobius lichenatus 37 58.5±1.9 2 53.8 -0.080 100 Hynobius maoershanensis 4 86.1 2 80.1 -0.069 101 Hynobius naevius 72.1 76.7 0.063 102 Hynobius nebulosus 14 48.3±2.9 12 50.4±2.1 0.043 Yes 96 Hynobius osumiensis 9 68.4±3.1 15 70.2±3.0 0.026 No 103 Hynobius quelpaertensis 41 52.5±3.8 4 61.3±4.1 0.167 Yes 104 Hynobius -

EC) No 338/97 on the Protection of Species of Wild Fauna and Flora by Regulating Trade Therein
12.8.2010 EN Official Journal of the European Union L 212/1 II (Non-legislative acts) REGULATIONS COMMISSION REGULATION (EU) No 709/2010 of 22 July 2010 amending Council Regulation (EC) No 338/97 on the protection of species of wild fauna and flora by regulating trade therein THE EUROPEAN COMMISSION, (7) The species Ctenosaura bakeri, C. oedirhina, C. melanosterna, C. palearis, Agalychnis spp., Dynastes satanas, Operculicarya hyphaenoides, O. pachypus, Zygosicyos pubescens, Z. Having regard to the Treaty on the Functioning of the European tripartitus, Aniba rosaeodora (with annotation), Adenia Union, olaboensis, Cyphostemma elephantopus, C. montagnacii and Bulnesia sarmientoi (with annotation) have been included in Appendix II. Having regard to Council Regulation (EC) No 338/97 of 9 December 1996 on the protection of species of wild fauna 1 and flora by regulating trade therein ( ), and in particular (8) The Appendix II listing of Beccariophoenix madagascariensis Article 19(5) thereof, and Neodypsis decaryi was extended to include seeds from Madagascar. Whereas: (9) The following species have been deleted from Appendix III to the Convention at the request of Malaysia: Arbo (1) Regulation (EC) No 338/97 lists animal and plant species rophila campbelli, Arborophila charltonii, Caloperdix oculeus, in respect of which trade is restricted or controlled. Lophura erythrophthalma, Lophura ignita, Melanoperdix niger, Those lists incorporate the lists set out in the Appendices Polyplectron inopinatum, Rhizothera dulitensis, Rhizothera to the Convention on International Trade in Endangered longirostris and Rollulus rouloul, and the species Haliotis Species of Wild Fauna and Flora, hereinafter ‘the midae has been deleted from Appendix III to the Convention’. -

A Conservation Reassessment of the Critically Endangered, Lorestan Newt Neurergus Kaiseri (Schmidt 1952) in Iran
Copyright: © This is an open-access article distributed under the terms of the Creative Commons Attribution–Non Commercial –No Derivs 3.0 Unported License, which permits conditional use for non-commercial and education purposes, provided the Amphibian and Reptile Conservation 9(1): 16- original author and source are credited, and prohibits the deposition of material from www.redlist -arc.org or affiliated websites 25. including www.redlist.arcme.org, including PDFs, images, or text, onto other websites without permission of the Amphibian and Reptile Conservation journal at www.redlist-arc.org A conservation reassessment of the Critically Endangered, Lorestan newt Neurergus kaiseri (Schmidt 1952) in Iran 1* 2 3 1 2 Asghar Mobaraki , Mohsen Amiri , Rahim Alvandi , Masoud Ebrahim Tehrani , Hossein Zarin Kia , Ali 2 4, 5 6 Khoshnamvand , Ali Bali Ehsan Forozanfar , and Robert K Browne 1: Department of Environment, Biodiversity and Wildlife Bureau, PO. Box 14155-7383, Tehran, Iran, 2: Department of Environment Provincial Office in Lorestan, Khoram Abad, Iran, 3: Department of Environment Provincial Office in Khuzestan, Ahvaz, Iran 4: Department of Environment, Habitats and Protected Areas Affairs Bureau, PO. Box 14155-7383, Tehran, Iran, 5: Faculty of Environment, Standard Square, Karaj, 6: Sustainability America, Sarteneja, Belize. Abstract The Lorestan newt (Neurergus kaiseri, Schmidt 1952) is an endemic salamander species to Iran, listed as “Critically Endangered” in the 2006 IUCN Red List due to population declines of 80%, over collection for the pet trade; area of occupancy less than 10 km2, fragmented populations, less than 1,000 adults, and continuing habitat degradation and loss. However, the Red List assessment was limited to surveys only around the type location of N. -

No Evidence for the 'Rate-Of-Living' Theory Across the Tetrapod Tree of Life
Received: 23 June 2019 | Revised: 30 December 2019 | Accepted: 7 January 2020 DOI: 10.1111/geb.13069 RESEARCH PAPER No evidence for the ‘rate-of-living’ theory across the tetrapod tree of life Gavin Stark1 | Daniel Pincheira-Donoso2 | Shai Meiri1,3 1School of Zoology, Faculty of Life Sciences, Tel Aviv University, Tel Aviv, Israel Abstract 2School of Biological Sciences, Queen’s Aim: The ‘rate-of-living’ theory predicts that life expectancy is a negative function of University Belfast, Belfast, United Kingdom the rates at which organisms metabolize. According to this theory, factors that accel- 3The Steinhardt Museum of Natural History, Tel Aviv University, Tel Aviv, Israel erate metabolic rates, such as high body temperature and active foraging, lead to organismic ‘wear-out’. This process reduces life span through an accumulation of bio- Correspondence Gavin Stark, School of Zoology, Faculty of chemical errors and the build-up of toxic metabolic by-products. Although the rate- Life Sciences, Tel Aviv University, Tel Aviv, of-living theory is a keystone underlying our understanding of life-history trade-offs, 6997801, Israel. Email: [email protected] its validity has been recently questioned. The rate-of-living theory has never been tested on a global scale in a phylogenetic framework, or across both endotherms and Editor: Richard Field ectotherms. Here, we test several of its fundamental predictions across the tetrapod tree of life. Location: Global. Time period: Present. Major taxa studied: Land vertebrates. Methods: Using a dataset spanning the life span data of 4,100 land vertebrate spe- cies (2,214 endotherms, 1,886 ectotherms), we performed the most comprehensive test to date of the fundamental predictions underlying the rate-of-living theory. -

Notes on the Distribution and Abundance of the Endangered Kaiser ’S Mountain Newt , Neurergus Kaiseri (C Audata : Salamandridae ), in Southwestern Iran
Herpetological Conservation and Biology 8(3):724−731. HSuebrpmeittotelodg: i1c aJlu Clyo 2n0se1r2v;a Aticocne apntedd B: 9io Jlouglyy 2013; Published: 31 December 2013. Notes oN the DistributioN aND abuNDaNce of the eNDaNgereD Kaiser ’s MouNtaiN Newt , Neurergus kaiseri (c auData : salaMaNDriDae ), iN southwesterN iraN Mozafar sharifi 1* , h osseiN farasat 1, h osseiN BaraNi -B eiraNv 2, soMaye vaissi 1, and ehsaN foroozaNfar 3 1Razi University Center for Environmental Studies, Department of Biology, Faculty of Sciences, Kermanshah 67149, Iran 2Department of Biology, Faculty of Sciences, Ferdowsi University of Mashhad, Mashhad, Iran 3Faculty of Environment, Ostandari Square, Karaj, Iran *Corresponding author, e-mail: [email protected] abstract. —the endangered Kaiser’s Mountain Newt, Neurergus kaiseri , is a species endemic to the southern Zagros range in iran. until now, N. kaiseri had been reported from only five localities. the present study describes eight new localities that increase the species’ known range from 212 km 2 (minimum convex polygon) to 789 km 2 at elevations between 930−1395 meters. the localities are generally dispersed; nearest neighbor distances among the 13 localities average 4.61 km (range, 0.93−12.39 km). the streams are separated from one another by steep and rocky terrain with vegetation cover of mature open oak woodland in the west and sparse scrubland or thin oak-pistachio woodland in the central area and east, thus potentially isolating many of the populations from each other. we surveyed 12 of the 13 localities with newts 1−2 times and counted a total of 1,277 adults, post-metamorphic sub-adults, and larvae (mean per stream, 106; range, 2−650) in a total of 4.23 km of stream reaches. -

First Survey of the Pathogenic Fungus Batrachochytrium Salamandrivorans in Wild and Captive Amphibians in the Czech Republic
SALAMANDRA 54(1) 87–91 15 February 2018 CorrespondenceISSN 0036–3375 Correspondence First survey of the pathogenic fungus Batrachochytrium salamandrivorans in wild and captive amphibians in the Czech Republic Vojtech Baláž1,2, Milič Solský3, David Lastra González3, Barbora Havlíková3, Juan Gallego Zamorano3, Cristina González Sevilleja3, Laura Torrent3 & Jiří Vojar3 1) Department of Ecology and Diseases of Game, Fish and Bees, Faculty of Veterinary Hygiene and Ecology, University of Veterinary and Pharmaceutical Sciences Brno, Palackého tř. 1/3, 612 42 Brno, Czech Republic 2) Department of Biology and Wildlife Diseases, Faculty of Veterinary Hygiene and Ecology, University of Veterinary and Pharmaceutical Sciences Brno, Palackého tř. 1/3, 612 42 Brno, Czech Republic 3) Department of Ecology, Faculty of Environmental Sciences, Czech University of Life Sciences Prague, Kamýcká 129, 165 21 Prague 6, Czech Republic Corresponding author: Jiří Vojar, e-mail: [email protected] Manuscript received: 8 June 2017 Accepted: 27 July 2017 by Jörn Köhler The recently discovered fungal pathogen Batrachochytri- exotic pet fairs take place on a regular basis (Havlíková um salamandrivorans (Martel et al. 2013) (hereinafter re- et al. 2015). Furthermore, Prague and its surroundings are ferred to as Bsal) has already received significant scientific known to harbour wild populations of at least four native and public attention (e.g., Martel et al. 2014, Van Rooij caudate species: fire salamander, smooth newt (Lissotriton et al. 2015, Yap et al. 2015, Stegen et al. 2017). The Bsal vulgaris), alpine newt (Ichthyosaura alpestris) and great epidemic has so far been limited to European newts and crested newt (Triturus cristatus) (Šťastný et al. -

Ecologia Balkanica
ECOLOGIA BALKANICA International Scientific Research Journal of Ecology Volume 4, Issue 1 June 2012 UNION OF SCIENTISTS IN BULGARIA – PLOVDIV UNIVERSITY OF PLOVDIV PUBLISHING HOUSE ii International Standard Serial Number Print ISSN 1314-0213; Online ISSN 1313-9940 Aim & Scope „Ecologia Balkanica” is an international scientific journal, in which original research articles in various fields of Ecology are published, including ecology and conservation of microorganisms, plants, aquatic and terrestrial animals, physiological ecology, behavioural ecology, population ecology, population genetics, community ecology, plant-animal interactions, ecosystem ecology, parasitology, animal evolution, ecological monitoring and bioindication, landscape and urban ecology, conservation ecology, as well as new methodical contributions in ecology. Studies conducted on the Balkans are a priority, but studies conducted in Europe or anywhere else in the World is accepted as well. Published by the Union of Scientists in Bulgaria – Plovdiv and the University of Plovdiv Publishing house – twice a year. Language: English. Peer review process All articles included in “Ecologia Balkanica” are peer reviewed. Submitted manuscripts are sent to two or three independent peer reviewers, unless they are either out of scope or below threshold for the journal. These manuscripts will generally be reviewed by experts with the aim of reaching a first decision as soon as possible. The journal uses the double anonymity standard for the peer-review process. Reviewers do not have to sign their reports and they do not know who the author(s) of the submitted manuscript are. We ask all authors to provide the contact details (including e-mail addresses) of at least four potential reviewers of their manuscript. -

Feeding in Amphibians: Evolutionary Transformations and Phenotypic Diversity As Drivers of Feeding System Diversity
Chapter 12 Feeding in Amphibians: Evolutionary Transformations and Phenotypic Diversity as Drivers of Feeding System Diversity Anthony Herrel, James C. O’Reilly, Anne-Claire Fabre, Carla Bardua, Aurélien Lowie, Renaud Boistel and Stanislav N. Gorb Abstract Amphibians are different from most other tetrapods because they have a biphasic life cycle, with larval forms showing a dramatically different cranial anatomy and feeding strategy compared to adults. Amphibians with their exceptional diversity in habitats, lifestyles and reproductive modes are also excellent models for studying the evolutionary divergence in feeding systems. In the present chapter, we review the literature on amphibian feeding anatomy and function published since 2000. We also present some novel unpublished data on caecilian feeding biome- chanics. This review shows that over the past two decades important new insights in our understanding of amphibian feeding anatomy and function have been made possible, thanks to a better understanding of the phylogenetic relationships between taxa, analyses of development and the use of biomechanical modelling. In terms of functional analyses, important advances involve the temperature-dependent nature of tongue projection mechanisms and the plasticity exhibited by animals when switch- A. Herrel (B) Département Adaptations du Vivant, Muséum national d’Histoire naturelle, UMR 7179 C.N.R.S/M.N.H.N, 55 rue Buffon, 75005, Paris Cedex 05, France e-mail: [email protected] J. C. O’Reilly Department of Biomedical Sciences, Ohio University, Cleveland Campus, Cleveland, Ohio 334C, USA A.-C. Fabre · C. Bardua Life Sciences Department, The Natural History Museum, Cromwell Road, London SW7 5BD, UK A. Lowie Department of Biology Evolutionary, Morphology of Vertebrates, Ghent University, K.L.