Identification and Validation of Novel Candidate Risk Genes in Endocytic Vesicular Trafficking Associated with Esophageal Atresia and Tracheoesophageal Fistulas
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Discovery of Driver Non-Coding Splice-Site-Creating Mutations in Cancer
ARTICLE https://doi.org/10.1038/s41467-020-19307-6 OPEN Discovery of driver non-coding splice-site-creating mutations in cancer Song Cao1,2, Daniel Cui Zhou 1,2, Clara Oh1,2, Reyka G. Jayasinghe1,2, Yanyan Zhao1, Christopher J. Yoon1,2, Matthew A. Wyczalkowski 1,2, Matthew H. Bailey 1,2, Terrence Tsou 1,2, Qingsong Gao1,2, ✉ ✉ Andrew Malone 1, Sheila Reynolds3, Ilya Shmulevich3, Michael C. Wendl2,4,5, Feng Chen1,6 & Li Ding1,2,4,6 Non-coding mutations can create splice sites, however the true extent of how such somatic 1234567890():,; non-coding mutations affect RNA splicing are largely unexplored. Here we use the MiSplice pipeline to analyze 783 cancer cases with WGS data and 9494 cases with WES data, discovering 562 non-coding mutations that lead to splicing alterations. Notably, most of these mutations create new exons. Introns associated with new exon creation are significantly larger than the genome-wide average intron size. We find that some mutation-induced splicing alterations are located in genes important in tumorigenesis (ATRX, BCOR, CDKN2B, MAP3K1, MAP3K4, MDM2, SMAD4, STK11, TP53 etc.), often leading to truncated proteins and affecting gene expression. The pattern emerging from these exon-creating mutations sug- gests that splice sites created by non-coding mutations interact with pre-existing potential splice sites that originally lacked a suitable splicing pair to induce new exon formation. Our study suggests the importance of investigating biological and clinical consequences of noncoding splice-inducing mutations that were previously neglected by conventional anno- tation pipelines. MiSplice will be useful for automatically annotating the splicing impact of coding and non-coding mutations in future large-scale analyses. -

Identification of Six Autophagy-Related-Lncrna Prognostic Biomarkers in Uveal Melanoma
Hindawi Disease Markers Volume 2021, Article ID 2401617, 12 pages https://doi.org/10.1155/2021/2401617 Research Article Identification of Six Autophagy-Related-lncRNA Prognostic Biomarkers in Uveal Melanoma Yao Chen , Lu Chen , Jinwei Wang , Jia Tan, and Sha Wang Hunan Key Laboratory of Ophthalmology, Eye Center of Xiangya Hospital, Central South University, China Correspondence should be addressed to Sha Wang; [email protected] Received 7 June 2021; Revised 16 July 2021; Accepted 23 July 2021; Published 13 August 2021 Academic Editor: Ting Su Copyright © 2021 Yao Chen et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Currently, no autophagy-related long noncoding RNA (lncRNA) has been reported to predict the prognosis of uveal melanoma patients. Our study screened for autophagy-related lncRNAs in 80 samples downloaded from The Cancer Genome Atlas (TCGA) database through lncRNA-mRNA coexpression. We used univariate Cox to further filter the lncRNAs. Multivariate Cox regression and LASSO regression were applied to construct an autophagy-associated lncRNA predictive model and calculate the risk score. Clinical risk factors were validated using Cox regression to determine whether they were independent prognostic indicators. Functional enrichment was performed using Gene Ontology and Kyoto Encyclopedia of Genes and Genomes. The model was built with six predictive autophagy-associated lncRNAs and clustered uveal melanoma patients into high- and low-risk groups. The risk score of our model was a significant independent prognostic factor (hazard ratio = 1:0; p <0:001). -

Dual Function of C/D Box Small Nucleolar Rnas in Rrna PNAS PLUS Modification and Alternative Pre-Mrna Splicing
Dual function of C/D box small nucleolar RNAs in rRNA PNAS PLUS modification and alternative pre-mRNA splicing Marina Falaleevaa, Amadis Pagesb, Zaneta Matuszeka, Sana Hidmic, Lily Agranat-Tamirc, Konstantin Korotkova, Yuval Nevod, Eduardo Eyrasb,e, Ruth Sperlingc, and Stefan Stamma,1 aDepartment of Molecular and Cellular Biochemistry, University of Kentucky, Lexington, KY 40536; bDepartment of Experimental and Health Sciences, Universitat Pompeu Fabra, E08003 Barcelona, Spain; cDepartment of Genetics, Hebrew University of Jerusalem, 91904 Jerusalem, Israel; dDepartment of Microbiology and Molecular Genetics, Computation Center at the Hebrew University and Hadassah Medical Center, 91904 Jerusalem, Israel; and eCatalan Institution for Research and Advanced Studies, E08010 Barcelona, Spain Edited by James E. Dahlberg, University of Wisconsin Medical School, Madison, WI, and approved February 5, 2016 (received for review October 1, 2015) C/D box small nucleolar RNAs (SNORDs) are small noncoding RNAs, SNORD fragments are not microRNAs, which have an average and their best-understood function is to target the methyltrans- length of 21–22 nt (13–16), and suggests that these fragments may ferase fibrillarin to rRNA (for example, SNORD27 performs 2′-O- have additional functions. methylation of A27 in 18S rRNA). Unexpectedly, we found a subset The association of some SNORDs with specific diseases sug- of SNORDs, including SNORD27, in soluble nuclear extract made gests that they may possess functions in addition to directing the under native conditions, where fibrillarin was not detected, indi- 2′-O-methylation of rRNA. For example, the loss of SNORD116 cating that a fraction of the SNORD27 RNA likely forms a protein expression is a decisive factor in Prader–Willi syndrome, the most complex different from canonical snoRNAs found in the insoluble common genetic cause for hyperphagia and obesity (17, 18). -

The Clinical Utility of Optical Genome Mapping for the Assessment of Genomic Aberrations in Acute Lymphoblastic Leukemia
cancers Article The Clinical Utility of Optical Genome Mapping for the Assessment of Genomic Aberrations in Acute Lymphoblastic Leukemia Jonathan Lukas Lühmann 1,† , Marie Stelter 1,†, Marie Wolter 1, Josephine Kater 1, Jana Lentes 1, Anke Katharina Bergmann 1, Maximilian Schieck 1 , Gudrun Göhring 1, Anja Möricke 2, Gunnar Cario 2, Markéta Žaliová 3 , Martin Schrappe 2, Brigitte Schlegelberger 1, Martin Stanulla 4 and Doris Steinemann 1,* 1 Department of Human Genetics, Hannover Medical School, 30625 Hannover, Germany; [email protected] (J.L.L.); [email protected] (M.S.); [email protected] (M.W.); [email protected] (J.K.); [email protected] (J.L.); [email protected] (A.K.B.); [email protected] (M.S.); [email protected] (G.G.); [email protected] (B.S.) 2 Department of Pediatrics I, ALL-BFM Study Group, Christian-Albrechts University Kiel and University Medical Center Schleswig-Holstein, 24105 Kiel, Germany; [email protected] (A.M.); [email protected] (G.C.); [email protected] (M.S.) 3 Department of Paediatric Haematology and Oncology, 2nd Faculty of Medicine, Charles University and University Hospital Motol, CZ-15006 Prague, Czech Republic; [email protected] 4 Pediatric Hematology and Oncology, Hannover Medical School, 30625 Hannover, Germany; [email protected] * Correspondence: [email protected] † These authors equally contributed to this work. Citation: Lühmann, J.L.; Stelter, M.; Wolter, M.; Kater, J.; Lentes, J.; Bergmann, A.K.; Schieck, M.; Simple Summary: The stratification of childhood ALL is currently based on various diagnostic Göhring, G.; Möricke, A.; Cario, G.; assays. -

Mouse Map4k3 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Map4k3 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Map4k3 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Map4k3 gene (NCBI Reference Sequence: NM_001290345 ; Ensembl: ENSMUSG00000024242 ) is located on Mouse chromosome 17. 34 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 34 (Transcript: ENSMUST00000025089). Exon 4~6 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Map4k3 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-187J7 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a gene trap allele exhibit decreased susceptibility to experimental autoimmune encephalomyelitis, decreased stimulated immunoglobin production, decreased stimulated T cell proliferation, and abnormal Th1, Th2, and Th17 differentiation. Exon 4 starts from about 9.17% of the coding region. The knockout of Exon 4~6 will result in frameshift of the gene. The size of intron 3 for 5'-loxP site insertion: 8070 bp, and the size of intron 6 for 3'-loxP site insertion: 2734 bp. The size of effective cKO region: ~2560 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 4 5 6 34 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Map4k3 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Functional Splicing Network Reveals Extensive Regulatory Potential of the Core Spliceosomal Machinery
Functional Splicing Network Reveals Extensive Regulatory Potential of the Core Spliceosomal Machinery Panagiotis Papasaikas,1,2,4 J. Ramo´ n Tejedor,1,2,4 Luisa Vigevani,1,2 and Juan Valca´ rcel1,2,3,* 1Centre de Regulacio´ Geno` mica, Dr. Aiguader 88, 08003 Barcelona, Spain 2Universitat Pompeu Fabra, Dr. Aiguader 88, 08003 Barcelona, Spain 3Institucio´ Catalana de Recerca i Estudis Avanc¸ ats, Passeig Lluis Companys 23, 08010 Barcelona, Spain 4Co-first authors SUMMARY quent binding of preassembled U4/5/6 tri-snRNP forms complex B, which after a series of conformational changes forms com- Pre-mRNA splicing relies on the poorly plexes Bact and C, concomitant with the activation of the two understood dynamic interplay between >150 catalytic steps that generate splicing intermediates and prod- protein compo- nents of the spliceosome. The ucts. Transition between spliceosomal subcomplexes involves steps at which splicing can be regulated remain profound dynamic changes in protein composition as well as largely unknown. We sys- tematically analyzed extensive rearrangements of base-pairing interactions between the effect of knocking down the components of snRNAs and between snRNAs and splice site sequences (Wahl the splicing machinery on alterna- tive splicing et al., 2009). RNA structures contributed by base-pairing interac- events relevant for cell proliferation and apoptosis tions between U2 and U6 snRNAs serve to coordinate metal ions and used this information to reconstruct a critical for splicing catalysis (Fica et al., 2013), implying that the network of functional interactions. The network spliceosome is an RNA enzyme whose catalytic center is only accurately captures known physical and established upon assembly of its individual components. -

Promoterless Transposon Mutagenesis Drives Solid Cancers Via Tumor Suppressor Inactivation
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.17.254565; this version posted August 17, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Promoterless Transposon Mutagenesis Drives Solid Cancers via Tumor Suppressor Inactivation 2 Aziz Aiderus1, Ana M. Contreras-Sandoval1, Amanda L. Meshey1, Justin Y. Newberg1,2, Jerrold M. Ward3, 3 Deborah Swing4, Neal G. Copeland2,3,4, Nancy A. Jenkins2,3,4, Karen M. Mann1,2,3,4,5,6,7, and Michael B. 4 Mann1,2,3,4,6,7,8,9 5 1Department of Molecular Oncology, Moffitt Cancer Center & Research Institute, Tampa, FL, USA 6 2Cancer Research Program, Houston Methodist Research Institute, Houston, Texas, USA 7 3Institute of Molecular and Cell Biology, Agency for Science, Technology and Research (A*STAR), 8 Singapore, Republic of Singapore 9 4Mouse Cancer Genetics Program, Center for Cancer Research, National Cancer Institute, Frederick, 10 Maryland, USA 11 5Departments of Gastrointestinal Oncology & Malignant Hematology, Moffitt Cancer Center & Research 12 Institute, Tampa, FL, USA 13 6Cancer Biology and Evolution Program, Moffitt Cancer Center & Research Institute, Tampa, FL, USA 14 7Department of Oncologic Sciences, Morsani College of Medicine, University of South Florida, Tampa, FL, 15 USA. 16 8Donald A. Adam Melanoma and Skin Cancer Research Center of Excellence, Moffitt Cancer Center, Tampa, 17 FL, USA 18 9Department of Cutaneous Oncology, Moffitt Cancer Center & Research Institute, Tampa, FL, USA 19 These authors contributed equally: Aziz Aiderus, Ana M. -
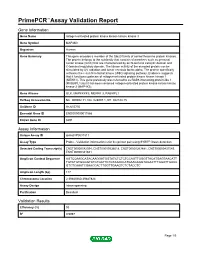
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name mitogen-activated protein kinase kinase kinase kinase 3 Gene Symbol MAP4K3 Organism Human Gene Summary This gene encodes a member of the Ste20 family of serine/threonine protein kinases. The protein belongs to the subfamily that consists of members such as germinal center kinase (GCK) that are characterized by an N-terminal catalytic domain and C-terminal regulatory domain. The kinase activity of the encoded protein can be stimulated by UV radiation and tumor necrosis factor-alpha. The protein specifically activates the c-Jun N-terminal kinase (JNK) signaling pathway. Evidence suggests that it functions upstream of mitogen-activated protein kinase kinase kinase 1 (MEKK1). This gene previously was referred to as RAB8-interacting protein-like 1 (RAB8IPL1) but it has been renamed mitogen-activated protein kinase kinase kinase kinase 3 (MAP4K3). Gene Aliases GLK, MAPKKKK3, MEKKK 3, RAB8IPL1 RefSeq Accession No. NC_000002.11, NG_028007.1, NT_022184.15 UniGene ID Hs.655750 Ensembl Gene ID ENSG00000011566 Entrez Gene ID 8491 Assay Information Unique Assay ID qHsaCIP0031811 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000542094, ENST00000536018, ENST00000263881, ENST00000437545, ENST00000341681 Amplicon Context Sequence AGTCCAAGCATACAAGGATGGTATCTCTCTCCAGTTGGGTTACATGAGTAACATT TGTCTGTGGGGTATCTGATTCTGTAAACCATGAAGAGGTAGAATTTGGATTGACC GTCTCAAATCGAACCACTTGGTTGAAGTCTCTACCTC Amplicon Length (bp) 117 Chromosome -

A Network Inference Approach to Understanding Musculoskeletal
A NETWORK INFERENCE APPROACH TO UNDERSTANDING MUSCULOSKELETAL DISORDERS by NIL TURAN A thesis submitted to The University of Birmingham for the degree of Doctor of Philosophy College of Life and Environmental Sciences School of Biosciences The University of Birmingham June 2013 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Musculoskeletal disorders are among the most important health problem affecting the quality of life and contributing to a high burden on healthcare systems worldwide. Understanding the molecular mechanisms underlying these disorders is crucial for the development of efficient treatments. In this thesis, musculoskeletal disorders including muscle wasting, bone loss and cartilage deformation have been studied using systems biology approaches. Muscle wasting occurring as a systemic effect in COPD patients has been investigated with an integrative network inference approach. This work has lead to a model describing the relationship between muscle molecular and physiological response to training and systemic inflammatory mediators. This model has shown for the first time that oxygen dependent changes in the expression of epigenetic modifiers and not chronic inflammation may be causally linked to muscle dysfunction. -

Mouse Map4k3 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Map4k3 Knockout Project (CRISPR/Cas9) Objective: To create a Map4k3 knockout Mouse model (C57BL/6N) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Map4k3 gene (NCBI Reference Sequence: NM_001290345 ; Ensembl: ENSMUSG00000024242 ) is located on Mouse chromosome 17. 34 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 34 (Transcript: ENSMUST00000025089). Exon 3~5 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a gene trap allele exhibit decreased susceptibility to experimental autoimmune encephalomyelitis, decreased stimulated immunoglobin production, decreased stimulated T cell proliferation, and abnormal Th1, Th2, and Th17 differentiation. Exon 3 starts from about 5.78% of the coding region. Exon 3~5 covers 7.9% of the coding region. The size of effective KO region: ~9671 bp. The KO region does not have any other known gene. Page 1 of 9 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 3 4 5 34 Legends Exon of mouse Map4k3 Knockout region Page 2 of 9 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 3 is aligned with itself to determine if there are tandem repeats. No significant tandem repeat is found in the dot plot matrix. So this region is suitable for PCR screening or sequencing analysis. -
Sheet1 Page 1 Gene Symbol Gene Description Entrez Gene ID
Sheet1 RefSeq ID ProbeSets Gene Symbol Gene Description Entrez Gene ID Sequence annotation Seed matches location(s) Ago-2 binding specific enrichment (replicate 1) Ago-2 binding specific enrichment (replicate 2) OE lysate log2 fold change (replicate 1) OE lysate log2 fold change (replicate 2) Probability Pulled down in Karginov? NM_005646 202813_at TARBP1 Homo sapiens TAR (HIV-1) RNA binding protein 1 (TARBP1), mRNA. 6894 TR(1..5130)CDS(1..4866) 4868..4874,5006..5013 3.73 2.53 -1.54 -0.44 1 Yes NM_001665 203175_at RHOG Homo sapiens ras homolog gene family, member G (rho G) (RHOG), mRNA. 391 TR(1..1332)CDS(159..734) 810..817,782..788,790..796,873..879 3.56 2.78 -1.62 -1 1 Yes NM_002742 205880_at PRKD1 Homo sapiens protein kinase D1 (PRKD1), mRNA. 5587 TR(1..3679)CDS(182..2920) 3538..3544,3202..3208 4.15 1.83 -2.55 -0.42 1 Yes NM_003068 213139_at SNAI2 Homo sapiens snail homolog 2 (Drosophila) (SNAI2), mRNA. 6591 TR(1..2101)CDS(165..971) 1410..1417,1814..1820,1610..1616 3.5 2.79 -1.38 -0.31 1 Yes NM_006270 212647_at RRAS Homo sapiens related RAS viral (r-ras) oncogene homolog (RRAS), mRNA. 6237 TR(1..1013)CDS(46..702) 871..877 3.82 2.27 -1.54 -0.55 1 Yes NM_025188 219923_at,242056_at TRIM45 Homo sapiens tripartite motif-containing 45 (TRIM45), mRNA. 80263 TR(1..3584)CDS(589..2331) 3408..3414,2437..2444,3425..3431,2781..2787 3.87 1.89 -0.62 -0.09 1 Yes NM_024684 221600_s_at,221599_at C11orf67 Homo sapiens chromosome 11 open reading frame 67 (C11orf67), mRNA. -
MAP4K3 (D1L4G) Rabbit Mab -20ºC Store At
MAP4K3 (D1L4G) Rabbit mAb -20ºC Store at Support: +1-978-867-2388 (U.S.) www.cellsignal.com/support Orders: 877-616-2355 (U.S.) [email protected] Entrez-Gene ID #8491 #92427 New 11/16 UniProt ID #Q8IVH8 For Research Use Only. Not For Use In Diagnostic Procedures. Applications Species Cross-Reactivity* Molecular Wt. Isotype Storage: Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 g/ml BSA, 50% glycerol and less than 0.02% W, IP H, Mk 101 kDa Rabbit IgG** µ Endogenous sodium azide. Store at –20°C. Do not aliquot the antibody. *Species cross-reactivity is determined by western blot. ** Anti-rabbit secondary antibodies must be used to Background: The mTORC1 kinase complex is a critical Specificity/Sensitivity: MAP4K3 (D1L4G) Rabbit mAb recog- detect this antibody. regulator of cell growth (1,2). Its activity is modulated by nizes endogenous levels of total MAP4K3 protein. growth factors and nutrients including amino acids (1,2). Recommended Antibody Dilutions: Source/Purification: Monoclonal antibody is produced by MAP4K3 is a mediator between nutrient signal and mTORC1 Western blotting 1:1000 immunizing animals with a synthetic peptide corresponding to (1). Research studies suggest that amino acid sufficiency leads Immunoprecipitation 1:50 residues surrounding Glu452 of human MAP4K3 protein. to the phosphorylation of Ser170 on MAP4K3, which activates mTORC1 (3). For product specific protocols and a complete listing of recommended companion products please see the Background References: product web page at www.cellsignal.com (1) Findlay, G.M. et al. (2007) Biochem J 403, 13-20. (2) Sancak, Y.