Elevated MCOLN1 Expression in P53-Deficient Bladder Cancer Is Necessary for Oncogene-Induced Cell Proliferation, Inflammation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Intracellular Ca2+ Release Channel TRPML1 Regulates Lower Urinary Tract Smooth Muscle Contractility
The intracellular Ca2+ release channel TRPML1 regulates lower urinary tract smooth muscle contractility Caoimhin S. Griffina, Michael G. Alvaradoa, Evan Yamasakia, Bernard T. Drummb,c, Vivek Krishnana, Sher Alia, Eleanor M. Naglea, Kenton M. Sandersb, and Scott Earleya,1 aDepartment of Pharmacology, Center for Molecular and Cellular Signaling in the Cardiovascular System, Reno School of Medicine, University of Nevada, Reno, NV 89557-0318; bDepartment of Physiology and Cell Biology, Reno School of Medicine, University of Nevada, Reno, NV 89557-0318; and cDepartment of Life & Health Sciences, Dundalk Institute of Technology, Louth, Ireland A91 K584 Edited by Mark T. Nelson, University of Vermont, Burlington, VT, and approved October 13, 2020 (received for review August 12, 2020) TRPML1 (transient receptor potential mucolipin 1) is a Ca2+-perme- including dense granulomembranous storage bodies in neurons, able, nonselective cation channel that is predominantly localized to elevated plasma gastrin, vacuolization in the gastric mucosa, and the membranes of late endosomes and lysosomes (LELs). Intracellular retinal degeneration (14). Interestingly, however, an anatomical release of Ca2+ through TRPML1 is thought to be pivotal for mainte- examination of these mice reveals dramatically distended bladders nance of intravesicular acidic pH as well as the maturation, fusion, and (14), leading us to question how TRPML1, an intracellular Ca2+- trafficking of LELs. Interestingly, genetic ablation of TRPML1 in mice release channel important in LEL function, affects bladder −/− (Mcoln1 ) induces a hyperdistended/hypertrophic bladder phenotype. physiology. Here, we investigated this phenomenon further by exploring an un- The lower urinary tract (LUT) is composed of the urinary conventional role for TRPML1 channels in the regulation of Ca2+-signal- bladder and urethra—structures that serve the simple, reciprocal ing activity and contractility in bladder and urethral smooth muscle cells functions of storing and voiding urine (15). -

TRP CHANNELS AS THERAPEUTIC TARGETS TRP CHANNELS AS THERAPEUTIC TARGETS from Basic Science to Clinical Use
TRP CHANNELS AS THERAPEUTIC TARGETS TRP CHANNELS AS THERAPEUTIC TARGETS From Basic Science to Clinical Use Edited by ARPAD SZALLASI MD, PHD Department of Pathology, Monmouth Medical Center, Long Branch, NJ, USA AMSTERDAM • BOSTON • HEIDELBERG • LONDON NEW YORK • OXFORD • PARIS • SAN DIEGO SAN FRANCISCO • SINGAPORE • SYDNEY • TOKYO Academic Press is an imprint of Elsevier Academic Press is an imprint of Elsevier 125 London Wall, London, EC2Y 5AS, UK 525 B Street, Suite 1800, San Diego, CA 92101-4495, USA 225 Wyman Street, Waltham, MA 02451, USA The Boulevard, Langford Lane, Kidlington, Oxford OX5 1GB, UK First published 2015 Copyright © 2015 Elsevier Inc. All rights reserved. No part of this publication may be reproduced or transmitted in any form or by any means, electronic or mechanical, including photocopying, recording, or any information storage and retrieval system, without permission in writing from the publisher. Details on how to seek permission, further information about the Publisher’s permissions policies and our arrangement with organizations such as the Copyright Clearance Center and the Copyright Licensing Agency, can be found at our website: www.elsevier.com/permissions This book and the individual contributions contained in it are protected under copyright by the Publisher (other than as may be noted herein). Notices Knowledge and best practice in this field are constantly changing. As new research and experience broaden our understanding, changes in research methods, professional practices, or medical treatment may become necessary. Practitioners and researchers must always rely on their own experience and knowledge in evaluating and using any information, methods, compounds, or experiments described herein. -
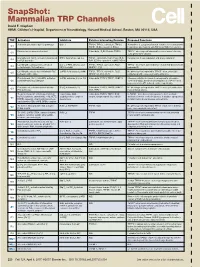
Snapshot: Mammalian TRP Channels David E
SnapShot: Mammalian TRP Channels David E. Clapham HHMI, Children’s Hospital, Department of Neurobiology, Harvard Medical School, Boston, MA 02115, USA TRP Activators Inhibitors Putative Interacting Proteins Proposed Functions Activation potentiated by PLC pathways Gd, La TRPC4, TRPC5, calmodulin, TRPC3, Homodimer is a purported stretch-sensitive ion channel; form C1 TRPP1, IP3Rs, caveolin-1, PMCA heteromeric ion channels with TRPC4 or TRPC5 in neurons -/- Pheromone receptor mechanism? Calmodulin, IP3R3, Enkurin, TRPC6 TRPC2 mice respond abnormally to urine-based olfactory C2 cues; pheromone sensing 2+ Diacylglycerol, [Ca ]I, activation potentiated BTP2, flufenamate, Gd, La TRPC1, calmodulin, PLCβ, PLCγ, IP3R, Potential role in vasoregulation and airway regulation C3 by PLC pathways RyR, SERCA, caveolin-1, αSNAP, NCX1 La (100 µM), calmidazolium, activation [Ca2+] , 2-APB, niflumic acid, TRPC1, TRPC5, calmodulin, PLCβ, TRPC4-/- mice have abnormalities in endothelial-based vessel C4 i potentiated by PLC pathways DIDS, La (mM) NHERF1, IP3R permeability La (100 µM), activation potentiated by PLC 2-APB, flufenamate, La (mM) TRPC1, TRPC4, calmodulin, PLCβ, No phenotype yet reported in TRPC5-/- mice; potentially C5 pathways, nitric oxide NHERF1/2, ZO-1, IP3R regulates growth cones and neurite extension 2+ Diacylglycerol, [Ca ]I, 20-HETE, activation 2-APB, amiloride, Cd, La, Gd Calmodulin, TRPC3, TRPC7, FKBP12 Missense mutation in human focal segmental glomerulo- C6 potentiated by PLC pathways sclerosis (FSGS); abnormal vasoregulation in TRPC6-/- -

Neuropathophysiology, Genetic Profile, and Clinical Manifestation of Mucolipidosis IV—A Review and Case Series
International Journal of Molecular Sciences Review Neuropathophysiology, Genetic Profile, and Clinical Manifestation of Mucolipidosis IV—A Review and Case Series 1, 2, 3, , Aleksandra Jezela-Stanek y , El˙zbietaCiara y and Karolina M. Stepien * y 1 Department of Genetics and Clinical Immunology, National Institute of Tuberculosis and Lung Diseases, 01-138 Warsaw, Poland; [email protected] 2 Department of Medical Genetics, The Children’s Memorial Heath Institute, 04-730 Warsaw, Poland; [email protected] 3 Adult Inherited Metabolic Diseases, Salford Royal NHS Foundation Trust, Salford M6 8HD, UK * Correspondence: [email protected] These authors contributed equally to this work. y Received: 31 May 2020; Accepted: 23 June 2020; Published: 26 June 2020 Abstract: Mucolipidosis type IV (MLIV) is an ultra-rare lysosomal storage disorder caused by biallelic mutations in MCOLN1 gene encoding the transient receptor potential channel mucolipin-1. So far, 35 pathogenic or likely pathogenic MLIV-related variants have been described. Clinical manifestations include severe intellectual disability, speech deficit, progressive visual impairment leading to blindness, and myopathy. The severity of the condition may vary, including less severe psychomotor delay and/or ocular findings. As no striking recognizable facial dysmorphism, skeletal anomalies, organomegaly, or lysosomal enzyme abnormalities in serum are common features of MLIV, the clinical diagnosis may be significantly improved because of characteristic ophthalmological anomalies. This review aims to outline the pathophysiology and genetic defects of this condition with a focus on the genotype–phenotype correlation amongst cases published in the literature. The authors will present their own clinical observations and long-term outcomes in adult MLIV cases. -

Download File
STRUCTURAL AND FUNCTIONAL STUDIES OF TRPML1 AND TRPP2 Nicole Marie Benvin Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2017 © 2017 Nicole Marie Benvin All Rights Reserved ABSTRACT Structural and Functional Studies of TRPML1 and TRPP2 Nicole Marie Benvin In recent years, the determination of several high-resolution structures of transient receptor potential (TRP) channels has led to significant progress within this field. The primary focus of this dissertation is to elucidate the structural characterization of TRPML1 and TRPP2. Mutations in TRPML1 cause mucolipidosis type IV (MLIV), a rare neurodegenerative lysosomal storage disorder. We determined the first high-resolution crystal structures of the human TRPML1 I-II linker domain using X-ray crystallography at pH 4.5, pH 6.0, and pH 7.5. These structures revealed a tetramer with a highly electronegative central pore which plays a role in the dual Ca2+/pH regulation of TRPML1. Notably, these physiologically relevant structures of the I-II linker domain harbor three MLIV-causing mutations. Our findings suggest that these pathogenic mutations destabilize not only the tetrameric structure of the I-II linker, but also the overall architecture of full-length TRPML1. In addition, TRPML1 proteins containing MLIV- causing mutations mislocalized in the cell when imaged by confocal fluorescence microscopy. Mutations in TRPP2 cause autosomal dominant polycystic kidney disease (ADPKD). Since novel technological advances in single-particle cryo-electron microscopy have now enabled the determination of high-resolution membrane protein structures, we set out to solve the structure of TRPP2 using this technique. -

Pflugers Final
CORE Metadata, citation and similar papers at core.ac.uk Provided by Serveur académique lausannois A comprehensive analysis of gene expression profiles in distal parts of the mouse renal tubule. Sylvain Pradervand2, Annie Mercier Zuber1, Gabriel Centeno1, Olivier Bonny1,3,4 and Dmitri Firsov1,4 1 - Department of Pharmacology and Toxicology, University of Lausanne, 1005 Lausanne, Switzerland 2 - DNA Array Facility, University of Lausanne, 1015 Lausanne, Switzerland 3 - Service of Nephrology, Lausanne University Hospital, 1005 Lausanne, Switzerland 4 – these two authors have equally contributed to the study to whom correspondence should be addressed: Dmitri FIRSOV Department of Pharmacology and Toxicology, University of Lausanne, 27 rue du Bugnon, 1005 Lausanne, Switzerland Phone: ++ 41-216925406 Fax: ++ 41-216925355 e-mail: [email protected] and Olivier BONNY Department of Pharmacology and Toxicology, University of Lausanne, 27 rue du Bugnon, 1005 Lausanne, Switzerland Phone: ++ 41-216925417 Fax: ++ 41-216925355 e-mail: [email protected] 1 Abstract The distal parts of the renal tubule play a critical role in maintaining homeostasis of extracellular fluids. In this review, we present an in-depth analysis of microarray-based gene expression profiles available for microdissected mouse distal nephron segments, i.e., the distal convoluted tubule (DCT) and the connecting tubule (CNT), and for the cortical portion of the collecting duct (CCD) (Zuber et al., 2009). Classification of expressed transcripts in 14 major functional gene categories demonstrated that all principal proteins involved in maintaining of salt and water balance are represented by highly abundant transcripts. However, a significant number of transcripts belonging, for instance, to categories of G protein-coupled receptors (GPCR) or serine-threonine kinases exhibit high expression levels but remain unassigned to a specific renal function. -

1 1 2 3 Cell Type-Specific Transcriptomics of Hypothalamic
1 2 3 4 Cell type-specific transcriptomics of hypothalamic energy-sensing neuron responses to 5 weight-loss 6 7 Fredrick E. Henry1,†, Ken Sugino1,†, Adam Tozer2, Tiago Branco2, Scott M. Sternson1,* 8 9 1Janelia Research Campus, Howard Hughes Medical Institute, 19700 Helix Drive, Ashburn, VA 10 20147, USA. 11 2Division of Neurobiology, Medical Research Council Laboratory of Molecular Biology, 12 Cambridge CB2 0QH, UK 13 14 †Co-first author 15 *Correspondence to: [email protected] 16 Phone: 571-209-4103 17 18 Authors have no competing interests 19 1 20 Abstract 21 Molecular and cellular processes in neurons are critical for sensing and responding to energy 22 deficit states, such as during weight-loss. AGRP neurons are a key hypothalamic population 23 that is activated during energy deficit and increases appetite and weight-gain. Cell type-specific 24 transcriptomics can be used to identify pathways that counteract weight-loss, and here we 25 report high-quality gene expression profiles of AGRP neurons from well-fed and food-deprived 26 young adult mice. For comparison, we also analyzed POMC neurons, an intermingled 27 population that suppresses appetite and body weight. We find that AGRP neurons are 28 considerably more sensitive to energy deficit than POMC neurons. Furthermore, we identify cell 29 type-specific pathways involving endoplasmic reticulum-stress, circadian signaling, ion 30 channels, neuropeptides, and receptors. Combined with methods to validate and manipulate 31 these pathways, this resource greatly expands molecular insight into neuronal regulation of 32 body weight, and may be useful for devising therapeutic strategies for obesity and eating 33 disorders. -

Transient Receptor Potential Channels As Drug Targets: from the Science of Basic Research to the Art of Medicine
1521-0081/66/3/676–814$25.00 http://dx.doi.org/10.1124/pr.113.008268 PHARMACOLOGICAL REVIEWS Pharmacol Rev 66:676–814, July 2014 Copyright © 2014 by The American Society for Pharmacology and Experimental Therapeutics ASSOCIATE EDITOR: DAVID R. SIBLEY Transient Receptor Potential Channels as Drug Targets: From the Science of Basic Research to the Art of Medicine Bernd Nilius and Arpad Szallasi KU Leuven, Department of Cellular and Molecular Medicine, Laboratory of Ion Channel Research, Campus Gasthuisberg, Leuven, Belgium (B.N.); and Department of Pathology, Monmouth Medical Center, Long Branch, New Jersey (A.S.) Abstract. ....................................................................................679 I. Transient Receptor Potential Channels: A Brief Introduction . ...............................679 A. Canonical Transient Receptor Potential Subfamily . .....................................682 B. Vanilloid Transient Receptor Potential Subfamily . .....................................686 C. Melastatin Transient Receptor Potential Subfamily . .....................................696 Downloaded from D. Ankyrin Transient Receptor Potential Subfamily .........................................700 E. Mucolipin Transient Receptor Potential Subfamily . .....................................702 F. Polycystic Transient Receptor Potential Subfamily . .....................................703 II. Transient Receptor Potential Channels: Hereditary Diseases (Transient Receptor Potential Channelopathies). ......................................................704 -

Joint Diseases
EuropeanO Krupkova Cells et al.and Materials Vol. 34 2017 (pages 180-201) DOI: 10.22203/eCM.v034a12 TRP channels in ISSN joint 1473-2262 diseases THE ROLE OF TRANSIENT RECEPTOR POTENTIAL CHANNELS IN JOINT DISEASES O. Krupkova1,*, J. Zvick1 and K. Wuertz-Kozak1,2,3,4 1 Department of Health Sciences and Technology, Institute for Biomechanics, ETH Zurich, Zurich, Switzerland 2 Department of Health Sciences, Institute of Sociology of Health and Physical Activity, University of Potsdam, Potsdam, Germany 3 Schön Clinic Munich Harlaching, Munich, Germany 4 Spine Centre, Academic Teaching Hospital and Spine Research, Paracelsus Medical University, Salzburg, Austria Abstract Transient receptor potential channels (TRP channels) are cation selective transmembrane receptors with diverse structures, activation mechanisms and physiological functions. TRP channels act as cellular sensors for a plethora of stimuli, including temperature, membrane voltage, oxidative stress, mechanical stimuli, pH and endogenous as well as exogenous ligands, thereby illustrating their versatility. As such, TRP channels regulate various functions in both excitable and non-excitable cells, mainly by mediating Ca2+ homeostasis. Dysregulation of TRP channels is implicated in many pathologies, including cardiovascular diseases, muscular dystrophies and hyperalgesia. However, the importance of TRP channel expression, physiological function and regulation in chondrocytes and intervertebral disc (IVD) cells is largely unexplored. Osteoarthritis (OA) and degenerative disc disease (DDD) are chronic age-related disorders that significantly affect the quality of life by causing pain, activity limitation and disability. Furthermore, currently available therapies cannot effectively slow-down or stop progression of these diseases. Both OA and DDD are characterised by reduced tissue cellularity, enhanced inflammatory responses and molecular, structural and mechanical alterations of the extracellular matrix, hence affecting load distribution and reducing joint flexibility. -

Igniting Ca2+ Sparks with TRPML1
COMMENTARY COMMENTARY + Igniting Ca2 sparks with TRPML1 Gerard P. Sergeanta,1, Mark A. Hollywooda, and Keith D. Thornburya Storage and voiding of urine in mammals is accom- leading to hyperpolarization of membrane potential + plished by a reciprocal contractile relationship be- and SMC relaxation (8, 10). Ca2 sparks originate from + tween the bladder and the urethra. During the storage ryanodine-sensitive intracellular Ca2 release channels phase, the urethra remains contracted to prevent (known as ryanodine receptors [RyRs]) on the sarco- leakage of urine, while the bladder is relaxed to accom- plasmic reticulum (SR) membrane (11). RyRs are acti- + modate the increased volume of urine. Conversely, vated by micromolar levels of intracellular Ca2 via + + during voiding, the urethra relaxes, while the bladder Ca2 -induced Ca2 release (CICR). In cardiac myo- + contracts to generate an intravesical pressure which cytes, RyR activation is tightly coupled to Ca2 influx exceeds that in the urethra (1). The cellular mecha- via VDCC (11); however, this is not the case in smooth nisms that govern these functions are only partly under- muscle, where there is only “loose coupling” between stood, and there is a particular paucity of data on plasmalemmal VDCCs and RyRs (12). the cellular processes that underlie tonic contraction The mechanisms responsible for the localized ac- + of urethral smooth muscle (USM). This issue is addressed tivation of RyRs that initiate Ca2 sparks in detrusor in PNAS by Griffin et al. (2), who demonstrate a crit- and urethral myocytes have remained elusive. This is- ical role for transient receptor potential mucolipin 1 sue is addressed by Griffin et al. -

Ca2+ Dyshomeostasis Disrupts Neuronal and Synaptic Function in Alzheimer's Disease
cells Review Ca2+ Dyshomeostasis Disrupts Neuronal and Synaptic Function in Alzheimer’s Disease John McDaid 1,2 , Sarah Mustaly-Kalimi 1,2 and Grace E. Stutzmann 1,2,3,* 1 Center for Neurodegenerative Disease and Therapeutics, Rosalind Franklin University of Medicine and Science, 3333 Green Bay Rd., North Chicago, IL 60064, USA; [email protected] (J.M.); [email protected] (S.M.-K.) 2 School of Graduate and Postdoctoral Studies, Rosalind Franklin University of Medicine and Science, 3333 Green Bay Rd., North Chicago, IL 60064, USA 3 Chicago Medical School, Rosalind Franklin University of Medicine and Science, 3333 Green Bay Rd., North Chicago, IL 60064, USA * Correspondence: [email protected]; Tel.: +1-847-578-8540 Received: 3 November 2020; Accepted: 7 December 2020; Published: 10 December 2020 Abstract: Ca2+ homeostasis is essential for multiple neuronal functions and thus, Ca2+ dyshomeostasis can lead to widespread impairment of cellular and synaptic signaling, subsequently contributing to dementia and Alzheimer’s disease (AD). While numerous studies implicate Ca2+ mishandling in AD, the cellular basis for loss of cognitive function remains under investigation. The process of synaptic degradation and degeneration in AD is slow, and constitutes a series of maladaptive processes each contributing to a further destabilization of the Ca2+ homeostatic machinery. Ca2+ homeostasis involves precise maintenance of cytosolic Ca2+ levels, despite extracellular influx via multiple synaptic Ca2+ channels, and intracellular release via organelles such as the endoplasmic reticulum (ER) via ryanodine receptor (RyRs) and IP3R, lysosomes via transient receptor potential mucolipin channel (TRPML) and two pore channel (TPC), and mitochondria via the permeability transition pore (PTP). -

Multiomics of Azacitidine-Treated AML Cells Reveals Variable And
Multiomics of azacitidine-treated AML cells reveals variable and convergent targets that remodel the cell-surface proteome Kevin K. Leunga, Aaron Nguyenb, Tao Shic, Lin Tangc, Xiaochun Nid, Laure Escoubetc, Kyle J. MacBethb, Jorge DiMartinob, and James A. Wellsa,1 aDepartment of Pharmaceutical Chemistry, University of California, San Francisco, CA 94143; bEpigenetics Thematic Center of Excellence, Celgene Corporation, San Francisco, CA 94158; cDepartment of Informatics and Predictive Sciences, Celgene Corporation, San Diego, CA 92121; and dDepartment of Informatics and Predictive Sciences, Celgene Corporation, Cambridge, MA 02140 Contributed by James A. Wells, November 19, 2018 (sent for review August 23, 2018; reviewed by Rebekah Gundry, Neil L. Kelleher, and Bernd Wollscheid) Myelodysplastic syndromes (MDS) and acute myeloid leukemia of DNA methyltransferases, leading to loss of methylation in (AML) are diseases of abnormal hematopoietic differentiation newly synthesized DNA (10, 11). It was recently shown that AZA with aberrant epigenetic alterations. Azacitidine (AZA) is a DNA treatment of cervical (12, 13) and colorectal (14) cancer cells methyltransferase inhibitor widely used to treat MDS and AML, can induce interferon responses through reactivation of endoge- yet the impact of AZA on the cell-surface proteome has not been nous retroviruses. This phenomenon, termed viral mimicry, is defined. To identify potential therapeutic targets for use in com- thought to induce antitumor effects by activating and engaging bination with AZA in AML patients, we investigated the effects the immune system. of AZA treatment on four AML cell lines representing different Although AZA treatment has demonstrated clinical benefit in stages of differentiation. The effect of AZA treatment on these AML patients, additional therapeutic options are needed (8, 9).