Recounting a Genetic Story Roger H
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Evidence for Gliadin Antibodies As Causative Agents in Schizophrenia
1 Evidence for gliadin antibodies as causative agents in schizophrenia. C.J.Carter PolygenicPathways, 20 Upper Maze Hill, Saint-Leonard’s on Sea, East Sussex, TN37 0LG [email protected] Tel: 0044 (0)1424 422201 I have no fax Abstract Antibodies to gliadin, a component of gluten, have frequently been reported in schizophrenia patients, and in some cases remission has been noted following the instigation of a gluten free diet. Gliadin is a highly immunogenic protein, and B cell epitopes along its entire immunogenic length are homologous to the products of numerous proteins relevant to schizophrenia (p = 0.012 to 3e-25). These include members of the DISC1 interactome, of glutamate, dopamine and neuregulin signalling networks, and of pathways involved in plasticity, dendritic growth or myelination. Antibodies to gliadin are likely to cross react with these key proteins, as has already been observed with synapsin 1 and calreticulin. Gliadin may thus be a causative agent in schizophrenia, under certain genetic and immunological conditions, producing its effects via antibody mediated knockdown of multiple proteins relevant to the disease process. Because of such homology, an autoimmune response may be sustained by the human antigens that resemble gliadin itself, a scenario supported by many reports of immune activation both in the brain and in lymphocytes in schizophrenia. Gluten free diets and removal of such antibodies may be of therapeutic benefit in certain cases of schizophrenia. 2 Introduction A number of studies from China, Norway, and the USA have reported the presence of gliadin antibodies in schizophrenia 1-5. Gliadin is a component of gluten, intolerance to which is implicated in coeliac disease 6. -

Ring 21 FTNW
Ring 21 rarechromo.org Sources Ring 21 The information Ring 21 is a rare genetic condition caused by having a in this leaflet ring-shaped chromosome. comes from the Almost half of the people with ring 21 chromosomes medical literature described in the medical literature are healthy and and from develop normally. Their unusual chromosomes are Unique’s discovered by chance, during tests for infertility or after members with repeated miscarriages or after having an affected baby. Ring 21 In other people the ring 21 chromosome affects (referenced U), development and learning and can also cause medical who were problems. In most of these people these effects are surveyed in slight but in some people they can be severe. The 2004. Unique is effects can even vary between different members of the very grateful to same family. The reason for these differences is not yet the families who fully understood. took part in the survey. What is a chromosome? The human body is made up of cells. Inside most cells is References a nucleus where genetic information is stored in genes which are grouped along chromosomes. Chromosomes The text contains are large enough to be studied under a microscope and references to come in different sizes, each with a short (p) and a long articles published (q) arm. They are numbered from largest to smallest in the medical according to their size, from number 1 to number 22, in press. The first- addition to the sex chromosomes, X and Y. A normal, named author healthy cell in the body has 46 chromosomes, 23 from and publication the mother and 23 from the father, including one date are given to chromosome 21 from each parent. -

Epigenetic Control of Mammalian Centromere Protein Binding: Does DNA Methylation Have a Role?
Journal of Cell Science 109, 2199-2206 (1996) 2199 Printed in Great Britain © The Company of Biologists Limited 1996 JCS3386 Epigenetic control of mammalian centromere protein binding: does DNA methylation have a role? Arthur R. Mitchell*, Peter Jeppesen, Linda Nicol†, Harris Morrison and David Kipling MRC Human Genetics Unit, Western General Hospital, Crewe Road, Edinburgh EH4 2XU, UK *Author for correspondence (internet [email protected]) †Present address: MRC Reproductive Biology Unit, Edinburgh, UK SUMMARY Chromosome 1 of the inbred mouse strain DBA/2 has a block of minor satellite DNA sequences on chromosome 1. polymorphism associated with the minor satellite DNA at The binding of the CENP-E protein does not appear to be its centromere. The more terminal block of satellite DNA affected by demethylation of the minor satellite sequences. sequences on this chromosome acts as the centromere as We present a model to explain these observations. This shown by the binding of CREST ACA serum, anti-CENP- model may also indicate the mechanism by which the B and anti-CENP-E polyclonal sera. Demethylation of the CENP-B protein recognises specific sites within the arrays minor satellite DNA sequences accomplished by growing of minor satellite DNA on mouse chromosomes. cells in the presence of the drug 5-aza-2′-deoxycytidine results in a redistribution of the CENP-B protein. This protein now binds to an enlarged area on the more terminal Key words: Centromere satellite DNA, Demethylation, Centromere block and in addition it now binds to the more internal antibody INTRODUCTION A common feature of many mammalian pericentromeric domains is that they contain families of repetitive DNA The centromere of mammalian chromosomes is recognised at sequences (Singer, 1982). -

The 50Th Anniversary of the Discovery of Trisomy 21: the Past, Present, and Future of Research and Treatment of Down Syndrome
REVIEW The 50th anniversary of the discovery of trisomy 21: The past, present, and future of research and treatment of Down syndrome Andre´Me´garbane´, MD, PhD1,2, Aime´ Ravel, MD1, Clotilde Mircher, MD1, Franck Sturtz, MD, PhD1,3, Yann Grattau, MD1, Marie-Odile Rethore´, MD1, Jean-Maurice Delabar, PhD4, and William C. Mobley, MD, PhD5 Abstract: Trisomy 21 or Down syndrome is a chromosomal disorder HISTORICAL REVIEW resulting from the presence of all or part of an extra Chromosome 21. Clinical description It is a common birth defect, the most frequent and most recognizable By examining artifacts from the Tumaco-La Tolita culture, form of mental retardation, appearing in about 1 of every 700 newborns. which existed on the border between current Colombia and Although the syndrome had been described thousands of years before, Ecuador approximately 2500 years ago, Bernal and Briceno2 it was named after John Langdon Down who reported its clinical suspected that certain figurines depicted individuals with Tri- description in 1866. The suspected association of Down syndrome with somy 21, making these potteries the earliest evidence for the a chromosomal abnormality was confirmed by Lejeune et al. in 1959. existence of the syndrome. Martinez-Frias3 identified the syn- Fifty years after the discovery of the origin of Down syndrome, the term drome in a terra-cotta head from the Tolteca culture of Mexico “mongolism” is still inappropriately used; persons with Down syn- in 500 patients with AD in which the facial features of Trisomy drome are still institutionalized. Health problems associated with that 21 are clearly displayed. -

Biol 1020: Chromosomal Genetics
Ch. 15: Chromosomal Abnormalities Abnormalities in Chromosomal Number Abnormalities in Chromosomal Structure: Rearrangements Fragile Sites . • Define: – nondisjunction – polyploidy – aneupoidy – trisomy – monosomy . Abnormalities in chromosomal number How does it happen? . Abnormalities in chromosomal number nondisjunction - mistake in cell division where chromosomes do not separate properly in anaphase usually in meiosis, although in mitosis occasionally in meiosis, can occur in anaphase I or II . Abnormalities in chromosomal number polyploidy – complete extra sets (3n, etc.) – fatal in humans, most animals aneuploidy – missing one copy or have an extra copy of a single chromosome three copies of a chromosome in your somatic cells: trisomy one copy of a chromosome in your somatic cells: monosomy most trisomies and monosomies are lethal well before birth in humans; exceptions will be covered . Abnormalities in chromosomal number generally, in humans autosomal aneuploids tend to be spontaneously aborted over 1/5 of human pregnancies are lost spontaneously after implantation (probably closer to 1/3) chromosomal abnormalities are the leading known cause of pregnancy loss data indicate that minimum 10-15% of conceptions have a chromosomal abnormality at least 95% of these conceptions spontaneously abort (often without being noticed) . • Define: – nondisjunction – polyploidy – aneupoidy – trisomy – monosomy . • Describe each of the aneuploidies that can be found in an appreciable number of human adults (chromosomal abnormality, common name of the syndrome if it has one, phenotypes) . aneuploidy in human sex chromosomes X_ female (Turner syndrome) short stature; sterile (immature sex organs); often reduced mental abilities about 1 in 2500 human female births XXY male (Klinefelter syndrome) often not detected until puberty, when female body characteristics develop sterile; sometimes reduced mental abilities; testosterone shots can be used as a partial treatment; about 1 in 500 human male births . -
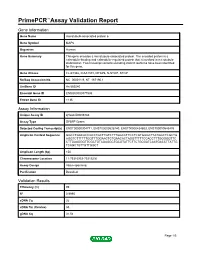
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name microtubule-associated protein 6 Gene Symbol MAP6 Organism Human Gene Summary This gene encodes a microtubule-associated protein. The encoded protein is a calmodulin-binding and calmodulin-regulated protein that is involved in microtubule stabilization. Two transcript variants encoding distinct isoforms have been identified for this gene. Gene Aliases FLJ41346, KIAA1878, MTAP6, N-STOP, STOP RefSeq Accession No. NC_000011.9, NT_167190.1 UniGene ID Hs.585540 Ensembl Gene ID ENSG00000171533 Entrez Gene ID 4135 Assay Information Unique Assay ID qHsaCID0006783 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000304771, ENST00000526740, ENST00000434603, ENST00000545476 Amplicon Context Sequence GGCCTGACACCGCCTGCTTGTCTTTGGCCTTCCTCGTGGGCTTATGGCTCGCTG AGGTCTTTTTTGGTTTGGAACTCTGAACACTAGGTTTTTCCACCTTTGGGGGTTC CTTGAAGGGTTCGCTGTAGAGGCTGCGTATTCTTCTGCGATCAATGACCTTATTG TCAGCTGTTGTTGGCT Amplicon Length (bp) 150 Chromosome Location 11:75316953-75319236 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 99 R2 0.9995 cDNA Cq 26 cDNA Tm (Celsius) 85 gDNA Cq 31.54 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report MAP6, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Dr. Fern Tsien, Dept. of Genetics, LSUHSC, NO, LA Down Syndrome
COMMON TYPES OF CHROMOSOME ABNORMALITIES Dr. Fern Tsien, Dept. of Genetics, LSUHSC, NO, LA A. Trisomy: instead of having the normal two copies of each chromosome, an individual has three of a particular chromosome. Which chromosome is trisomic determines the type and severity of the disorder. Down syndrome or Trisomy 21, is the most common trisomy, occurring in 1 per 800 births (about 3,400) a year in the United States. It is one of the most common genetic birth defects. According to the National Down Syndrome Society, there are more than 400,000 individuals with Down syndrome in the United States. Patients with Down syndrome have three copies of their 21 chromosomes instead of the normal two. The major clinical features of Down syndrome patients include low muscle tone, small stature, an upward slant to the eyes, a single deep crease across the center of the palm, mental retardation, and physical abnormalities, including heart and intestinal defects, and increased risk of leukemia. Every person with Down syndrome is a unique individual and may possess these characteristics to different degrees. Down syndrome patients Karyotype of a male patient with trisomy 21 What are the causes of Down syndrome? • 95% of all Down syndrome patients have a trisomy due to nondisjunction before fertilization • 1-2% have a mosaic karyotype due to nondisjunction after fertilization • 3-4% are due to a translocation 1. Nondisjunction refers to the failure of chromosomes to separate during cell division in the formation of the egg, sperm, or the fetus, causing an abnormal number of chromosomes. As a result, the baby may have an extra chromosome (trisomy). -

Cytogenetics, Chromosomal Genetics
Cytogenetics Chromosomal Genetics Sophie Dahoun Service de Génétique Médicale, HUG Geneva, Switzerland [email protected] Training Course in Sexual and Reproductive Health Research Geneva 2010 Cytogenetics is the branch of genetics that correlates the structure, number, and behaviour of chromosomes with heredity and diseases Conventional cytogenetics Molecular cytogenetics Molecular Biology I. Karyotype Definition Chromosomal Banding Resolution limits Nomenclature The metaphasic chromosome telomeres p arm q arm G-banded Human Karyotype Tjio & Levan 1956 Karyotype: The characterization of the chromosomal complement of an individual's cell, including number, form, and size of the chromosomes. A photomicrograph of chromosomes arranged according to a standard classification. A chromosome banding pattern is comprised of alternating light and dark stripes, or bands, that appear along its length after being stained with a dye. A unique banding pattern is used to identify each chromosome Chromosome banding techniques and staining Giemsa has become the most commonly used stain in cytogenetic analysis. Most G-banding techniques require pretreating the chromosomes with a proteolytic enzyme such as trypsin. G- banding preferentially stains the regions of DNA that are rich in adenine and thymine. R-banding involves pretreating cells with a hot salt solution that denatures DNA that is rich in adenine and thymine. The chromosomes are then stained with Giemsa. C-banding stains areas of heterochromatin, which are tightly packed and contain -

Down's Syndrome Phenotype and Autosomal Gene Inactivation in a Child with Presumed
J Med Genet: first published as 10.1136/jmg.19.2.144 on 1 April 1982. Downloaded from 144 Case reports Down's syndrome phenotype and Case report autosomal gene inactivation in a The proband, a 3 2-year-old white female, was referred for evaluation of developmental delay and child with presumed (X;21) de novo dysmorphic features. She was the 1790 g,. 39 week translocation gestation product of a gravida 1, para 0, 15-year-old female. The pregnancy was complicated with SUMMARY A 32-year-old female with clinical recurrent urinary tract infections. The mother used features of Down's syndrome was found to have alcohol and tobacco in small quantities during the extra chromosome material on the long arm of pregnancy. The labour lasted ten hours and the one of the X chromosomes, 46,XXq+. The delivery was vaginal with vertex presentation. The parental karyotypes were normal. In the light baby breathed and cried spontaneously. Her of the clinical features of the proband and the immediate neonatal course was uneventful, but her subsequent weight gain was poor. She had several banding characteristics of the extra chromosome admissions to hospital for repeated diarrhoea, otitis material, the patient was thought to have a de media, and pneumonia. She had two 'febrile' seizures novo (X;21) translocation. The results of late for which she was placed on phenobarbital. Her replication studies with BUdR and enzyme development was markedly delayed. She smiled at superoxide dismutase (SOD) assays in the 4 months, turned over at 7 months, walked at 18 proband suggest that: (1) the presumed (X;21) months, and was not yet toilet trained. -

Microtubule Organization and Microtubule- Associated Proteins (Maps)
Chapter 3 Microtubule Organization and Microtubule- Associated Proteins (MAPs) Elena Tortosa, Lukas C. Kapitein, and Casper C. Hoogenraad Abstract Dendrites have a unique microtubule organization. In vertebrates, den- dritic microtubules are organized in antiparallel bundles, oriented with their plus ends either pointing away or toward the soma. The mixed microtubule arrays control intracellular trafficking and local signaling pathways, and are essential for dendrite development and function. The organization of microtubule arrays largely depends on the combined function of different microtubule regulatory factors or generally named microtubule-associated proteins (MAPs). Classical MAPs, also called structural MAPs, were identified more than 20 years ago based on their ability to bind to and copurify with microtubules. Most classical MAPs bind along the microtubule lattice and regulate microtubule polymerization, bundling, and stabilization. Recent evidences suggest that classical MAPs also guide motor protein transport, interact with the actin cytoskeleton, and act in various neuronal signaling networks. Here, we give an overview of microtubule organization in dendrites and the role of classical MAPs in dendrite development, dendritic spine formation, and synaptic plasticity. Keywords Neuron • Dendrite • Cytoskeleton • Microtubule • Microtubule- associated protein • MAP1 • MAP2 • MAP4 • MAP6 • MAP7 • MAP9 • Tau 3.1 Introduction Microtubules (MTs) are cytoskeletal structures that play essential roles in all eukaryotic cells. MTs are important not only during cell division but also in non-dividing cells, where they are critical structures in numerous cellular processes such as cell motility, migration, differentiation, intracellular transport and organelle positioning. MTs are composed of two proteins, α- and β-tubulin, that form heterodimers and organize themselves in a head-to-tail manner. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Mutational Mechanisms That Activate Wnt Signaling and Predict Outcomes in Colorectal Cancer Patients William Hankey1, Michael A
Published OnlineFirst December 6, 2017; DOI: 10.1158/0008-5472.CAN-17-1357 Cancer Genome and Epigenome Research Mutational Mechanisms That Activate Wnt Signaling and Predict Outcomes in Colorectal Cancer Patients William Hankey1, Michael A. McIlhatton1, Kenechi Ebede2, Brian Kennedy3, Baris Hancioglu3, Jie Zhang4, Guy N. Brock3, Kun Huang4, and Joanna Groden1 Abstract APC biallelic loss-of-function mutations are the most prevalent also exhibiting unique changes in pathways related to prolifera- genetic changes in colorectal tumors, but it is unknown whether tion, cytoskeletal organization, and apoptosis. Apc-mutant ade- these mutations phenocopy gain-of-function mutations in the nomas were characterized by increased expression of the glial CTNNB1 gene encoding b-catenin that also activate canonical nexin Serpine2, the human ortholog, which was increased in WNT signaling. Here we demonstrate that these two mutational advanced human colorectal tumors. Our results support the mechanisms are not equivalent. Furthermore, we show how hypothesis that APC-mutant colorectal tumors are transcription- differences in gene expression produced by these different ally distinct from APC-wild-type colorectal tumors with canonical mechanisms can stratify outcomes in more advanced human WNT signaling activated by other mechanisms, with possible colorectal cancers. Gene expression profiling in Apc-mutant and implications for stratification and prognosis. Ctnnb1-mutant mouse colon adenomas identified candidate Significance: These findings suggest that colon adenomas genes for subsequent evaluation of human TCGA (The Cancer driven by APC mutations are distinct from those driven by WNT Genome Atlas) data for colorectal cancer outcomes. Transcrip- gain-of-function mutations, with implications for identifying tional patterns exhibited evidence of activated canonical Wnt at-risk patients with advanced disease based on gene expression signaling in both types of adenomas, with Apc-mutant adenomas patterns.