Subgroup-Specific Prognostic Signaling and Metabolic Pathways
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Uncovering the Signaling Landscape Controlling Breast Cancer Cell Migration Identifies Novel Metastasis Driver Genes
ARTICLE https://doi.org/10.1038/s41467-019-11020-3 OPEN Uncovering the signaling landscape controlling breast cancer cell migration identifies novel metastasis driver genes Esmee Koedoot1,4, Michiel Fokkelman 1,4, Vasiliki-Maria Rogkoti1,4, Marcel Smid2, Iris van de Sandt1, Hans de Bont1, Chantal Pont1, Janna E. Klip1, Steven Wink 1, Mieke A. Timmermans2, Erik A.C. Wiemer2, Peter Stoilov3, John A. Foekens2, Sylvia E. Le Dévédec 1, John W.M. Martens 2 & Bob van de Water1 1234567890():,; Ttriple-negative breast cancer (TNBC) is an aggressive and highly metastatic breast cancer subtype. Enhanced TNBC cell motility is a prerequisite of TNBC cell dissemination. Here, we apply an imaging-based RNAi phenotypic cell migration screen using two highly motile TNBC cell lines (Hs578T and MDA-MB-231) to provide a repository of signaling determinants that functionally drive TNBC cell motility. We have screened ~4,200 target genes individually and discovered 133 and 113 migratory modulators of Hs578T and MDA-MB-231, respectively, which are linked to signaling networks predictive for breast cancer progression. The splicing factors PRPF4B and BUD31 and the transcription factor BPTF are essential for cancer cell migration, amplified in human primary breast tumors and associated with metastasis-free survival. Depletion of PRPF4B, BUD31 and BPTF causes primarily down regulation of genes involved in focal adhesion and ECM-interaction pathways. PRPF4B is essential for TNBC metastasis formation in vivo, making PRPF4B a candidate for further drug development. 1 Division of Drug Discovery and Safety, LACDR, Leiden University, Einsteinweg 55, Leiden 2333 CC, Netherlands. 2 Department of Medical Oncology and Cancer Genomics Netherlands, Erasmus MC Cancer Institute, Erasmus University Medical Center, Rotterdam 3008 AE, Netherlands. -

Nuclear and Mitochondrial Genome Defects in Autisms
UC Irvine UC Irvine Previously Published Works Title Nuclear and mitochondrial genome defects in autisms. Permalink https://escholarship.org/uc/item/8vq3278q Journal Annals of the New York Academy of Sciences, 1151(1) ISSN 0077-8923 Authors Smith, Moyra Spence, M Anne Flodman, Pamela Publication Date 2009 DOI 10.1111/j.1749-6632.2008.03571.x License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California THE YEAR IN HUMAN AND MEDICAL GENETICS 2009 Nuclear and Mitochondrial Genome Defects in Autisms Moyra Smith, M. Anne Spence, and Pamela Flodman Department of Pediatrics, University of California, Irvine, California In this review we will evaluate evidence that altered gene dosage and structure im- pacts neurodevelopment and neural connectivity through deleterious effects on synap- tic structure and function, and evidence that the latter are key contributors to the risk for autism. We will review information on alterations of structure of mitochondrial DNA and abnormal mitochondrial function in autism and indications that interactions of the nuclear and mitochondrial genomes may play a role in autism pathogenesis. In a final section we will present data derived using Affymetrixtm SNP 6.0 microar- ray analysis of DNA of a number of subjects and parents recruited to our autism spectrum disorders project. We include data on two sets of monozygotic twins. Col- lectively these data provide additional evidence of nuclear and mitochondrial genome imbalance in autism and evidence of specific candidate genes in autism. We present data on dosage changes in genes that map on the X chromosomes and the Y chro- mosome. -

TLR3-Dependent Activation of TLR2 Endogenous Ligands Via the Myd88 Signaling Pathway Augments the Innate Immune Response
cells Article TLR3-Dependent Activation of TLR2 Endogenous Ligands via the MyD88 Signaling Pathway Augments the Innate Immune Response 1 2, 1 3 Hellen S. Teixeira , Jiawei Zhao y, Ethan Kazmierski , Denis F. Kinane and Manjunatha R. Benakanakere 2,* 1 Department of Orthodontics, School of Dental Medicine, University of Pennsylvania, Philadelphia, PA 19004, USA; [email protected] (H.S.T.); [email protected] (E.K.) 2 Department of Periodontics, School of Dental Medicine, University of Pennsylvania, Philadelphia, PA 19004, USA; [email protected] 3 Periodontology Department, Bern Dental School, University of Bern, 3012 Bern, Switzerland; [email protected] * Correspondence: [email protected] Present address: Department of Pathology, Wayne State University School of Medicine, y 541 East Canfield Ave., Scott Hall 9215, Detroit, MI 48201, USA. Received: 30 June 2020; Accepted: 12 August 2020; Published: 17 August 2020 Abstract: The role of the adaptor molecule MyD88 is thought to be independent of Toll-like receptor 3 (TLR3) signaling. In this report, we demonstrate a previously unknown role of MyD88 in TLR3 signaling in inducing endogenous ligands of TLR2 to elicit innate immune responses. Of the various TLR ligands examined, the TLR3-specific ligand polyinosinic:polycytidylic acid (poly I:C), significantly induced TNF production and the upregulation of other TLR transcripts, in particular, TLR2. Accordingly, TLR3 stimulation also led to a significant upregulation of endogenous TLR2 ligands mainly, HMGB1 and Hsp60. By contrast, the silencing of TLR3 significantly downregulated MyD88 and TLR2 gene expression and pro-inflammatory IL1β, TNF, and IL8 secretion. The silencing of MyD88 similarly led to the downregulation of TLR2, IL1β, TNF and IL8, thus suggesting MyD88 / to somehow act downstream of TLR3. -

RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT
RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT Cat. no. 330231 PAHS-3012ZE For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Applied Biosystems® models 7900HT (384-well block), Format E ViiA™ 7 (384-well block); Bio-Rad CFX384™ RT² Profiler PCR Array, Roche® LightCycler® 480 (384-well block) Format G Description The Human Apoptosis RT² Profiler PCR Array profiles the expression of 370 key genes involved in apoptosis, or programmed cell death. The array includes the TNF ligands and their receptors; members of the bcl-2 family, BIR (baculoviral IAP repeat) domain proteins, CARD domain (caspase recruitment domain) proteins, death domain proteins, TRAF (TNF receptor-associated factor) domain proteins and caspases. These 370 genes are also grouped by their functional contribution to apoptosis, either anti-apoptosis or induction of apoptosis. Using real-time PCR, you can easily and reliably analyze expression of a focused panel of genes related to apoptosis with this array. For further details, consult the RT² Profiler PCR Array Handbook. Sample & Assay Technologies Shipping and storage RT² Profiler PCR Arrays in formats E and G are shipped at ambient temperature, on dry ice, or blue ice packs depending on destination and accompanying products. For long term storage, keep plates at –20°C. Note: Ensure that you have the correct RT² Profiler PCR Array format for your real-time cycler (see table above). Note: Open the package and store the products appropriately immediately -

MYD88 L265P Is a Marker Highly Characteristic Of, but Not Restricted To, Waldenstro¨M’S Macroglobulinemia
Leukemia (2013) 27, 1722–1728 & 2013 Macmillan Publishers Limited All rights reserved 0887-6924/13 www.nature.com/leu ORIGINAL ARTICLE MYD88 L265P is a marker highly characteristic of, but not restricted to, Waldenstro¨m’s macroglobulinemia C Jime´ nez1, E Sebastia´n1, MC Chillo´n1,2, P Giraldo3, J Mariano Herna´ndez4, F Escalante5, TJ Gonza´lez-Lo´ pez6, C Aguilera7, AG de Coca8, I Murillo3, M Alcoceba1, A Balanzategui1, ME Sarasquete1,2, R Corral1, LA Marı´n1, B Paiva1,2, EM Ocio1,2, NC Gutie´ rrez1,2, M Gonza´lez1,2, JF San Miguel1,2 and R Garcı´a-Sanz1,2 We evaluated the MYD88 L265P mutation in Waldenstro¨m’s macroglobulinemia (WM) and B-cell lymphoproliferative disorders by specific polymerase chain reaction (PCR) (sensitivity B10 À 3). No mutation was seen in normal donors, while it was present in 101/117 (86%) WM patients, 27/31 (87%) IgM monoclonal gammapathies of uncertain significance (MGUS), 3/14 (21%) splenic marginal zone lymphomas and 9/48 (19%) non-germinal center (GC) diffuse large B-cell lymphomas (DLBCLs). The mutation was absent in all 28 GC-DLBCLs, 13 DLBCLs not subclassified, 35 hairy cell leukemias, 39 chronic lymphocyticleukemias(16withM-component), 25 IgA or IgG-MGUS, 24 multiple myeloma (3 with an IgM isotype), 6 amyloidosis, 9 lymphoplasmacytic lymphomas and 1 IgM-related neuropathy. Among WM and IgM- MGUS, MYD88 L265P mutation was associated with some differences in clinical and biological characteristics, although usually minor; wild-type MYD88 cases had smaller M-component (1.77 vs 2.72 g/dl, P ¼ 0.022), more lymphocytosis (24 vs 5%, P ¼ 0.006), higher lactate dehydrogenase level (371 vs 265 UI/L, P ¼ 0.002), atypical immunophenotype (CD23 À CD27 þþFMC7 þþ), less Immunoglobulin Heavy Chain Variable gene (IGHV) somatic hypermutation (57 vs 97%, P ¼ 0.012) and less IGHV3–23 gene selection (9 vs 27%, P ¼ 0.014). -

Molecular Structure of Promoter-Bound Yeast TFIID
ARTICLE DOI: 10.1038/s41467-018-07096-y OPEN Molecular structure of promoter-bound yeast TFIID Olga Kolesnikova 1,2,3,4, Adam Ben-Shem1,2,3,4, Jie Luo5, Jeff Ranish 5, Patrick Schultz 1,2,3,4 & Gabor Papai 1,2,3,4 Transcription preinitiation complex assembly on the promoters of protein encoding genes is nucleated in vivo by TFIID composed of the TATA-box Binding Protein (TBP) and 13 TBP- associate factors (Tafs) providing regulatory and chromatin binding functions. Here we present the cryo-electron microscopy structure of promoter-bound yeast TFIID at a resolu- 1234567890():,; tion better than 5 Å, except for a flexible domain. We position the crystal structures of several subunits and, in combination with cross-linking studies, describe the quaternary organization of TFIID. The compact tri lobed architecture is stabilized by a topologically closed Taf5-Taf6 tetramer. We confirm the unique subunit stoichiometry prevailing in TFIID and uncover a hexameric arrangement of Tafs containing a histone fold domain in the Twin lobe. 1 Department of Integrated Structural Biology, Equipe labellisée Ligue Contre le Cancer, Institut de Génétique et de Biologie Moléculaire et Cellulaire, Illkirch 67404, France. 2 Centre National de la Recherche Scientifique, UMR7104, 67404 Illkirch, France. 3 Institut National de la Santé et de la Recherche Médicale, U1258, 67404 Illkirch, France. 4 Université de Strasbourg, Illkirch 67404, France. 5 Institute for Systems Biology, Seattle, WA 98109, USA. These authors contributed equally: Olga Kolesnikova, Adam Ben-Shem. Correspondence and requests for materials should be addressed to P.S. (email: [email protected]) or to G.P. -

MYD88 and Beyond: Novel Opportunities for Diagnosis, Prognosis and Treatment in Waldenstro¨M’S Macroglobulinemia
Leukemia (2014) 28, 1799–1803 & 2014 Macmillan Publishers Limited All rights reserved 0887-6924/14 www.nature.com/leu CONCISE REVIEW MYD88 and beyond: novel opportunities for diagnosis, prognosis and treatment in Waldenstro¨m’s Macroglobulinemia O Landgren and N Tageja Waldenstro¨m’s Macroglobulinemia (WM) is a rare disease of the elderly with a median age of 63–68 years at diagnosis. Despite recent progress in biological insights and therapeutics, WM remains clinically challenging to diagnose and is difficult to manage with significant morbidity and lack of established curative therapies. Recently, the use of whole-genome sequencing has helped to identify a highly recurrent somatic mutation, myeloid differentiation factor 88 [MYD88] L265P in WM. This has fueled major interest in the field and as newer evidence accumulates, it is clear that that discovery of MYD88 L265P mutation may represent an important breakthrough in understanding the pathogenesis of WM and lymphoproliferative disorders. Recent scientific work in this field has also guided the identification of new targets such as CXCR4 and PI3K-delta that may have major implications in the future treatment of WM. This review discusses the role of MYD88 L265P mutations as well as targets beyond MYD88 in the setting of pathogenesis and development of future rational therapeutic trials focusing on patients diagnosed with WM. Leukemia (2014) 28, 1799–1803; doi:10.1038/leu.2014.88 INTRODUCTION transduces signals to the NF-kB transcription factors in response to Waldenstro¨m’s Macroglobulinemia (WM) is a rare hematological IL-1R1 signaling. MYD88 has a modular structure with a Toll/IL-1R malignancy with a reported age-adjusted incidence rate of 3.4 per (TIR) domain at its COOH terminus and a death domain at its 12 11 million among men and 1.7 per million among women in the NH2 terminus. -

The Role of Protein Clearance Mechanisms in Organismal Ageing and Age-Related Diseases
REVIEW Received 18 Mar 2014 | Accepted 24 Oct 2014 | Published 8 Dec 2014 DOI: 10.1038/ncomms6659 The role of protein clearance mechanisms in organismal ageing and age-related diseases David Vilchez1, Isabel Saez1 & Andrew Dillin2,3 The ability to maintain a functional proteome, or proteostasis, declines during the ageing process. Damaged and misfolded proteins accumulate with age, impairing cell function and tissue homeostasis. The accumulation of damaged proteins contributes to multiple age- related diseases such as Alzheimer’s, Parkinson’s or Huntington’s disease. Damaged proteins are degraded by the ubiquitin–proteasome system or through autophagy-lysosome, key components of the proteostasis network. Modulation of either proteasome activity or autophagic-lysosomal potential extends lifespan and protects organisms from symptoms associated with proteostasis disorders, suggesting that protein clearance mechanisms are directly linked to ageing and age-associated diseases. he integrity of the proteome, or proteostasis, is challenged during the ageing process. Damaged proteins accumulate as a consequence of ageing and may ensue from the Taccumulation of reactive oxygen species and a progressive decline in the ability to maintain a functional proteome1. This demise in proteostasis is considered one of the hallmarks of ageing1 and contributes to multiple age-related diseases such as Alzheimer’s (AD)2, Parkinson’s (PD)3 or Huntington’s disease (HD)4. Proteostasis is maintained by a network of cellular mechanisms that monitors folding, concentration, cellular localization and interactions of proteins from their synthesis through their degradation5. Chaperones assure the proper folding of proteins throughout their life cycle and under stress conditions but their activity declines with age (reviewed in refs 6–10). -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

And Sepsis-Induced Lung Inflammation and Mediates Myd88
The Journal of Immunology Caveolin-1 Tyr14 Phosphorylation Induces Interaction with TLR4 in Endothelial Cells and Mediates MyD88-Dependent Signaling and Sepsis-Induced Lung Inflammation Hao Jiao,*,†,1 Yang Zhang,*,†,1 Zhibo Yan,* Zhen-Guo Wang,* Gongjian Liu,† Richard D. Minshall,*,‡ Asrar B. Malik,‡ and Guochang Hu*,‡ Activation of TLR4 by the endotoxin LPS is a critical event in the pathogenesis of Gram-negative sepsis. Caveolin-1, the signaling protein associated with caveolae, is implicated in regulating the lung inflammatory response to LPS; however, the mechanism is not understood. In this study, we investigated the role of caveolin-1 in regulating TLR4 signaling in endothelial cells. We observed that LPS interaction with CD14 in endothelial cells induced Src-dependent caveolin-1 phosphorylation at Tyr14. Using a TLR4-MD2- CD14–transfected HEK-293 cell line and caveolin-1–deficient (cav-12/2) mouse lung microvascular endothelial cells, we demon- strated that caveolin-1 phosphorylation at Tyr14 following LPS exposure induced caveolin-1 and TLR4 interaction and, thereby, TLR4 activation of MyD88, leading to NF-kB activation and generation of proinflammatory cytokines. Exogenous expression of phosphorylation-deficient Y14F caveolin-1 mutant in cav-12/2 mouse pulmonary vasculature rendered the mice resistant to LPS compared with reintroduction of wild-type caveolin-1. Thus, caveolin-1 Y14 phosphorylation was required for the interaction with TLR4 and activation of TLR4-MyD88 signaling and sepsis-induced lung inflammation. Inhibiting caveolin-1 Tyr14 phosphoryla- tion and resultant inactivation of TLR4 signaling in pulmonary vascular endothelial cells represent a novel strategy for preventing sepsis-induced lung inflammation and injury. -

Pathway Analysis of Breast, Colorectal, Pancreatic Cancers and Glioblastoma
Pathway analysis ofbreast, colorectal, pancreatic cancers and glioblastoma Dongyan Song Degree project inbiology, Bachelor ofscience, 2009 Examensarbete ibiologi 30 hp tillkandidatexamen, 2009 Biology Education Centre, Uppsala University and Department ofGenetics and Pathology Supervisor: Tobias Sjöblom Pathway analysis of breast, colorectal, pancreatic cancers and glioblastoma Summary Cancer is a genetic disease, and due to the multi-step progression, mutated genes accumulate in genome, leading to a gene spectrum with a few frequently mutated genes and a bunch of infrequently mutated genes. Because of the complexity of mutation profile in gene level, this study tries to analyze cancer mutations in a pathway level, implemented as clusters in this case. Initial attempts used phylogenetic analysis, which gave a result that breast and colorectal cancers cannot be distinguished from each other only by information of mutated genes but pancreatic cancers and glioblastoma formed a cluster distinguishing with patients with pancreatic cancer or glioblastoma. Thus, to some extent, the pattern of mutated genes can distinguish different cancers or different patients with same cancer type, but the results are not clear enough to give a conclusion. So, I implemented network methodology to study mutational pathway patterns in breast, colorectal, pancreatic cancers and glioblastoma. The initial network was constructed based on gene-gene interaction pairs identified from literature mining and the STRING database. PubMed IDs were retrieved with Entrez Gene IDs as queries, and via checking the overlap of PubMed IDs, 135,863,922 potential gene-gene interaction pairs were found. Negative logarithms of co-occurrence probabilities were calculated and the number of pairs was reduced by setting the significance level at 99.7% in the Poisson distribution. -
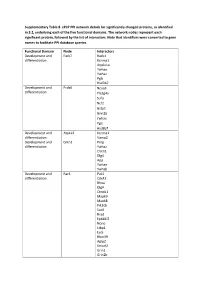
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4