Identi Cation of Molecular and Immune Prognostic Classi Ers of Acute Myeloid Leukemia
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Table 1. Complete Gene Lists and GO Terms from Figure 3C
Supplemental Table 1. Complete gene lists and GO terms from Figure 3C. Path 1 Genes: RP11-34P13.15, RP4-758J18.10, VWA1, CHD5, AZIN2, FOXO6, RP11-403I13.8, ARHGAP30, RGS4, LRRN2, RASSF5, SERTAD4, GJC2, RHOU, REEP1, FOXI3, SH3RF3, COL4A4, ZDHHC23, FGFR3, PPP2R2C, CTD-2031P19.4, RNF182, GRM4, PRR15, DGKI, CHMP4C, CALB1, SPAG1, KLF4, ENG, RET, GDF10, ADAMTS14, SPOCK2, MBL1P, ADAM8, LRP4-AS1, CARNS1, DGAT2, CRYAB, AP000783.1, OPCML, PLEKHG6, GDF3, EMP1, RASSF9, FAM101A, STON2, GREM1, ACTC1, CORO2B, FURIN, WFIKKN1, BAIAP3, TMC5, HS3ST4, ZFHX3, NLRP1, RASD1, CACNG4, EMILIN2, L3MBTL4, KLHL14, HMSD, RP11-849I19.1, SALL3, GADD45B, KANK3, CTC- 526N19.1, ZNF888, MMP9, BMP7, PIK3IP1, MCHR1, SYTL5, CAMK2N1, PINK1, ID3, PTPRU, MANEAL, MCOLN3, LRRC8C, NTNG1, KCNC4, RP11, 430C7.5, C1orf95, ID2-AS1, ID2, GDF7, KCNG3, RGPD8, PSD4, CCDC74B, BMPR2, KAT2B, LINC00693, ZNF654, FILIP1L, SH3TC1, CPEB2, NPFFR2, TRPC3, RP11-752L20.3, FAM198B, TLL1, CDH9, PDZD2, CHSY3, GALNT10, FOXQ1, ATXN1, ID4, COL11A2, CNR1, GTF2IP4, FZD1, PAX5, RP11-35N6.1, UNC5B, NKX1-2, FAM196A, EBF3, PRRG4, LRP4, SYT7, PLBD1, GRASP, ALX1, HIP1R, LPAR6, SLITRK6, C16orf89, RP11-491F9.1, MMP2, B3GNT9, NXPH3, TNRC6C-AS1, LDLRAD4, NOL4, SMAD7, HCN2, PDE4A, KANK2, SAMD1, EXOC3L2, IL11, EMILIN3, KCNB1, DOK5, EEF1A2, A4GALT, ADGRG2, ELF4, ABCD1 Term Count % PValue Genes regulation of pathway-restricted GDF3, SMAD7, GDF7, BMPR2, GDF10, GREM1, BMP7, LDLRAD4, SMAD protein phosphorylation 9 6.34 1.31E-08 ENG pathway-restricted SMAD protein GDF3, SMAD7, GDF7, BMPR2, GDF10, GREM1, BMP7, LDLRAD4, phosphorylation -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Single-Cell RNA-Seq of Dopaminergic Neurons Informs Candidate Gene Selection for Sporadic
bioRxiv preprint doi: https://doi.org/10.1101/148049; this version posted August 31, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Single-cell RNA-seq of dopaminergic neurons informs candidate gene selection for sporadic 2 Parkinson's disease 3 4 Paul W. Hook1, Sarah A. McClymont1, Gabrielle H. Cannon1, William D. Law1, A. Jennifer 5 Morton2, Loyal A. Goff1,3*, Andrew S. McCallion1,4,5* 6 7 1McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins University School of 8 Medicine, Baltimore, Maryland, United States of America 9 2Department of Physiology Development and Neuroscience, University of Cambridge, 10 Cambridge, United Kingdom 11 3Department of Neuroscience, Johns Hopkins University School of Medicine, Baltimore, 12 Maryland, United States of America 13 4Department of Comparative and Molecular Pathobiology, Johns Hopkins University School of 14 Medicine, Baltimore, Maryland, United States of America 15 5Department of Medicine, Johns Hopkins University School of Medicine, Baltimore, Maryland, 16 United States of America 17 *, To whom correspondence should be addressed: [email protected] and [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/148049; this version posted August 31, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. -

Age-Related DNA Hydroxymethylation Is Enriched for Gene Expression and Immune System Processes in Human Peripheral Blood
Epigenetics ISSN: 1559-2294 (Print) 1559-2308 (Online) Journal homepage: https://www.tandfonline.com/loi/kepi20 Age-related DNA hydroxymethylation is enriched for gene expression and immune system processes in human peripheral blood Nicholas D. Johnson, Luoxiu Huang, Ronghua Li, Yun Li, Yuchen Yang, Hye Rim Kim, Crystal Grant, Hao Wu, Eric A. Whitsel, Douglas P. Kiel, Andrea A. Baccarelli, Peng Jin, Joanne M. Murabito & Karen N. Conneely To cite this article: Nicholas D. Johnson, Luoxiu Huang, Ronghua Li, Yun Li, Yuchen Yang, Hye Rim Kim, Crystal Grant, Hao Wu, Eric A. Whitsel, Douglas P. Kiel, Andrea A. Baccarelli, Peng Jin, Joanne M. Murabito & Karen N. Conneely (2019): Age-related DNA hydroxymethylation is enriched for gene expression and immune system processes in human peripheral blood, Epigenetics, DOI: 10.1080/15592294.2019.1666651 To link to this article: https://doi.org/10.1080/15592294.2019.1666651 View supplementary material Accepted author version posted online: 11 Sep 2019. Published online: 26 Sep 2019. Submit your article to this journal Article views: 179 View related articles View Crossmark data Full Terms & Conditions of access and use can be found at https://www.tandfonline.com/action/journalInformation?journalCode=kepi20 EPIGENETICS https://doi.org/10.1080/15592294.2019.1666651 RESEARCH PAPER Age-related DNA hydroxymethylation is enriched for gene expression and immune system processes in human peripheral blood Nicholas D. Johnsona,b, Luoxiu Huanga, Ronghua Lia, Yun Li c,d, Yuchen Yangc, Hye Rim Kim a,e, Crystal Grant a,f, Hao Wug, Eric A. Whitsel h,i, Douglas P. Kiel j, Andrea A. Baccarellik, Peng Jin a, Joanne M. -

Neural Development
31 October 2007 NEURAL DEVELOPMENT www.neuraldevelopment.com Analysis of Lrrn1 expression and its relationship to neuromeric boundaries during chick neural development Laura C Andreae et al. Neural Development 2007, 2:22 http://www.neuraldevelopment.com/content/2/1/22 Neural Development BioMed Central Research article Open Access Analysis of Lrrn1 expression and its relationship to neuromeric boundaries during chick neural development LauraCAndreae1,2, Daniela Peukert1, Andrew Lumsden1 and Jonathan D Gilthorpe*1 Address: 1MRC Centre for Developmental Neurobiology, King's College London, New Hunt's House, Guy's Campus, London, UK, SE1 1UL and 2Department of Neurophysiology, National Institute for Medical Research, The Ridgeway, Mill Hill, London, UK, NW7 1AA Email: Laura C Andreae - [email protected]; Daniela Peukert - [email protected]; Andrew Lumsden - [email protected]; Jonathan D Gilthorpe* - [email protected] * Corresponding author Published: 31 October 2007 Received: 26 March 2007 Accepted: 31 October 2007 Neural Development 2007, 2:22 doi:10.1186/1749-8104-2-22 This article is available from: http://www.neuraldevelopment.com/content/2/1/22 © 2007 Andreae et al.; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The Drosophila leucine-rich repeat proteins Tartan (TRN) and Capricious (CAPS) mediate cell affinity differences during compartition of the wing imaginal disc. This study aims to identify and characterize the expression of a chick orthologue of TRN/CAPS and examine its potential function in relation to compartment boundaries in the vertebrate central nervous system. -

Genetic and Genomics Laboratory Tools and Approaches
Genetic and Genomics Laboratory Tools and Approaches Meredith Yeager, PhD Cancer Genomics Research Laboratory Division of Cancer Epidemiology and Genetics [email protected] DCEG Radiation Epidemiology and Dosimetry Course 2019 www.dceg.cancer.gov/RadEpiCourse (Recent) history of genetics 2 Sequencing of the Human Genome Science 291, 1304-1351 (2001) 3 The Human Genome – 2019 • ~3.3 billion bases (A, C, G, T) • ~20,000 protein-coding genes, many non-coding RNAs (~2% of the genome) • Annotation ongoing – the initial sequencing in 2001 is still being refined, assembled and annotated, even now – hg38 • Variation (polymorphism) present within humans – Population-specific – Cosmopolitan 4 Types of polymorphisms . Single nucleotide polymorphisms (SNPs) . Common SNPs are defined as > 5% in at least one population . Abundant in genome (~50 million and counting) ATGGAACGA(G/C)AGGATA(T/A)TACGCACTATGAAG(C/A)CGGTGAGAGG . Repeats of DNA (long, short, complex, simple), insertions/deletions . A small fraction of SNPs and other types of variation are very or slightly deleterious and may contribute by themselves or with other genetic or environmental factors to a phenotype or disease 5 Different mutation rates at the nucleotide level Mutation type Mutation rate (per generation) Transition on a CpG 1.6X10-7 Transversion on a CpG 4.4X10-8 Transition: purine to purine Transition out of CpG 1.2X10-8 Transversion: purine to pyrimidine Transversion out of CpG 5.5X10-9 Substitution (average) 2.3X10-8 A and G are purines Insertion/deletion (average) 2.3X10-9 C and T are pyrimidines Mutation rate (average) 2.4X10-8 . Size of haploid genome : 3.3X109 nucleotides . -

Islet-1 Is Essential for Pancreatic Β-Cell Function Benjamin N. Ediger
Page 1 of 68 Diabetes June 20, 2014 Diabetes Islet-1 is essential for pancreatic β-cell function Benjamin N. Ediger1,5, Aiping Du1, Jingxuan Liu1, Chad S. Hunter3, Erik R. Walp1, Jonathan Schug4, Klaus H. Kaestner4, Roland Stein3, Doris A. Stoffers5* and Catherine Lee May*,1,2 1Department of Pathology and Laboratory Medicine, Children’s Hospital of Philadelphia, 2Department of Pathology and Laboratory Medicine, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA 3Department of Molecular Physiology and Biophysics, Vanderbilt University Medical Center, Nashville, Tennessee 37232, USA 4Department of Genetics and Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA 5Department of Medicine and Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, USA * These authors contributed equally. Running Title: Isl-1 regulates β-cell function Address correspondence to: Catherine Lee May, Ph.D. 3615 Civic Center Blvd, Room 516E Philadelphia, PA 19104 Phone: 267-426-0116 E-mail: [email protected] And Doris A. Stoffers, M.D., Ph.D. 3400 Civic Center Boulevard, 12-124 SCTR Philadelphia, PA 19104 Phone: (215) 573-5413 E-mail: [email protected] Fax: 215-590-3709 Word Count: 4065 Number of Tables: 6 (all are supplemental) Number of Figures: 9 (3 are supplemental) Diabetes Publish Ahead of Print, published online July 15, 2014 Diabetes Page 2 of 68 Abstract Isl-1 is essential for the survival and ensuing differentiation of pancreatic endocrine progenitors. Isl-1 remains expressed in all adult pancreatic endocrine lineages; however, its specific function in the postnatal pancreas is unclear. -

Prognostic Stratification of Adult Primary Glioblastoma Multiforme Patients Based on Their Tumor Gene Amplification Profiles
www.oncotarget.com Oncotarget, 2018, Vol. 9, (No. 46), pp: 28083-28102 Research Paper Prognostic stratification of adult primary glioblastoma multiforme patients based on their tumor gene amplification profiles María González-Tablas1, Inês Crespo2,3, Ana Luísa Vital2,3, Álvaro Otero4, Ana Belén Nieto5, Pablo Sousa4, María Carmen Patino-Alonso5, Luis Antonio Corchete6, Hermínio Tão7, Olinda Rebelo8, Marcos Barbosa7,9, Maria Rosário Almeida2, Ana Filipa Guedes2, María Celeste Lopes2,3, Pim J. French10, Alberto Orfao1,11,* and María Dolores Tabernero1,11,* 1Centre for Cancer Research (CIC IBMCC-CSIC/USAL), Department of Medicine, CIBERONC, University of Salamanca, Salamanca, Spain 2Centre for Neuroscience and Cell Biology, University of Coimbra, Coimbra, Portugal 3Faculty of Pharmacy, University of Coimbra, Coimbra, Portugal 4Servicio de Neurocirugía, Hospital Universitario e Instituto Biosanitario de Salamanca (IBSAL), Salamanca, Spain 5Department of Statistics, University of Salamanca, Salamanca, Spain 6Departamento de Hematología, Hospital Universitario, IBSAL, IBMCC (USAL-CSIC), Salamanca, Spain 7Neurosurgery Service, University Hospital of Coimbra, Coimbra, Portugal 8Neuropathology Laboratory, Neurology Service, University Hospital of Coimbra, Coimbra, Portugal 9Faculty of Medicine, University of Coimbra, Coimbra, Portugal 10Department of Neurology, Erasmus MC, Rotterdam, The Netherlands 11Instituto Biosanitario de Salamanca (IBSAL), Salamanca, Spain *Both authors have equally contributed to this work and should be considered as last authors Correspondence to: María Dolores Tabernero, email: [email protected] Keywords: glioblastoma; classification; subtypes; gene amplification; survival Received: February 28, 2018 Accepted: May 14, 2018 Published: June 15, 2018 Copyright: González-Tablas et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. -

NIH Public Access Author Manuscript Science
NIH Public Access Author Manuscript Science. Author manuscript; available in PMC 2013 August 23. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Science. 2013 June 21; 340(6139): 1467–1471. doi:10.1126/science.1235488. GWAS of 126,559 Individuals Identifies Genetic Variants Associated with Educational Attainment Cornelius A. Rietveld1,2, Sarah E. Medland3, Jaime Derringer4, Jian Yang5, Tõnu Esko6, Nicolas W. Martin3,7, Harm-Jan Westra8, Konstantin Shakhbazov5,9, Abdel Abdellaoui10, Arpana Agrawal11, Eva Albrecht12, Behrooz Z. Alizadeh13, Najaf Amin14, John Barnard15, Sebastian E. Baumeister16, Kelly S. Benke17, Lawrence F. Bielak18, Jeffrey A. Boatman19, Patricia A. Boyle20, Gail Davies21, Christiaan de Leeuw22, Niina Eklund24,25, Daniel S. Evans26, Rudolf Ferhmann8, Krista Fischer6, Christian Gieger12, Håkon K. Gjessing27, Sara Hägg28,29,30, Jennifer R. Harris27, Caroline Hayward31, Christina Holzapfel32,33, Carla A. Ibrahim-Verbaas14,34, Erik Ingelsson28,29,30, Bo Jacobsson27,35, Peter K. Joshi36, Astanand Jugessur27, Marika Kaakinen37,38, Stavroula Kanoni39, Juha Karjalainen8, Ivana Kolcic40, Kati Kristiansson24,25, Zoltán Kutalik41,42, Jari Lahti43, Sang H. Lee3, Peng Lin11, Penelope A. Lind3, Yongmei Liu44, Kurt Lohman45, Marisa Loitfelder46, George McMahon47, Pedro Marques Vidal48, Osorio Meirelles49, Lili Milani6, Ronny Myhre27, Marja-Liisa Nuotio24,25, Christopher J. Oldmeadow50, Katja E. Petrovic51, Wouter J. Peyrot52, Ozren Polašek40, Lydia Quaye53, Eva Reinmaa6, John P. Rice11, Thais S. Rizzi22, Helena Schmidt54, Reinhold Schmidt46, Albert V. Smith55,56, Jennifer A. Smith18, Toshiko Tanaka49, Antonio Terracciano49,57, Matthijs J.H.M. van der Loos1,2, Veronique Vitart31, Henry Völzke16, Jürgen Wellmann58, Lei Yu20, Wei Zhao18, Jüri Allik59, John R. -

Genome-Wide Association Study of Cognitive Flexibility Assessed by the Wisconsin Card Sorting Test H
Donald and Barbara Zucker School of Medicine Journal Articles Academic Works 2018 Genome-wide association study of cognitive flexibility assessed by the Wisconsin Card Sorting Test H. Zhang H. Zhou T. Lencz Zucker School of Medicine at Hofstra/Northwell L. A. Farrer H. R. Kranzler See next page for additional authors Follow this and additional works at: https://academicworks.medicine.hofstra.edu/articles Part of the Psychiatry Commons Recommended Citation Zhang H, Zhou H, Lencz T, Farrer LA, Kranzler HR, Gelernter J. Genome-wide association study of cognitive flexibility assessed by the Wisconsin Card Sorting Test. 2018 Jan 01; 177(5):Article 4033 [ p.]. Available from: https://academicworks.medicine.hofstra.edu/articles/4033. Free full text article. This Article is brought to you for free and open access by Donald and Barbara Zucker School of Medicine Academic Works. It has been accepted for inclusion in Journal Articles by an authorized administrator of Donald and Barbara Zucker School of Medicine Academic Works. For more information, please contact [email protected]. Authors H. Zhang, H. Zhou, T. Lencz, L. A. Farrer, H. R. Kranzler, and J. Gelernter This article is available at Donald and Barbara Zucker School of Medicine Academic Works: https://academicworks.medicine.hofstra.edu/articles/4033 HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Am J Med Manuscript Author Genet B Neuropsychiatr Manuscript Author Genet. Author manuscript; available in PMC 2019 July 01. Published in final edited form as: Am J Med Genet B Neuropsychiatr Genet. 2018 July ; 177(5): 511–519. doi:10.1002/ajmg.b.32642. Genome-wide association study of cognitive flexibility assessed by the Wisconsin Card Sorting Test Huiping Zhang, Ph.D.1,2, Hang Zhou, Ph.D.5, Todd Lencz, Ph.D.8, Lindsay A. -

Predict AID Targeting in Non-Ig Genes Multiple Transcription Factor
Downloaded from http://www.jimmunol.org/ by guest on September 26, 2021 is online at: average * The Journal of Immunology published online 20 March 2013 from submission to initial decision 4 weeks from acceptance to publication Multiple Transcription Factor Binding Sites Predict AID Targeting in Non-Ig Genes Jamie L. Duke, Man Liu, Gur Yaari, Ashraf M. Khalil, Mary M. Tomayko, Mark J. Shlomchik, David G. Schatz and Steven H. Kleinstein J Immunol http://www.jimmunol.org/content/early/2013/03/20/jimmun ol.1202547 Submit online. Every submission reviewed by practicing scientists ? is published twice each month by http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://www.jimmunol.org/content/suppl/2013/03/20/jimmunol.120254 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 26, 2021. Published March 20, 2013, doi:10.4049/jimmunol.1202547 The Journal of Immunology Multiple Transcription Factor Binding Sites Predict AID Targeting in Non-Ig Genes Jamie L. Duke,* Man Liu,†,1 Gur Yaari,‡ Ashraf M. Khalil,x Mary M. Tomayko,{ Mark J. Shlomchik,†,x David G. -
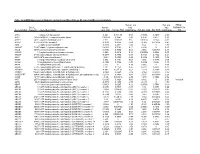
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde