Homologue-Specific Chromosome Sequencing Characterizes Translocation Junctions and Permits Allelic Assignment Fumio Kasai1,2,*, Jorge C
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
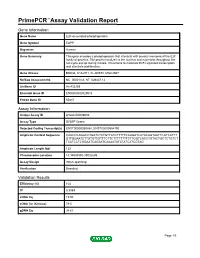
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name E2F-associated phosphoprotein Gene Symbol EAPP Organism Human Gene Summary This gene encodes a phosphoprotein that interacts with several members of the E2F family of proteins. The protein localizes to the nucleus and is present throughout the cell cycle except during mitosis. It functions to modulate E2F-regulated transcription and stimulate proliferation. Gene Aliases BM036, C14orf11, FLJ20578, MGC4957 RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.433269 Ensembl Gene ID ENSG00000129518 Entrez Gene ID 55837 Assay Information Unique Assay ID qHsaCID0008053 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000250454, ENST00000554792 Amplicon Context Sequence CAACCCAGGCCTGATCTCTGTTATCTTTTTCAGGATCATACAGTAATTCGTCATTT GTTGGAATCTTGTGTTGTTTCTTCTTTTTTTTCTTGGTCACCTGTACTGCTCTGTCT TCATCCTCGGAATCAGAATCAAAATATATATCATCGTAG Amplicon Length (bp) 122 Chromosome Location 14:34998580-35002699 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 104 R2 0.9989 cDNA Cq 19.93 cDNA Tm (Celsius) 79.5 gDNA Cq 34.63 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report EAPP, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Evidence for Differential Alternative Splicing in Blood of Young Boys With
Stamova et al. Molecular Autism 2013, 4:30 http://www.molecularautism.com/content/4/1/30 RESEARCH Open Access Evidence for differential alternative splicing in blood of young boys with autism spectrum disorders Boryana S Stamova1,2,5*, Yingfang Tian1,2,4, Christine W Nordahl1,3, Mark D Shen1,3, Sally Rogers1,3, David G Amaral1,3 and Frank R Sharp1,2 Abstract Background: Since RNA expression differences have been reported in autism spectrum disorder (ASD) for blood and brain, and differential alternative splicing (DAS) has been reported in ASD brains, we determined if there was DAS in blood mRNA of ASD subjects compared to typically developing (TD) controls, as well as in ASD subgroups related to cerebral volume. Methods: RNA from blood was processed on whole genome exon arrays for 2-4–year-old ASD and TD boys. An ANCOVA with age and batch as covariates was used to predict DAS for ALL ASD (n=30), ASD with normal total cerebral volumes (NTCV), and ASD with large total cerebral volumes (LTCV) compared to TD controls (n=20). Results: A total of 53 genes were predicted to have DAS for ALL ASD versus TD, 169 genes for ASD_NTCV versus TD, 1 gene for ASD_LTCV versus TD, and 27 genes for ASD_LTCV versus ASD_NTCV. These differences were significant at P <0.05 after false discovery rate corrections for multiple comparisons (FDR <5% false positives). A number of the genes predicted to have DAS in ASD are known to regulate DAS (SFPQ, SRPK1, SRSF11, SRSF2IP, FUS, LSM14A). In addition, a number of genes with predicted DAS are involved in pathways implicated in previous ASD studies, such as ROS monocyte/macrophage, Natural Killer Cell, mTOR, and NGF signaling. -

Cell-Type–Specific Eqtl of Primary Melanocytes Facilitates Identification of Melanoma Susceptibility Genes
Downloaded from genome.cshlp.org on November 19, 2018 - Published by Cold Spring Harbor Laboratory Press Research Cell-type–specific eQTL of primary melanocytes facilitates identification of melanoma susceptibility genes Tongwu Zhang,1,7 Jiyeon Choi,1,7 Michael A. Kovacs,1 Jianxin Shi,2 Mai Xu,1 NISC Comparative Sequencing Program,9 Melanoma Meta-Analysis Consortium,10 Alisa M. Goldstein,3 Adam J. Trower,4 D. Timothy Bishop,4 Mark M. Iles,4 David L. Duffy,5 Stuart MacGregor,5 Laufey T. Amundadottir,1 Matthew H. Law,5 Stacie K. Loftus,6 William J. Pavan,6,8 and Kevin M. Brown1,8 1Laboratory of Translational Genomics, Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892, USA; 2Biostatistics Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892, USA; 3Clinical Genetics Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892, USA; 4Section of Epidemiology and Biostatistics, Leeds Institute of Cancer and Pathology, University of Leeds, Leeds, LS9 7TF, United Kingdom; 5Statistical Genetics, QIMR Berghofer Medical Research Institute, Brisbane, Queensland, 4006, Australia; 6Genetic Disease Research Branch, National Human Genome Research Institute, National Institutes of Health, Bethesda, Maryland 20892, USA Most expression quantitative trait locus (eQTL) studies to date have been performed in heterogeneous tissues as opposed to specific cell types. To better understand the cell-type–specific regulatory landscape of human melanocytes, which give rise to melanoma but account for <5% of typical human skin biopsies, we performed an eQTL analysis in primary melanocyte cul- tures from 106 newborn males. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Prospective Diagnostic Analysis of Copy Number Variants Using SNP Microarrays in Individuals with Autism Spectrum Disorders
European Journal of Human Genetics (2013), 1–8 & 2013 Macmillan Publishers Limited All rights reserved 1018-4813/13 www.nature.com/ejhg ARTICLE Prospective diagnostic analysis of copy number variants using SNP microarrays in individuals with autism spectrum disorders Caroline Nava1,2,3,4,19, Boris Keren5,19, Cyril Mignot4,6,7,8, Agne`s Rastetter1,2,3, Sandra Chantot-Bastaraud9, Anne Faudet4, Eric Fonteneau5, Claire Amiet10, Claudine Laurent1,2,10, Aure´lia Jacquette4,7,8, Sandra Whalen4, Alexandra Afenjar4,6,7,8,11, Didier Pe´risse10,12, Diane Doummar6, Nathalie Dorison6,11, Marion Leboyer13,14,15,16, Jean-Pierre Siffroi9, David Cohen10,17, Alexis Brice*,1,2,3,4,18, Delphine He´ron4,6,7,8 and Christel Depienne*,1,2,3,4,18 Copy number variants (CNVs) have repeatedly been found to cause or predispose to autism spectrum disorders (ASDs). For diagnostic purposes, we screened 194 individuals with ASDs for CNVs using Illumina SNP arrays. In several probands, we also analyzed candidate genes located in inherited deletions to unmask autosomal recessive variants. Three CNVs, a de novo triplication of chromosome 15q11–q12 of paternal origin, a deletion on chromosome 9p24 and a de novo 3q29 deletion, were identified as the cause of the disorder in one individual each. An autosomal recessive cause was considered possible in two patients: a homozygous 1p31.1 deletion encompassing PTGER3 and a deletion of the entire DOCK10 gene associated with a rare hemizygous missense variant. We also identified multiple private or recurrent CNVs, the majority of which were inherited from asymptomatic parents. Although highly penetrant CNVs or variants inherited in an autosomal recessive manner were detected in rare cases, our results mainly support the hypothesis that most CNVs contribute to ASDs in association with other CNVs or point variants located elsewhere in the genome. -
![CBWD1 Mouse Monoclonal Antibody [Clone ID: OTI3F9] Product Data](https://docslib.b-cdn.net/cover/2738/cbwd1-mouse-monoclonal-antibody-clone-id-oti3f9-product-data-1162738.webp)
CBWD1 Mouse Monoclonal Antibody [Clone ID: OTI3F9] Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for CF501612 CBWD1 Mouse Monoclonal Antibody [Clone ID: OTI3F9] Product data: Product Type: Primary Antibodies Clone Name: OTI3F9 Applications: FC, IF, IHC, WB Recommended Dilution: WB 1:500~2000, IHC 1:150, IF 1:100, FLOW 1:100 Reactivity: Human Host: Mouse Isotype: IgG2b Clonality: Monoclonal Immunogen: Full length human recombinant protein of human CBWD1 (NP_060961) produced in HEK293T cell. Formulation: Lyophilized powder (original buffer 1X PBS, pH 7.3, 8% trehalose) Reconstitution Method: For reconstitution, we recommend adding 100uL distilled water to a final antibody concentration of about 1 mg/mL. To use this carrier-free antibody for conjugation experiment, we strongly recommend performing another round of desalting process. (OriGene recommends Zeba Spin Desalting Columns, 7KMWCO from Thermo Scientific) Purification: Purified from mouse ascites fluids or tissue culture supernatant by affinity chromatography (protein A/G) Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Predicted Protein Size: 43.9 kDa Gene Name: Homo sapiens COBW domain containing 1 (CBWD1), transcript variant 1, mRNA. Database Link: NP_060961 Entrez Gene 55871 Human Q9BRT8 Synonyms: COBP This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 7 CBWD1 Mouse Monoclonal Antibody [Clone ID: OTI3F9] – CF501612 Product images: HEK293T cells were transfected with the pCMV6- ENTRY control (Cat# [PS100001], Left lane) or pCMV6-ENTRY CBWD1 (Cat# [RC222790], Right lane) cDNA for 48 hrs and lysed. -

Mapping of a Chromosome 12 Region Associated with Airway Hyperresponsiveness in a Recombinant Congenic Mouse Strain and Selectio
Mapping of a Chromosome 12 Region Associated with Airway Hyperresponsiveness in a Recombinant Congenic Mouse Strain and Selection of Potential Candidate Genes by Expression and Sequence Variation Analyses Cynthia Kanagaratham1*, Rafael Marino2, Pierre Camateros2, John Ren3, Daniel Houle4, Robert Sladek1,2,5, Silvia M. Vidal1,3, Danuta Radzioch1,2 1 Department of Human Genetics, McGill University, Montreal, Quebec, Canada, 2 Faculty of Medicine, Division of Experimental Medicine, McGill University, Montreal, Quebec, Canada, 3 Department of Microbiology and Immunology, McGill University, Montreal, Quebec, Canada, 4 Research Institute of the McGill University Health Center, Montreal, Quebec, Canada, 5 McGill University and Genome Quebec Innovation Centre, Montreal, Quebec, Canada Abstract In a previous study we determined that BcA86 mice, a strain belonging to a panel of AcB/BcA recombinant congenic strains, have an airway responsiveness phenotype resembling mice from the airway hyperresponsive A/J strain. The majority of the BcA86 genome is however from the hyporesponsive C57BL/6J strain. The aim of this study was to identify candidate regions and genes associated with airway hyperresponsiveness (AHR) by quantitative trait locus (QTL) analysis using the BcA86 strain. Airway responsiveness of 205 F2 mice generated from backcrossing BcA86 strain to C57BL/6J strain was measured and used for QTL analysis to identify genomic regions in linkage with AHR. Consomic mice for the QTL containing chromosomes were phenotyped to study the contribution of each chromosome to lung responsiveness. Candidate genes within the QTL were selected based on expression differences in mRNA from whole lungs, and the presence of coding non- synonymous mutations that were predicted to have a functional effect by amino acid substitution prediction tools. -

Triangulating Molecular Evidence to Prioritise Candidate Causal Genes at Established Atopic Dermatitis Loci
medRxiv preprint doi: https://doi.org/10.1101/2020.11.30.20240838; this version posted November 30, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-ND 4.0 International license . Triangulating molecular evidence to prioritise candidate causal genes at established atopic dermatitis loci Maria K Sobczyk1, Tom G Richardson1, Verena Zuber2,3, Josine L Min1, eQTLGen Consortium4, BIOS Consortium5, GoDMC, Tom R Gaunt1, Lavinia Paternoster1* 1) MRC Integrative Epidemiology Unit, Bristol Medical School, University of Bristol, Bristol, UK 2) Department of Epidemiology and Biostatistics, School of Public Health, Imperial College London, London, UK 3) MRC Biostatistics Unit, School of Clinical Medicine, University of Cambridge, Cambridge, UK 4) Members of the eQTLGen Consortium are listed in: Supplementary_Consortium_members.docx 5) Members of the BIOS Consortium are listed in: Supplementary_Consortium_members.docx Abstract Background: Genome-wide association studies for atopic dermatitis (AD, eczema) have identified 25 reproducible loci associated in populations of European descent. We attempt to prioritise candidate causal genes at these loci using a multifaceted bioinformatic approach and extensive molecular resources compiled into a novel pipeline: ADGAPP (Atopic Dermatitis GWAS Annotation & Prioritisation Pipeline). Methods: We identified a comprehensive list of 103 accessible -

NRF1) Coordinates Changes in the Transcriptional and Chromatin Landscape Affecting Development and Progression of Invasive Breast Cancer
Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 11-7-2018 Decipher Mechanisms by which Nuclear Respiratory Factor One (NRF1) Coordinates Changes in the Transcriptional and Chromatin Landscape Affecting Development and Progression of Invasive Breast Cancer Jairo Ramos [email protected] Follow this and additional works at: https://digitalcommons.fiu.edu/etd Part of the Clinical Epidemiology Commons Recommended Citation Ramos, Jairo, "Decipher Mechanisms by which Nuclear Respiratory Factor One (NRF1) Coordinates Changes in the Transcriptional and Chromatin Landscape Affecting Development and Progression of Invasive Breast Cancer" (2018). FIU Electronic Theses and Dissertations. 3872. https://digitalcommons.fiu.edu/etd/3872 This work is brought to you for free and open access by the University Graduate School at FIU Digital Commons. It has been accepted for inclusion in FIU Electronic Theses and Dissertations by an authorized administrator of FIU Digital Commons. For more information, please contact [email protected]. FLORIDA INTERNATIONAL UNIVERSITY Miami, Florida DECIPHER MECHANISMS BY WHICH NUCLEAR RESPIRATORY FACTOR ONE (NRF1) COORDINATES CHANGES IN THE TRANSCRIPTIONAL AND CHROMATIN LANDSCAPE AFFECTING DEVELOPMENT AND PROGRESSION OF INVASIVE BREAST CANCER A dissertation submitted in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY in PUBLIC HEALTH by Jairo Ramos 2018 To: Dean Tomás R. Guilarte Robert Stempel College of Public Health and Social Work This dissertation, Written by Jairo Ramos, and entitled Decipher Mechanisms by Which Nuclear Respiratory Factor One (NRF1) Coordinates Changes in the Transcriptional and Chromatin Landscape Affecting Development and Progression of Invasive Breast Cancer, having been approved in respect to style and intellectual content, is referred to you for judgment. -

Bach1 Is Critical for the Transformation of Mouse Embryonic Fibroblasts by Rasv12 and Maintains ERK Signaling
Oncogene (2013) 32, 3231–3245 & 2013 Macmillan Publishers Limited All rights reserved 0950-9232/13 www.nature.com/onc ORIGINAL ARTICLE Bach1 is critical for the transformation of mouse embryonic fibroblasts by RasV12 and maintains ERK signaling A Nakanome1,2, A Brydun1,3, M Matsumoto1,4,KOta1, R Funayama5,6, K Nakayama5, M Ono7, K Shiga2, T Kobayashi2 and K Igarashi1,3,6 Reactive oxygen species (ROS), by-products of aerobic respiration, promote genetic instability and contribute to the malignant transformation of cells. Among the genes related to ROS metabolism, Bach1 is a repressor of the oxidative stress response, and a negative regulator of ROS-induced cellular senescence directed by p53 in higher eukaryotes. While ROS are intimately involved in carcinogenesis, it is not clear whether Bach1 is involved in this process. We found that senescent Bach1-deficient mouse embryonic fibroblasts (MEFs) underwent spontaneous immortalization the same as did the wild-type cells. When transduced with constitutively active Ras (H-RasV12), the proliferation and colony formation of these cells in vitro were markedly reduced. When transplanted into athymic nude mice, the growth and vascularization of tumors derived from Bach1-deficient cells were also decreased. Gene expression profiling of the MEFs revealed a new H-RasV12 signature, which was distinct from the previously reported signatures in epithelial tumors, and was partly dependent on Bach1. The Bach1-deficient cells showed diminished phosphorylation of MEK and ERK1/2 in response to H-RasV12, which was consistent with the alterations in the gene expression profile, including phosphatase genes. Finally, Bach1-deficient mice were less susceptible to 4-nitroquinoline-1-oxidide (4-NQO)- induced tongue carcinoma than wild-type mice. -

Identifying Potential Regions of Copy Number Variation for Bipolar Disorder
Microarrays 2014, 3, 52-71; doi:10.3390/microarrays3010052 OPEN ACCESS microarrays ISSN 2076-3905 www.mdpi.com/journal/microarrays Article Identifying Potential Regions of Copy Number Variation for Bipolar Disorder Yi-Hsuan Chen 1, Ru-Band Lu 2, Hung Hung 1,3 and Po-Hsiu Kuo 1,3,* 1 Department of Public Health & Institute of Epidemiology and Preventive Medicine, College of Public Health, National Taiwan University, Taipei 100, Taiwan; E-Mails: [email protected] (Y.-H.C.); [email protected] (H.H.) 2 Department of Psychiatry, College of Medicine & Hospital, National Cheng Kung University, Tainan 704, Taiwan; E-Mail: [email protected] 3 Research Center for Genes, Environment and Human Health, National Taiwan University, Taipei 100, Taiwan * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +886-2-3366-8015; Fax: +886-2-2351-1955. Received: 1 December 2013; in revised form: 10 February 2014 / Accepted: 12 February 2014 / Published: 28 February 2014 Abstract: Bipolar disorder is a complex psychiatric disorder with high heritability, but its genetic determinants are still largely unknown. Copy number variation (CNV) is one of the sources to explain part of the heritability. However, it is a challenge to estimate discrete values of the copy numbers using continuous signals calling from a set of markers, and to simultaneously perform association testing between CNVs and phenotypic outcomes. The goal of the present study is to perform a series of data filtering and analysis procedures using a DNA pooling strategy to identify potential CNV regions that are related to bipolar disorder.