Supplementary Figure 4
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Unrip (STRAP) (NM 007178) Human Tagged ORF Clone Lentiviral Particle Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC209149L3V Unrip (STRAP) (NM_007178) Human Tagged ORF Clone Lentiviral Particle Product data: Product Type: Lentiviral Particles Product Name: Unrip (STRAP) (NM_007178) Human Tagged ORF Clone Lentiviral Particle Symbol: STRAP Synonyms: MAWD; PT-WD; UNRIP Vector: pLenti-C-Myc-DDK-P2A-Puro (PS100092) ACCN: NM_007178 ORF Size: 1050 bp ORF Nucleotide The ORF insert of this clone is exactly the same as(RC209149). Sequence: OTI Disclaimer: The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g. polymorphisms), each with its own valid existence. This clone is substantially in agreement with the reference, but a complete review of all prevailing variants is recommended prior to use. More info OTI Annotation: This clone was engineered to express the complete ORF with an expression tag. Expression varies depending on the nature of the gene. RefSeq: NM_007178.2 RefSeq Size: 1924 bp RefSeq ORF: 1053 bp Locus ID: 11171 UniProt ID: Q9Y3F4 Domains: WD40 Protein Families: Druggable Genome MW: 38.4 kDa This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 Unrip (STRAP) (NM_007178) Human Tagged ORF Clone Lentiviral Particle – RC209149L3V Gene Summary: The SMN complex plays a catalyst role in the assembly of small nuclear ribonucleoproteins (snRNPs), the building blocks of the spliceosome. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
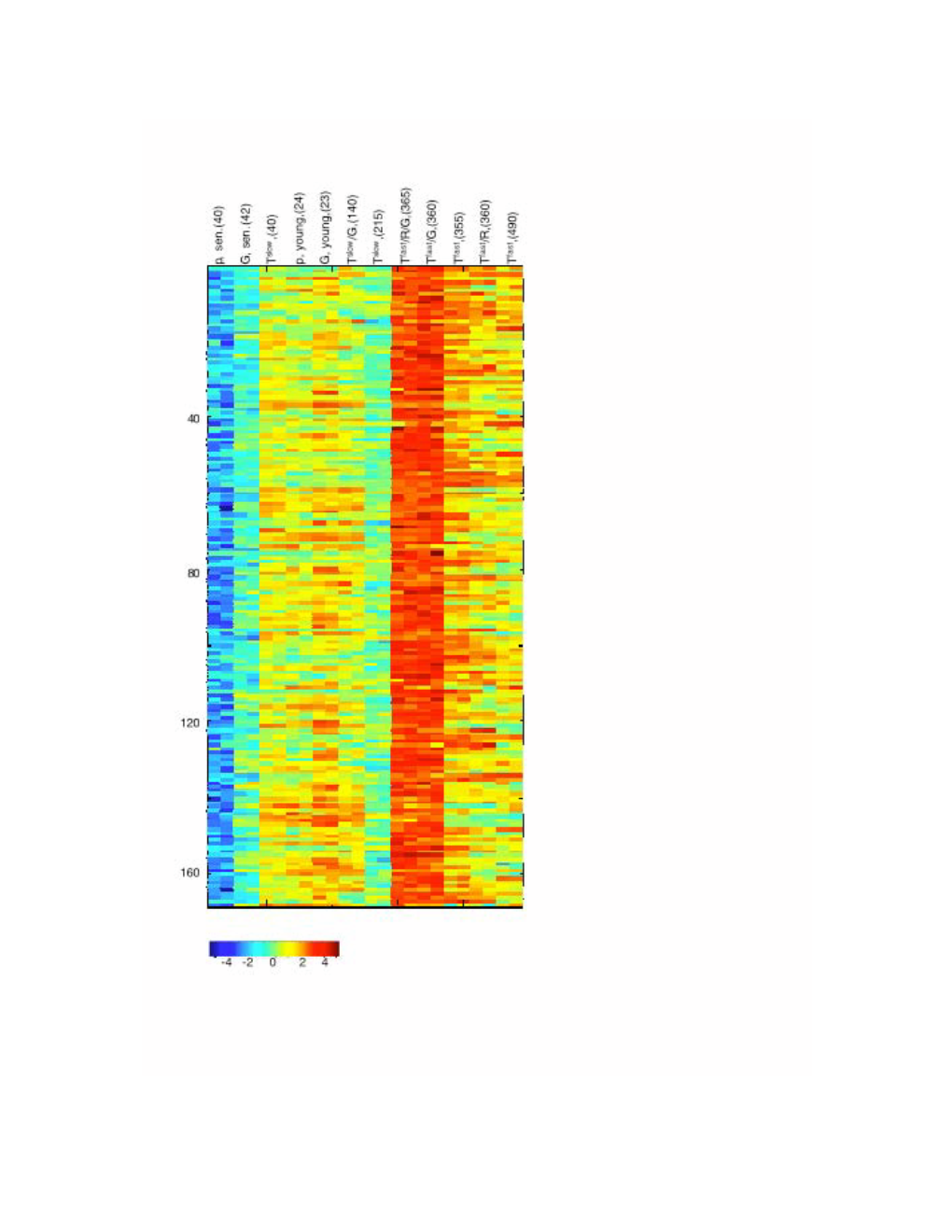
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Zhou Et Al POLQ Inhibitor.Docx
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.23.111658; this version posted May 26, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Polymerase Theta Inhibition Kills Homologous Recombination Deficient Tumors Jia Zhou1, Camille Gelot2, Constantia Pantelidou3, Adam Li1, Hatice Yücel2, Rachel E. Davis4, Anniina Farkkila1, Bose Kochupurakkal1, Aleem Syed5, Geoffrey I. Shapiro3,6, John A. Tainer5, Brian S. J. Blagg4, Raphael Ceccaldi2,7* and Alan D. D’Andrea1,6,7* 1Department of Radiation Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA 02215, USA. 2Inserm U830, PSL Research University, Institut Curie, 75005, Paris, France. 3Department of Medical Oncology, Dana-Farber Cancer Institute and Department of Medicine, Harvard Medical School, Boston, Massachusetts, USA. 4Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, IN 46556, USA. 5Departments of Cancer Biology and of Molecular and Cellular Oncology, University of Texas MD Anderson Cancer Center, Houston, TX 77030, USA. 6Center for DNA Damage and Repair, Dana-Farber Cancer Institute, Boston, MA, USA. 7Co-senior authors. * Co-corresponding authors. * Corresponding authors: Alan D. D’Andrea, M.D. Director, Susan F. Smith Center for Women’s Cancers (SFSCWC) Director, Center for DNA Damage and Repair Dana-Farber Cancer Institute The Fuller-American Cancer Society Professor Harvard Medical School Phone: 617-632-2080 Email: [email protected] Raphael Ceccaldi Institut Curie, 75005, Paris, France Phone: +33 (0)1 56 24 69 49 Email: [email protected] Key Words: Novobiocin, Polymerase theta (POLθ), Homologous Recombination, PARP inhibitor resistance. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

ARTICLE Doi:10.1038/Nature10523
ARTICLE doi:10.1038/nature10523 Spatio-temporal transcriptome of the human brain Hyo Jung Kang1*, Yuka Imamura Kawasawa1*, Feng Cheng1*, Ying Zhu1*, Xuming Xu1*, Mingfeng Li1*, Andre´ M. M. Sousa1,2, Mihovil Pletikos1,3, Kyle A. Meyer1, Goran Sedmak1,3, Tobias Guennel4, Yurae Shin1, Matthew B. Johnson1,Zˇeljka Krsnik1, Simone Mayer1,5, Sofia Fertuzinhos1, Sheila Umlauf6, Steven N. Lisgo7, Alexander Vortmeyer8, Daniel R. Weinberger9, Shrikant Mane6, Thomas M. Hyde9,10, Anita Huttner8, Mark Reimers4, Joel E. Kleinman9 & Nenad Sˇestan1 Brain development and function depend on the precise regulation of gene expression. However, our understanding of the complexity and dynamics of the transcriptome of the human brain is incomplete. Here we report the generation and analysis of exon-level transcriptome and associated genotyping data, representing males and females of different ethnicities, from multiple brain regions and neocortical areas of developing and adult post-mortem human brains. We found that 86 per cent of the genes analysed were expressed, and that 90 per cent of these were differentially regulated at the whole-transcript or exon level across brain regions and/or time. The majority of these spatio-temporal differences were detected before birth, with subsequent increases in the similarity among regional transcriptomes. The transcriptome is organized into distinct co-expression networks, and shows sex-biased gene expression and exon usage. We also profiled trajectories of genes associated with neurobiological categories and diseases, and identified associations between single nucleotide polymorphisms and gene expression. This study provides a comprehensive data set on the human brain transcriptome and insights into the transcriptional foundations of human neurodevelopment. -

Supplement 1 Microarray Studies
EASE Categories Significantly Enriched in vs MG vs vs MGC4-2 Pt1-C vs C4-2 Pt1-C UP-Regulated Genes MG System Gene Category EASE Global MGRWV Pt1-N RWV Pt1-N Score FDR GO Molecular Extracellular matrix cellular construction 0.0008 0 110 genes up- Function Interpro EGF-like domain 0.0009 0 regulated GO Molecular Oxidoreductase activity\ acting on single dono 0.0015 0 Function GO Molecular Calcium ion binding 0.0018 0 Function Interpro Laminin-G domain 0.0025 0 GO Biological Process Cell Adhesion 0.0045 0 Interpro Collagen Triple helix repeat 0.0047 0 KEGG pathway Complement and coagulation cascades 0.0053 0 KEGG pathway Immune System – Homo sapiens 0.0053 0 Interpro Fibrillar collagen C-terminal domain 0.0062 0 Interpro Calcium-binding EGF-like domain 0.0077 0 GO Molecular Cell adhesion molecule activity 0.0105 0 Function EASE Categories Significantly Enriched in Down-Regulated Genes System Gene Category EASE Global Score FDR GO Biological Process Copper ion homeostasis 2.5E-09 0 Interpro Metallothionein 6.1E-08 0 Interpro Vertebrate metallothionein, Family 1 6.1E-08 0 GO Biological Process Transition metal ion homeostasis 8.5E-08 0 GO Biological Process Heavy metal sensitivity/resistance 1.9E-07 0 GO Biological Process Di-, tri-valent inorganic cation homeostasis 6.3E-07 0 GO Biological Process Metal ion homeostasis 6.3E-07 0 GO Biological Process Cation homeostasis 2.1E-06 0 GO Biological Process Cell ion homeostasis 2.1E-06 0 GO Biological Process Ion homeostasis 2.1E-06 0 GO Molecular Helicase activity 2.3E-06 0 Function GO Biological -

Cell Growth-Regulated Expression of Mammalian MCM5 and MCM6 Genes Mediated by the Transcription Factor E2F
Oncogene (1999) 18, 2299 ± 2309 ã 1999 Stockton Press All rights reserved 0950 ± 9232/99 $12.00 http://www.stockton-press.co.uk/onc Cell growth-regulated expression of mammalian MCM5 and MCM6 genes mediated by the transcription factor E2F Kiyoshi Ohtani1, Ritsuko Iwanaga1, Masataka Nakamura*,1, Masa-aki Ikeda2, Norikazu Yabuta3, Hiromichi Tsuruga3 and Hiroshi Nojima3 1Human Gene Sciences Center, Tokyo Medical and Dental University, Tokyo 113-8510, Japan 2Department of Developmental Biology, Graduate School of Dentistry, Tokyo Medical and Dental University, Tokyo 113-8549, Japan; 3Department of Molecular Genetics, Research Institute for Microbial Diseases, Osaka University, Suita 565-0871, Japan Initiation of DNA replication requires the function of family (MCM2-7) that have been identi®ed in yeast, MCM gene products, which participate in ensuring that Xenopus, and human. Mcm proteins seem to regulate DNA replication occurs only once in the cell cycle. the initiation at the replication origin where the loading Expression of all mammalian genes of the MCM family of the proteins onto the origin recognition complex is induced by growth stimulation, unlike yeast, and the (ORC) is regulated by Cdc6 and cyclin-dependent mRNA levels peak at G1/S boundary. In this study, we kinases (Donovan et al., 1997; Tanaka et al., 1997). examined the transcriptional activities of isolated human However, the mechanism(s) by which Mcm proteins MCM gene promoters. Human MCM5 and MCM6 control the initiation of DNA replication remains promoters with mutation in the E2F sites failed in unclear. promoter regulation following serum stimulation and Xenopus Mcm proteins seem to be able to access exogenous E2F expression. -

HNRPH1 (HNRNPH1) (NM 005520) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC201834 HNRPH1 (HNRNPH1) (NM_005520) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: HNRPH1 (HNRNPH1) (NM_005520) Human Tagged ORF Clone Tag: Myc-DDK Symbol: HNRNPH1 Synonyms: hnRNPH; HNRPH; HNRPH1 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 6 HNRPH1 (HNRNPH1) (NM_005520) Human Tagged ORF Clone – RC201834 ORF Nucleotide >RC201834 ORF sequence Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGATGTTGGGCACGGAAGGTGGAGAGGGATTCGTGGTGAAGGTCCGGGGCTTGCCCTGGTCTTGCTCGG CCGATGAAGTGCAGAGGTTTTTTTCTGACTGCAAAATTCAAAATGGGGCTCAAGGTATTCGTTTCATCTA CACCAGAGAAGGCAGACCAAGTGGCGAGGCTTTTGTTGAACTTGAATCAGAAGATGAAGTCAAATTGGCC CTGAAAAAAGACAGAGAAACTATGGGACACAGATATGTTGAAGTATTCAAGTCAAACAACGTTGAAATGG ATTGGGTGTTGAAGCATACTGGTCCAAATAGTCCTGACACGGCCAATGATGGCTTTGTACGGCTTAGAGG ACTTCCCTTTGGATGTAGCAAGGAAGAAATTGTTCAGTTCTTCTCAGGGTTGGAAATCGTGCCAAATGGG ATAACATTGCCGGTGGACTTCCAGGGGAGGAGTACGGGGGAGGCCTTCGTGCAGTTTGCTTCACAGGAAA TAGCTGAAAAGGCTCTAAAGAAACACAAGGAAAGAATAGGGCACAGGTATATTGAAATCTTTAAGAGCAG TAGAGCTGAAGTTAGAACTCATTATGATCCACCACGAAAGCTTATGGCCATGCAGCGGCCAGGTCCTTAT -

Supplementary Table S5. Differentially Expressed Gene Lists of PD-1High CD39+ CD8 Tils According to 4-1BB Expression Compared to PD-1+ CD39- CD8 Tils
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Supplementary Table S5. Differentially expressed gene lists of PD-1high CD39+ CD8 TILs according to 4-1BB expression compared to PD-1+ CD39- CD8 TILs Up- or down- regulated genes in Up- or down- regulated genes Up- or down- regulated genes only PD-1high CD39+ CD8 TILs only in 4-1BBneg PD-1high CD39+ in 4-1BBpos PD-1high CD39+ CD8 compared to PD-1+ CD39- CD8 CD8 TILs compared to PD-1+ TILs compared to PD-1+ CD39- TILs CD39- CD8 TILs CD8 TILs IL7R KLRG1 TNFSF4 ENTPD1 DHRS3 LEF1 ITGA5 MKI67 PZP KLF3 RYR2 SIK1B ANK3 LYST PPP1R3B ETV1 ADAM28 H2AC13 CCR7 GFOD1 RASGRP2 ITGAX MAST4 RAD51AP1 MYO1E CLCF1 NEBL S1PR5 VCL MPP7 MS4A6A PHLDB1 GFPT2 TNF RPL3 SPRY4 VCAM1 B4GALT5 TIPARP TNS3 PDCD1 POLQ AKAP5 IL6ST LY9 PLXND1 PLEKHA1 NEU1 DGKH SPRY2 PLEKHG3 IKZF4 MTX3 PARK7 ATP8B4 SYT11 PTGER4 SORL1 RAB11FIP5 BRCA1 MAP4K3 NCR1 CCR4 S1PR1 PDE8A IFIT2 EPHA4 ARHGEF12 PAICS PELI2 LAT2 GPRASP1 TTN RPLP0 IL4I1 AUTS2 RPS3 CDCA3 NHS LONRF2 CDC42EP3 SLCO3A1 RRM2 ADAMTSL4 INPP5F ARHGAP31 ESCO2 ADRB2 CSF1 WDHD1 GOLIM4 CDK5RAP1 CD69 GLUL HJURP SHC4 GNLY TTC9 HELLS DPP4 IL23A PITPNC1 TOX ARHGEF9 EXO1 SLC4A4 CKAP4 CARMIL3 NHSL2 DZIP3 GINS1 FUT8 UBASH3B CDCA5 PDE7B SOGA1 CDC45 NR3C2 TRIB1 KIF14 TRAF5 LIMS1 PPP1R2C TNFRSF9 KLRC2 POLA1 CD80 ATP10D CDCA8 SETD7 IER2 PATL2 CCDC141 CD84 HSPA6 CYB561 MPHOSPH9 CLSPN KLRC1 PTMS SCML4 ZBTB10 CCL3 CA5B PIP5K1B WNT9A CCNH GEM IL18RAP GGH SARDH B3GNT7 C13orf46 SBF2 IKZF3 ZMAT1 TCF7 NECTIN1 H3C7 FOS PAG1 HECA SLC4A10 SLC35G2 PER1 P2RY1 NFKBIA WDR76 PLAUR KDM1A H1-5 TSHZ2 FAM102B HMMR GPR132 CCRL2 PARP8 A2M ST8SIA1 NUF2 IL5RA RBPMS UBE2T USP53 EEF1A1 PLAC8 LGR6 TMEM123 NEK2 SNAP47 PTGIS SH2B3 P2RY8 S100PBP PLEKHA7 CLNK CRIM1 MGAT5 YBX3 TP53INP1 DTL CFH FEZ1 MYB FRMD4B TSPAN5 STIL ITGA2 GOLGA6L10 MYBL2 AHI1 CAND2 GZMB RBPJ PELI1 HSPA1B KCNK5 GOLGA6L9 TICRR TPRG1 UBE2C AURKA Leem G, et al. -

Growth and Molecular Profile of Lung Cancer Cells Expressing Ectopic LKB1: Down-Regulation of the Phosphatidylinositol 3-Phosphate Kinase/PTEN Pathway1
[CANCER RESEARCH 63, 1382–1388, March 15, 2003] Growth and Molecular Profile of Lung Cancer Cells Expressing Ectopic LKB1: Down-Regulation of the Phosphatidylinositol 3-Phosphate Kinase/PTEN Pathway1 Ana I. Jimenez, Paloma Fernandez, Orlando Dominguez, Ana Dopazo, and Montserrat Sanchez-Cespedes2 Molecular Pathology Program [A. I. J., P. F., M. S-C.], Genomics Unit [O. D.], and Microarray Analysis Unit [A. D.], Spanish National Cancer Center, 28029 Madrid, Spain ABSTRACT the cell cycle in G1 (8, 9). However, the intrinsic mechanism by which LKB1 activity is regulated in cells and how it leads to the suppression Germ-line mutations in LKB1 gene cause the Peutz-Jeghers syndrome of cell growth is still unknown. It has been proposed that growth (PJS), a genetic disease with increased risk of malignancies. Recently, suppression by LKB1 is mediated through p21 in a p53-dependent LKB1-inactivating mutations have been identified in one-third of sporadic lung adenocarcinomas, indicating that LKB1 gene inactivation is critical in mechanism (7). In addition, it has been observed that LKB1 binds to tumors other than those of the PJS syndrome. However, the in vivo brahma-related gene 1 protein (BRG1) and this interaction is required substrates of LKB1 and its role in cancer development have not been for BRG1-induced growth arrest (10). Similar to what happens in the completely elucidated. Here we show that overexpression of wild-type PJS, Lkb1 heterozygous knockout mice show gastrointestinal hamar- LKB1 protein in A549 lung adenocarcinomas cells leads to cell-growth tomatous polyposis and frequent hepatocellular carcinomas (11, 12). suppression. To examine changes in gene expression profiles subsequent to Interestingly, the hamartomas, but not the malignant tumors, arising in exogenous wild-type LKB1 in A549 cells, we used cDNA microarrays. -
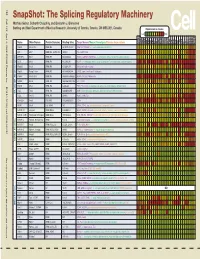
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes.