Admixture Mapping Identifies an Amerindian Ancestry Locus
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Analysis of the Indacaterol-Regulated Transcriptome in Human Airway
Supplemental material to this article can be found at: http://jpet.aspetjournals.org/content/suppl/2018/04/13/jpet.118.249292.DC1 1521-0103/366/1/220–236$35.00 https://doi.org/10.1124/jpet.118.249292 THE JOURNAL OF PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS J Pharmacol Exp Ther 366:220–236, July 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Analysis of the Indacaterol-Regulated Transcriptome in Human Airway Epithelial Cells Implicates Gene Expression Changes in the s Adverse and Therapeutic Effects of b2-Adrenoceptor Agonists Dong Yan, Omar Hamed, Taruna Joshi,1 Mahmoud M. Mostafa, Kyla C. Jamieson, Radhika Joshi, Robert Newton, and Mark A. Giembycz Departments of Physiology and Pharmacology (D.Y., O.H., T.J., K.C.J., R.J., M.A.G.) and Cell Biology and Anatomy (M.M.M., R.N.), Snyder Institute for Chronic Diseases, Cumming School of Medicine, University of Calgary, Calgary, Alberta, Canada Received March 22, 2018; accepted April 11, 2018 Downloaded from ABSTRACT The contribution of gene expression changes to the adverse and activity, and positive regulation of neutrophil chemotaxis. The therapeutic effects of b2-adrenoceptor agonists in asthma was general enriched GO term extracellular space was also associ- investigated using human airway epithelial cells as a therapeu- ated with indacaterol-induced genes, and many of those, in- tically relevant target. Operational model-fitting established that cluding CRISPLD2, DMBT1, GAS1, and SOCS3, have putative jpet.aspetjournals.org the long-acting b2-adrenoceptor agonists (LABA) indacaterol, anti-inflammatory, antibacterial, and/or antiviral activity. Numer- salmeterol, formoterol, and picumeterol were full agonists on ous indacaterol-regulated genes were also induced or repressed BEAS-2B cells transfected with a cAMP-response element in BEAS-2B cells and human primary bronchial epithelial cells by reporter but differed in efficacy (indacaterol $ formoterol . -

ADHD) Gene Networks in Children of Both African American and European American Ancestry
G C A T T A C G G C A T genes Article Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry Yichuan Liu 1 , Xiao Chang 1, Hui-Qi Qu 1 , Lifeng Tian 1 , Joseph Glessner 1, Jingchun Qu 1, Dong Li 1, Haijun Qiu 1, Patrick Sleiman 1,2 and Hakon Hakonarson 1,2,3,* 1 Center for Applied Genomics, Children’s Hospital of Philadelphia, Philadelphia, PA 19104, USA; [email protected] (Y.L.); [email protected] (X.C.); [email protected] (H.-Q.Q.); [email protected] (L.T.); [email protected] (J.G.); [email protected] (J.Q.); [email protected] (D.L.); [email protected] (H.Q.); [email protected] (P.S.) 2 Division of Human Genetics, Department of Pediatrics, The Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104, USA 3 Department of Human Genetics, Children’s Hospital of Philadelphia, Philadelphia, PA 19104, USA * Correspondence: [email protected]; Tel.: +1-267-426-0088 Abstract: Attention-deficit hyperactivity disorder (ADHD) is a neurodevelopmental disorder with poorly understood molecular mechanisms that results in significant impairment in children. In this study, we sought to assess the role of rare recurrent variants in non-European populations and outside of coding regions. We generated whole genome sequence (WGS) data on 875 individuals, Citation: Liu, Y.; Chang, X.; Qu, including 205 ADHD cases and 670 non-ADHD controls. The cases included 116 African Americans H.-Q.; Tian, L.; Glessner, J.; Qu, J.; Li, (AA) and 89 European Americans (EA), and the controls included 408 AA and 262 EA. -

Systematic Elucidation of Neuron-Astrocyte Interaction in Models of Amyotrophic Lateral Sclerosis Using Multi-Modal Integrated Bioinformatics Workflow
ARTICLE https://doi.org/10.1038/s41467-020-19177-y OPEN Systematic elucidation of neuron-astrocyte interaction in models of amyotrophic lateral sclerosis using multi-modal integrated bioinformatics workflow Vartika Mishra et al.# 1234567890():,; Cell-to-cell communications are critical determinants of pathophysiological phenotypes, but methodologies for their systematic elucidation are lacking. Herein, we propose an approach for the Systematic Elucidation and Assessment of Regulatory Cell-to-cell Interaction Net- works (SEARCHIN) to identify ligand-mediated interactions between distinct cellular com- partments. To test this approach, we selected a model of amyotrophic lateral sclerosis (ALS), in which astrocytes expressing mutant superoxide dismutase-1 (mutSOD1) kill wild-type motor neurons (MNs) by an unknown mechanism. Our integrative analysis that combines proteomics and regulatory network analysis infers the interaction between astrocyte-released amyloid precursor protein (APP) and death receptor-6 (DR6) on MNs as the top predicted ligand-receptor pair. The inferred deleterious role of APP and DR6 is confirmed in vitro in models of ALS. Moreover, the DR6 knockdown in MNs of transgenic mutSOD1 mice attenuates the ALS-like phenotype. Our results support the usefulness of integrative, systems biology approach to gain insights into complex neurobiological disease processes as in ALS and posit that the proposed methodology is not restricted to this biological context and could be used in a variety of other non-cell-autonomous communication -

Comprehensive Analysis Reveals Novel Gene Signature in Head and Neck Squamous Cell Carcinoma: Predicting Is Associated with Poor Prognosis in Patients
5892 Original Article Comprehensive analysis reveals novel gene signature in head and neck squamous cell carcinoma: predicting is associated with poor prognosis in patients Yixin Sun1,2#, Quan Zhang1,2#, Lanlin Yao2#, Shuai Wang3, Zhiming Zhang1,2 1Department of Breast Surgery, The First Affiliated Hospital of Xiamen University, School of Medicine, Xiamen University, Xiamen, China; 2School of Medicine, Xiamen University, Xiamen, China; 3State Key Laboratory of Cellular Stress Biology, School of Life Sciences, Xiamen University, Xiamen, China Contributions: (I) Conception and design: Y Sun, Q Zhang; (II) Administrative support: Z Zhang; (III) Provision of study materials or patients: Y Sun, Q Zhang; (IV) Collection and assembly of data: Y Sun, L Yao; (V) Data analysis and interpretation: Y Sun, S Wang; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Zhiming Zhang. Department of Surgery, The First Affiliated Hospital of Xiamen University, Xiamen, China. Email: [email protected]. Background: Head and neck squamous cell carcinoma (HNSC) remains an important public health problem, with classic risk factors being smoking and excessive alcohol consumption and usually has a poor prognosis. Therefore, it is important to explore the underlying mechanisms of tumorigenesis and screen the genes and pathways identified from such studies and their role in pathogenesis. The purpose of this study was to identify genes or signal pathways associated with the development of HNSC. Methods: In this study, we downloaded gene expression profiles of GSE53819 from the Gene Expression Omnibus (GEO) database, including 18 HNSC tissues and 18 normal tissues. -

Identifying Rare Genetic Variation in Obsessive-Compulsive Disorder
Yale University EliScholar – A Digital Platform for Scholarly Publishing at Yale Yale Medicine Thesis Digital Library School of Medicine January 2020 Identifying Rare Genetic Variation In Obsessive-Compulsive Disorder Sarah Abdallah Follow this and additional works at: https://elischolar.library.yale.edu/ymtdl Recommended Citation Abdallah, Sarah, "Identifying Rare Genetic Variation In Obsessive-Compulsive Disorder" (2020). Yale Medicine Thesis Digital Library. 3876. https://elischolar.library.yale.edu/ymtdl/3876 This Open Access Thesis is brought to you for free and open access by the School of Medicine at EliScholar – A Digital Platform for Scholarly Publishing at Yale. It has been accepted for inclusion in Yale Medicine Thesis Digital Library by an authorized administrator of EliScholar – A Digital Platform for Scholarly Publishing at Yale. For more information, please contact [email protected]. Identifying Rare Genetic Variation in Obsessive-Compulsive Disorder A Thesis Submitted to the Yale University School of Medicine in Partial Fulfillment of the Requirements for the Degree of Doctor of Medicine by Sarah Barbara Abdallah 2020 ABSTRACT IDENTIFYING RARE GENETIC VARIATION IN OBSESSIVE-COMPULSIVE DISORDER Sarah B. Abdallah, Carolina Cappi, Emily Olfson, and Thomas V. Fernandez. Child Study Center, Yale University School of Medicine, New Haven, CT Obsessive-compulsive disorder (OCD) is a neuropsychiatric developmental disorder with known heritability (estimates ranging from 27%-80%) but poorly understood etiology. Current treatments are not fully effective in addressing chronic functional impairments and distress caused by the disorder, providing an impetus to study the genetic basis of OCD in the hopes of identifying new therapeutic targets. We previously demonstrated a significant contribution to OCD risk from likely damaging de novo germline DNA sequence variants, which arise spontaneously in the parental germ cells or zygote instead of being inherited from a parent, and we successfully used these identified variants to implicate new OCD risk genes. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -
Genome Informatics
Joint Cold Spring Harbor Laboratory/Wellcome Trust Conference GENOME INFORMATICS September 15–September 19, 2010 View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Cold Spring Harbor Laboratory Institutional Repository Joint Cold Spring Harbor Laboratory/Wellcome Trust Conference GENOME INFORMATICS September 15–September 19, 2010 Arranged by Inanc Birol, BC Cancer Agency, Canada Michele Clamp, BioTeam, Inc. James Kent, University of California, Santa Cruz, USA SCHEDULE AT A GLANCE Wednesday 15th September 2010 17.00-17.30 Registration – finger buffet dinner served from 17.30-19.30 19.30-20:50 Session 1: Epigenomics and Gene Regulation 20.50-21.10 Break 21.10-22.30 Session 1, continued Thursday 16th September 2010 07.30-09.00 Breakfast 09.00-10.20 Session 2: Population and Statistical Genomics 10.20-10:40 Morning Coffee 10:40-12:00 Session 2, continued 12.00-14.00 Lunch 14.00-15.20 Session 3: Environmental and Medical Genomics 15.20-15.40 Break 15.40-17.00 Session 3, continued 17.00-19.00 Poster Session I and Drinks Reception 19.00-21.00 Dinner Friday 17th September 2010 07.30-09.00 Breakfast 09.00-10.20 Session 4: Databases, Data Mining, Visualization and Curation 10.20-10.40 Morning Coffee 10.40-12.00 Session 4, continued 12.00-14.00 Lunch 14.00-16.00 Free afternoon 16.00-17.00 Keynote Speaker: Alex Bateman 17.00-19.00 Poster Session II and Drinks Reception 19.00-21.00 Dinner Saturday 18th September 2010 07.30-09.00 Breakfast 09.00-10.20 Session 5: Sequencing Pipelines and Assembly 10.20-10.40 -

Microarray Bioinformatics and Its Applications to Clinical Research
Microarray Bioinformatics and Its Applications to Clinical Research A dissertation presented to the School of Electrical and Information Engineering of the University of Sydney in fulfillment of the requirements for the degree of Doctor of Philosophy i JLI ··_L - -> ...·. ...,. by Ilene Y. Chen Acknowledgment This thesis owes its existence to the mercy, support and inspiration of many people. In the first place, having suffering from adult-onset asthma, interstitial cystitis and cold agglutinin disease, I would like to express my deepest sense of appreciation and gratitude to Professors Hong Yan and David Levy for harbouring me these last three years and providing me a place at the University of Sydney to pursue a very meaningful course of research. I am also indebted to Dr. Craig Jin, who has been a source of enthusiasm and encouragement on my research over many years. In the second place, for contexts concerning biological and medical aspects covered in this thesis, I am very indebted to Dr. Ling-Hong Tseng, Dr. Shian-Sehn Shie, Dr. Wen-Hung Chung and Professor Chyi-Long Lee at Change Gung Memorial Hospital and University of Chang Gung School of Medicine (Taoyuan, Taiwan) as well as Professor Keith Lloyd at University of Alabama School of Medicine (AL, USA). All of them have contributed substantially to this work. In the third place, I would like to thank Mrs. Inge Rogers and Mr. William Ballinger for their helpful comments and suggestions for the writing of my papers and thesis. In the fourth place, I would like to thank my swim coach, Hirota Homma. -
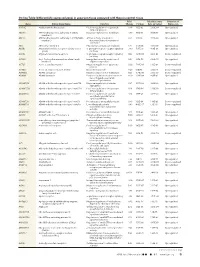
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Transcriptome Profiling Reveals the Complexity of Pirfenidone Effects in IPF
ERJ Express. Published on August 30, 2018 as doi: 10.1183/13993003.00564-2018 Early View Original article Transcriptome profiling reveals the complexity of pirfenidone effects in IPF Grazyna Kwapiszewska, Anna Gungl, Jochen Wilhelm, Leigh M. Marsh, Helene Thekkekara Puthenparampil, Katharina Sinn, Miroslava Didiasova, Walter Klepetko, Djuro Kosanovic, Ralph T. Schermuly, Lukasz Wujak, Benjamin Weiss, Liliana Schaefer, Marc Schneider, Michael Kreuter, Andrea Olschewski, Werner Seeger, Horst Olschewski, Malgorzata Wygrecka Please cite this article as: Kwapiszewska G, Gungl A, Wilhelm J, et al. Transcriptome profiling reveals the complexity of pirfenidone effects in IPF. Eur Respir J 2018; in press (https://doi.org/10.1183/13993003.00564-2018). This manuscript has recently been accepted for publication in the European Respiratory Journal. It is published here in its accepted form prior to copyediting and typesetting by our production team. After these production processes are complete and the authors have approved the resulting proofs, the article will move to the latest issue of the ERJ online. Copyright ©ERS 2018 Copyright 2018 by the European Respiratory Society. Transcriptome profiling reveals the complexity of pirfenidone effects in IPF Grazyna Kwapiszewska1,2, Anna Gungl2, Jochen Wilhelm3†, Leigh M. Marsh1, Helene Thekkekara Puthenparampil1, Katharina Sinn4, Miroslava Didiasova5, Walter Klepetko4, Djuro Kosanovic3, Ralph T. Schermuly3†, Lukasz Wujak5, Benjamin Weiss6, Liliana Schaefer7, Marc Schneider8†, Michael Kreuter8†, Andrea Olschewski1, -

Mecp2 Knockdown Reveals DNA Methylation-Independent Gene Repression of Target Genes in Living Cells and a Bias in the Cellular Location of Target Gene Products
Genes Genet. Syst. (2008) 83, p. 199–208 MeCP2 knockdown reveals DNA methylation-independent gene repression of target genes in living cells and a bias in the cellular location of target gene products Shinya Yakabe1,2, Hidenobu Soejima1*, Hitomi Yatsuki1, Hirotaka Tominaga3, Wei Zhao4, Ken Higashimoto1, Keiichiro Joh1, Shinichi Kudo5, Kohji Miyazaki2 and Tsunehiro Mukai6 1Division of Molecular Genetics and Epigenetics, Department of Biomolecular Sciences, Faculty of Medicine, Saga University, 5-1-1 Nabeshima, Saga 849-8501, Japan 2Division of General Surgery, Department of Surgery, Faculty of Medicine, Saga University, 5-1-1 Nabeshima, Saga 849-8501, Japan 3Section of Clinical Cooperation System, Center for Comprehensive Community Medicine, Faculty of Medicine, Saga University, 5-1-1 Nabeshima, Saga 849-8501, Japan 4Department of Cardiovascular Medicine, Shanghai Shuguang Hospital Affiliated with Shanghai University of T.C.M., No. 185 Pu An Road, Shanghai 200021, R. P. China 5Hokkaido Institute of Public Health, Sapporo 060-0819, Japan 6Saga University, 1 Honjo, Saga, Japan (Received 9 February 2008, accepted 21 February 2008) MeCP2, a methyl-CpG binding domain (MBD) protein, is known to bind to meth- ylated CpG sites via a conserved MBD, leading to transcriptional repression. However, studies in cell-free system for gene repression and MeCP2 binding have suggested that DNA methylation-independent repression also occurs in living cells. It has been difficult to characterize the target genes of MeCP2 because a limited number have been identified to date. In this context, we screened for MeCP2 target genes using knockdown (KD) experiments combined with microar- ray gene expression analyses. Of the 49 genes that showed a more than three- fold increase in expression in two independent KD experiments conducted with different siRNA sets, unexpectedly, half (24 genes) did not contain promoter CpG islands (CGIs). -

Transposon Mutagenesis Identifies Genetic Drivers of Brafv600e Melanoma
ARTICLES Transposon mutagenesis identifies genetic drivers of BrafV600E melanoma Michael B Mann1,2, Michael A Black3, Devin J Jones1, Jerrold M Ward2,12, Christopher Chin Kuan Yew2,12, Justin Y Newberg1, Adam J Dupuy4, Alistair G Rust5,12, Marcus W Bosenberg6,7, Martin McMahon8,9, Cristin G Print10,11, Neal G Copeland1,2,13 & Nancy A Jenkins1,2,13 Although nearly half of human melanomas harbor oncogenic BRAFV600E mutations, the genetic events that cooperate with these mutations to drive melanogenesis are still largely unknown. Here we show that Sleeping Beauty (SB) transposon-mediated mutagenesis drives melanoma progression in BrafV600E mutant mice and identify 1,232 recurrently mutated candidate cancer genes (CCGs) from 70 SB-driven melanomas. CCGs are enriched in Wnt, PI3K, MAPK and netrin signaling pathway components and are more highly connected to one another than predicted by chance, indicating that SB targets cooperative genetic networks in melanoma. Human orthologs of >500 CCGs are enriched for mutations in human melanoma or showed statistically significant clinical associations between RNA abundance and survival of patients with metastatic melanoma. We also functionally validate CEP350 as a new tumor-suppressor gene in human melanoma. SB mutagenesis has thus helped to catalog the cooperative molecular mechanisms driving BRAFV600E melanoma and discover new genes with potential clinical importance in human melanoma. Substantial sun exposure and numerous genetic factors, including including BrafV600E, recapitulate the genetic and histological hallmarks skin type and family history, are the most important melanoma risk of human melanoma. In these models, increased MEK-ERK signaling factors. Familial melanoma, which accounts for <10% of cases, is asso- initiates clonal expansion of melanocytes, which is limited by oncogene- ciated with mutations in CDKN2A1, MITF2 and POT1 (refs.