Diversity of Xylanolytic Bacteria Isolated from Thai Sources
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Comparative Genomic Analysis of Three Pseudomonas
microorganisms Article Comparative Genomic Analysis of Three Pseudomonas Species Isolated from the Eastern Oyster (Crassostrea virginica) Tissues, Mantle Fluid, and the Overlying Estuarine Water Column Ashish Pathak 1, Paul Stothard 2 and Ashvini Chauhan 1,* 1 Environmental Biotechnology Laboratory, School of the Environment, 1515 S. Martin Luther King Jr. Blvd., Suite 305B, FSH Science Research Center, Florida A&M University, Tallahassee, FL 32307, USA; [email protected] 2 Department of Agricultural, Food and Nutritional Science, University of Alberta, Edmonton, AB T6G2P5, Canada; [email protected] * Correspondence: [email protected]; Tel.: +1-850-412-5119; Fax: +1-850-561-2248 Abstract: The eastern oysters serve as important keystone species in the United States, especially in the Gulf of Mexico estuarine waters, and at the same time, provide unparalleled economic, ecological, environmental, and cultural services. One ecosystem service that has garnered recent attention is the ability of oysters to sequester impurities and nutrients, such as nitrogen (N), from the estuarine water that feeds them, via their exceptional filtration mechanism coupled with microbially-mediated denitrification processes. It is the oyster-associated microbiomes that essentially provide these myriads of ecological functions, yet not much is known on these microbiota at the genomic scale, especially from warm temperate and tropical water habitats. Among the suite of bacterial genera that appear to interplay with the oyster host species, pseudomonads deserve further assessment because Citation: Pathak, A.; Stothard, P.; of their immense metabolic and ecological potential. To obtain a comprehensive understanding on Chauhan, A. Comparative Genomic this aspect, we previously reported on the isolation and preliminary genomic characterization of Analysis of Three Pseudomonas Species three Pseudomonas species isolated from minced oyster tissue (P. -

Corynebacterium Sp.|NML98-0116
1 Limnochorda_pilosa~GCF_001544015.1@NZ_AP014924=Bacteria-Firmicutes-Limnochordia-Limnochordales-Limnochordaceae-Limnochorda-Limnochorda_pilosa 0,9635 Ammonifex_degensii|KC4~GCF_000024605.1@NC_013385=Bacteria-Firmicutes-Clostridia-Thermoanaerobacterales-Thermoanaerobacteraceae-Ammonifex-Ammonifex_degensii 0,985 Symbiobacterium_thermophilum|IAM14863~GCF_000009905.1@NC_006177=Bacteria-Firmicutes-Clostridia-Clostridiales-Symbiobacteriaceae-Symbiobacterium-Symbiobacterium_thermophilum Varibaculum_timonense~GCF_900169515.1@NZ_LT827020=Bacteria-Actinobacteria-Actinobacteria-Actinomycetales-Actinomycetaceae-Varibaculum-Varibaculum_timonense 1 Rubrobacter_aplysinae~GCF_001029505.1@NZ_LEKH01000003=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_aplysinae 0,975 Rubrobacter_xylanophilus|DSM9941~GCF_000014185.1@NC_008148=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_xylanophilus 1 Rubrobacter_radiotolerans~GCF_000661895.1@NZ_CP007514=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_radiotolerans Actinobacteria_bacterium_rbg_16_64_13~GCA_001768675.1@MELN01000053=Bacteria-Actinobacteria-unknown_class-unknown_order-unknown_family-unknown_genus-Actinobacteria_bacterium_rbg_16_64_13 1 Actinobacteria_bacterium_13_2_20cm_68_14~GCA_001914705.1@MNDB01000040=Bacteria-Actinobacteria-unknown_class-unknown_order-unknown_family-unknown_genus-Actinobacteria_bacterium_13_2_20cm_68_14 1 0,9803 Thermoleophilum_album~GCF_900108055.1@NZ_FNWJ01000001=Bacteria-Actinobacteria-Thermoleophilia-Thermoleophilales-Thermoleophilaceae-Thermoleophilum-Thermoleophilum_album -

Report on 31 Unrecorded Bacterial Species in Korea That Belong to the Phylum Actinobacteria
Journal of Species Research 5(1):113, 2016 Report on 31 unrecorded bacterial species in Korea that belong to the phylum Actinobacteria JungHye Choi1, JuHee Cha1, JinWoo Bae2, JangCheon Cho3, Jongsik Chun4, WanTaek Im5, Kwang Yeop Jahng6, Che Ok Jeon7, Kiseong Joh8, Seung Bum Kim9, Chi Nam Seong10, JungHoon Yoon11 and ChangJun Cha1,* 1Department of Systems Biotechnology, Chung-Ang University, Anseong 17546, Korea 2Department of Biology, Kyung Hee University, Seoul 02447, Korea 3Department of Biological Sciences, Inha University, Incheon 22212, Korea 4School of Biological Sciences, Seoul National University, Seoul 08826, Korea 5Department of Biotechnology, Hankyong National University, Anseong 17579, Korea 6Department of Life Sciences, Chonbuk National University, Jeonju-si 54896, Korea 7Department of Life Science, Chung-Ang University, Seoul 06974, Korea 8Department of Bioscience and Biotechnology, Hankuk University of Foreign Studies, Gyeonggi 17035, Korea 9Department of Microbiology, Chungnam National University, Daejeon 34134, Korea 10Department of Biology, Sunchon National University, Suncheon 57922, Korea 11Department of Food Science and Biotechnology, Sungkyunkwan University, Suwon 16419, Korea *Correspondent: [email protected] To discover and characterize indigenous species in Korea, a total of 31 bacterial strains that belong to the phylum Actinobacteria were isolated from various niches in Korea. Each strain showed the high sequence similarity (>99.1%) with the closest bacterial species, forming a robust phylogenetic clade. These strains have not been previously recorded in Korea. According to the recently updated taxonomy of the phylum Actinobacteria based upon 16S rRNA trees, we report 25 genera of 13 families within 5 orders of the class Actinobacteria as actinobacterial species found in Korea. -

Rapport Nederlands
Moleculaire detectie van bacteriën in dekaarde Dr. J.J.P. Baars & dr. G. Straatsma Plant Research International B.V., Wageningen December 2007 Rapport nummer 2007-10 © 2007 Wageningen, Plant Research International B.V. Alle rechten voorbehouden. Niets uit deze uitgave mag worden verveelvoudigd, opgeslagen in een geautomatiseerd gegevensbestand, of openbaar gemaakt, in enige vorm of op enige wijze, hetzij elektronisch, mechanisch, door fotokopieën, opnamen of enige andere manier zonder voorafgaande schriftelijke toestemming van Plant Research International B.V. Exemplaren van dit rapport kunnen bij de (eerste) auteur worden besteld. Bij toezending wordt een factuur toegevoegd; de kosten (incl. verzend- en administratiekosten) bedragen € 50 per exemplaar. Plant Research International B.V. Adres : Droevendaalsesteeg 1, Wageningen : Postbus 16, 6700 AA Wageningen Tel. : 0317 - 47 70 00 Fax : 0317 - 41 80 94 E-mail : [email protected] Internet : www.pri.wur.nl Inhoudsopgave pagina 1. Samenvatting 1 2. Inleiding 3 3. Methodiek 8 Algemene werkwijze 8 Bestudeerde monsters 8 Monsters uit praktijkteelten 8 Monsters uit proefteelten 9 Alternatieve analyse m.b.v. DGGE 10 Vaststellen van verschillen tussen de bacterie-gemeenschappen op myceliumstrengen en in de omringende dekaarde. 11 4. Resultaten 13 Monsters uit praktijkteelten 13 Monsters uit proefteelten 16 Alternatieve analyse m.b.v. DGGE 23 Vaststellen van verschillen tussen de bacterie-gemeenschappen op myceliumstrengen en in de omringende dekaarde. 25 5. Discussie 28 6. Conclusies 33 7. Suggesties voor verder onderzoek 35 8. Gebruikte literatuur. 37 Bijlage I. Bacteriesoorten geïsoleerd uit dekaarde en van mycelium uit commerciële teelten I-1 Bijlage II. Bacteriesoorten geïsoleerd uit dekaarde en van mycelium uit experimentele teelten II-1 1 1. -

Succession of Lignocellulolytic Bacterial Consortia Bred Anaerobically from Lake Sediment
bs_bs_banner Succession of lignocellulolytic bacterial consortia bred anaerobically from lake sediment Elisa Korenblum,*† Diego Javier Jimenez and A total of 160 strains was isolated from the enrich- Jan Dirk van Elsas ments. Most of the strains tested (78%) were able to Department of Microbial Ecology,Groningen Institute for grow anaerobically on carboxymethyl cellulose and Evolutionary Life Sciences,University of Groningen, xylan. The final consortia yield attractive biological Groningen,The Netherlands. tools for the depolymerization of recalcitrant ligno- cellulosic materials and are proposed for the produc- tion of precursors of biofuels. Summary Anaerobic bacteria degrade lignocellulose in various Introduction anoxic and organically rich environments, often in a syntrophic process. Anaerobic enrichments of bacte- Lignocellulose is naturally depolymerized by enzymes of rial communities on a recalcitrant lignocellulose microbial communities that develop in soil as well as in source were studied combining polymerase chain sediments of lakes and rivers (van der Lelie et al., reaction–denaturing gradient gel electrophoresis, 2012). Sediments in organically rich environments are amplicon sequencing of the 16S rRNA gene and cul- usually waterlogged and anoxic, already within a cen- turing. Three consortia were constructed using the timetre or less of the sediment water interface. There- microbiota of lake sediment as the starting inoculum fore, much of the organic detritus is probably degraded and untreated switchgrass (Panicum virgatum) (acid by anaerobic processes in such systems (Benner et al., or heat) or treated (with either acid or heat) as the 1984). Whereas fungi are well-known lignocellulose sole source of carbonaceous compounds. Addition- degraders in toxic conditions, due to their oxidative ally, nitrate was used in order to limit sulfate reduc- enzymes (Wang et al., 2013), in anoxic environments tion and methanogenesis. -

Thermophilic and Alkaliphilic Actinobacteria: Biology and Potential Applications
REVIEW published: 25 September 2015 doi: 10.3389/fmicb.2015.01014 Thermophilic and alkaliphilic Actinobacteria: biology and potential applications L. Shivlata and Tulasi Satyanarayana * Department of Microbiology, University of Delhi, New Delhi, India Microbes belonging to the phylum Actinobacteria are prolific sources of antibiotics, clinically useful bioactive compounds and industrially important enzymes. The focus of the current review is on the diversity and potential applications of thermophilic and alkaliphilic actinobacteria, which are highly diverse in their taxonomy and morphology with a variety of adaptations for surviving and thriving in hostile environments. The specific metabolic pathways in these actinobacteria are activated for elaborating pharmaceutically, agriculturally, and biotechnologically relevant biomolecules/bioactive Edited by: compounds, which find multifarious applications. Wen-Jun Li, Sun Yat-Sen University, China Keywords: Actinobacteria, thermophiles, alkaliphiles, polyextremophiles, bioactive compounds, enzymes Reviewed by: Erika Kothe, Friedrich Schiller University Jena, Introduction Germany Hongchen Jiang, The phylum Actinobacteria is one of the most dominant phyla in the bacteria domain (Ventura Miami University, USA et al., 2007), that comprises a heterogeneous Gram-positive and Gram-variable genera. The Qiuyuan Huang, phylum also includes a few Gram-negative species such as Thermoleophilum sp. (Zarilla and Miami University, USA Perry, 1986), Gardenerella vaginalis (Gardner and Dukes, 1955), Saccharomonospora -

Diversity of Cultivated Aerobic Poly-Hydrolytic Bacteria in Saline Alkaline Soils
Diversity of cultivated aerobic poly-hydrolytic bacteria in saline alkaline soils Dimitry Y. Sorokin1,2, Tatiana V. Kolganova3, Tatiana V. Khijniak1, Brian E. Jones4 and Ilya V. Kublanov1,5 1 Winogradsky Institute of Microbiology, Research Centre of Biotechnology, Russian Academy of Sciences, Moscow, Russia 2 Department of Biotechnology, Delft University of Technology, Delft, Netherlands 3 Institute of Bioengineering, Research Centre of Biotechnology, Russian Academy of Sciences, Moscow, Russia 4 DuPont Industrial Biosciences/Genencor International BV, Leiden, Netherlands 5 Immanuel Kant Baltic Federal University, Kaliningrad, Russia ABSTRACT Alkaline saline soils, known also as ``soda solonchaks'', represent a natural soda habitat which differs from soda lake sediments by higher aeration and lower humidity. The microbiology of soda soils, in contrast to the more intensively studied soda lakes, remains poorly explored. In this work we investigate the diversity of culturable aerobic haloalkalitolerant bacteria with various hydrolytic activities from soda soils at different locations in Central Asia, Africa, and North America. In total, 179 pure cultures were obtained by using media with various polymers at pH 10 and 0.6 M total Na+. According to the 16S rRNA gene sequence analysis, most of the isolates belonged to Firmicutes and Actinobacteria. Most isolates possessed multiple hydrolytic activities, including endoglucanase, xylanase, amylase and protease. The pH profiling of selected representatives of actinobacteria and endospore-forming bacteria showed, that the former were facultative alkaliphiles, while the latter were mostly obligate alkaliphiles. The hydrolases of selected representatives from both groups were active at a broad pH range from six to 11. Overall, this work demonstrates the presence of a rich hydrolytic Submitted 9 June 2017 bacterial community in soda soils which might be explored further for production of Accepted 21 August 2017 haloalkalistable hydrolases. -
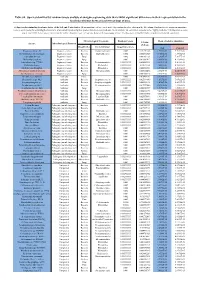
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp. -

Of the 16 Strains Obtained from Salty Soil in Mongolia, Six Strains Were
African Journal of Microbiology Research Vol. 7(4), pp. 298-308, 22 January, 2013 Available online at http://www.academicjournals.org/AJMR DOI: 10.5897/AJMR12.498 ISSN 1996 0808 ©2013 Academic Journals Full Length Research Paper Isolation, classification, phylogenetic analysis and scanning electron microscopy of halophilic, halotolerant and alkaliphilic actinomycetes isolated from hypersaline soil Ismet Ara1,2*, D. Daram4, T. Baljinova4, H. Yamamura5, W. N. Hozzein3, M. A. Bakir1, M. Suto2 and K. Ando2 1Department of Botany and Microbiology, College of Science, King Saud University, P. O. Box 22452, Riyadh-11495, Kingdom of Saudi Arabia. 2Biotechnology Development Center, Department of Biotechnology (DOB), National Institute of Technology and Evaluation (NITE), Kazusakamatari 2-5-8, Kisarazu, Chiba, 292-0818, Japan. 3Bioproducts Research Chair (BRC), College of Science, King Saud University. 4Division of Applied Biological Sciences, Interdisciplinary Graduate School of Medicine and Engineering, University of Yamanashi, Takeda-4, Kofu 400-8511, Japan. Accepted 9 January, 2013 Actinomycetes were isolated by plating of serially diluted samples onto humic acid-vitamin agar prepared with or without NaCl and characterized by physiological and phylogenetical studies. A total of 16 strains were isolated from hypersaline soil in Mongolia. All strains showed alkaliphilic characteristics and were able to abundantly grow in media with pH 9.0. Six strains were halotolerant actinomycetes (0 to 12.0% NaCl) and 10 strains were moderately halophiles (3.0 to 12.0% NaCl). Most strains required moderately high salt (5.0 to 10.0%) for optimal growth but were able to grow at lower NaCl concentrations. Among the 16 strains, 5 were able to grow at 45°C. -

A Novel Approach to the Discovery of Natural Products from Actinobacteria Rahmy Tawfik University of South Florida, [email protected]
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School 3-24-2017 A Novel Approach to the Discovery of Natural Products From Actinobacteria Rahmy Tawfik University of South Florida, [email protected] Follow this and additional works at: http://scholarcommons.usf.edu/etd Part of the Microbiology Commons Scholar Commons Citation Tawfik, Rahmy, "A Novel Approach to the Discovery of Natural Products From Actinobacteria" (2017). Graduate Theses and Dissertations. http://scholarcommons.usf.edu/etd/6766 This Thesis is brought to you for free and open access by the Graduate School at Scholar Commons. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Scholar Commons. For more information, please contact [email protected]. A Novel Approach to the Discovery of Natural Products From Actinobacteria by Rahmy Tawfik A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science Department of Cell Biology, Microbiology & Molecular Biology College of Arts and Sciences University of South Florida Major Professor: Lindsey N. Shaw, Ph.D. Edward Turos, Ph.D. Bill J. Baker, Ph.D. Date of Approval: March 22, 2017 Keywords: Secondary Metabolism, Soil, HPLC, Mass Spectrometry, Antibiotic Copyright © 2017, Rahmy Tawfik Acknowledgements I would like to express my gratitude to the people who have helped and supported me throughout this degree for both scientific and personal. First, I would like to thank my mentor and advisor, Dr. Lindsey Shaw. Although my academics were lacking prior to entering graduate school, you were willing to look beyond my shortcomings and focus on my strengths. -

Appendix 1. Validly Published Names, Conserved and Rejected Names, And
Appendix 1. Validly published names, conserved and rejected names, and taxonomic opinions cited in the International Journal of Systematic and Evolutionary Microbiology since publication of Volume 2 of the Second Edition of the Systematics* JEAN P. EUZÉBY New phyla Alteromonadales Bowman and McMeekin 2005, 2235VP – Valid publication: Validation List no. 106 – Effective publication: Names above the rank of class are not covered by the Rules of Bowman and McMeekin (2005) the Bacteriological Code (1990 Revision), and the names of phyla are not to be regarded as having been validly published. These Anaerolineales Yamada et al. 2006, 1338VP names are listed for completeness. Bdellovibrionales Garrity et al. 2006, 1VP – Valid publication: Lentisphaerae Cho et al. 2004 – Valid publication: Validation List Validation List no. 107 – Effective publication: Garrity et al. no. 98 – Effective publication: J.C. Cho et al. (2004) (2005xxxvi) Proteobacteria Garrity et al. 2005 – Valid publication: Validation Burkholderiales Garrity et al. 2006, 1VP – Valid publication: Vali- List no. 106 – Effective publication: Garrity et al. (2005i) dation List no. 107 – Effective publication: Garrity et al. (2005xxiii) New classes Caldilineales Yamada et al. 2006, 1339VP VP Alphaproteobacteria Garrity et al. 2006, 1 – Valid publication: Campylobacterales Garrity et al. 2006, 1VP – Valid publication: Validation List no. 107 – Effective publication: Garrity et al. Validation List no. 107 – Effective publication: Garrity et al. (2005xv) (2005xxxixi) VP Anaerolineae Yamada et al. 2006, 1336 Cardiobacteriales Garrity et al. 2005, 2235VP – Valid publica- Betaproteobacteria Garrity et al. 2006, 1VP – Valid publication: tion: Validation List no. 106 – Effective publication: Garrity Validation List no. 107 – Effective publication: Garrity et al. -

Manuscript with Highlighted Changes 06032020
1 Diversity and distribution of Nitrogen Fixation Genes in the Oxygen Minimum Zones of the 2 World Oceans 3 1Amal Jayakumar and 1Bess B. Ward 4 5 1Department of Geosciences 6 Princeton University 7 Princeton, NJ 08544 8 9 Abstract 10 Diversity and community composition of nitrogen fixing microbes in the three main oxygen 11 minimum zones (OMZs) of the world ocean were investigated using operational taxonomic unit 12 (OTU) analysis of nifH clone libraries. Representatives of the all four main clusters of nifH genes 13 were detected. Cluster I sequences were most diverse in the surface waters and the most abundant 14 OTUs were affiliated with Alpha- and Gammaproteobacteria. Cluster II, III, IV assemblages were 15 most diverse at oxygen depleted depths and none of the sequences were closely related to sequences 16 from cultivated organisms. The OTUs were biogeographically distinct for the most part – there was 17 little overlap among regions, between depths or between cDNA and DNA. Only a few 18 cyanobacterial sequences were detected. The prevalence and diversity of microbes that harbour nifH 19 genes in the OMZ regions, where low rates of N fixation are reported, remains an enigma. 20 21 Introduction 22 Nitrogen fixation is the biological process that introduces new biologically available 23 nitrogen into the ocean, and thus constrains the overall productivity of large regions of the ocean 24 where N is limiting to primary production. The most abundant and most important diazotrophs 25 in the ocean are cyanobacteria, members of the filamentous genus Trichodesmium and several 1 26 unicellular genera, including Chrocosphaera sp.