Investigating DNA Junction Structure and Dynamics Using a Coarse-Grained Implicit Ion Model for DNA
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
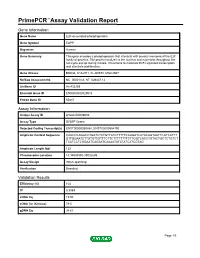
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name E2F-associated phosphoprotein Gene Symbol EAPP Organism Human Gene Summary This gene encodes a phosphoprotein that interacts with several members of the E2F family of proteins. The protein localizes to the nucleus and is present throughout the cell cycle except during mitosis. It functions to modulate E2F-regulated transcription and stimulate proliferation. Gene Aliases BM036, C14orf11, FLJ20578, MGC4957 RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.433269 Ensembl Gene ID ENSG00000129518 Entrez Gene ID 55837 Assay Information Unique Assay ID qHsaCID0008053 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000250454, ENST00000554792 Amplicon Context Sequence CAACCCAGGCCTGATCTCTGTTATCTTTTTCAGGATCATACAGTAATTCGTCATTT GTTGGAATCTTGTGTTGTTTCTTCTTTTTTTTCTTGGTCACCTGTACTGCTCTGTCT TCATCCTCGGAATCAGAATCAAAATATATATCATCGTAG Amplicon Length (bp) 122 Chromosome Location 14:34998580-35002699 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 104 R2 0.9989 cDNA Cq 19.93 cDNA Tm (Celsius) 79.5 gDNA Cq 34.63 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report EAPP, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Evidence for Differential Alternative Splicing in Blood of Young Boys With
Stamova et al. Molecular Autism 2013, 4:30 http://www.molecularautism.com/content/4/1/30 RESEARCH Open Access Evidence for differential alternative splicing in blood of young boys with autism spectrum disorders Boryana S Stamova1,2,5*, Yingfang Tian1,2,4, Christine W Nordahl1,3, Mark D Shen1,3, Sally Rogers1,3, David G Amaral1,3 and Frank R Sharp1,2 Abstract Background: Since RNA expression differences have been reported in autism spectrum disorder (ASD) for blood and brain, and differential alternative splicing (DAS) has been reported in ASD brains, we determined if there was DAS in blood mRNA of ASD subjects compared to typically developing (TD) controls, as well as in ASD subgroups related to cerebral volume. Methods: RNA from blood was processed on whole genome exon arrays for 2-4–year-old ASD and TD boys. An ANCOVA with age and batch as covariates was used to predict DAS for ALL ASD (n=30), ASD with normal total cerebral volumes (NTCV), and ASD with large total cerebral volumes (LTCV) compared to TD controls (n=20). Results: A total of 53 genes were predicted to have DAS for ALL ASD versus TD, 169 genes for ASD_NTCV versus TD, 1 gene for ASD_LTCV versus TD, and 27 genes for ASD_LTCV versus ASD_NTCV. These differences were significant at P <0.05 after false discovery rate corrections for multiple comparisons (FDR <5% false positives). A number of the genes predicted to have DAS in ASD are known to regulate DAS (SFPQ, SRPK1, SRSF11, SRSF2IP, FUS, LSM14A). In addition, a number of genes with predicted DAS are involved in pathways implicated in previous ASD studies, such as ROS monocyte/macrophage, Natural Killer Cell, mTOR, and NGF signaling. -

Mapping of a Chromosome 12 Region Associated with Airway Hyperresponsiveness in a Recombinant Congenic Mouse Strain and Selectio
Mapping of a Chromosome 12 Region Associated with Airway Hyperresponsiveness in a Recombinant Congenic Mouse Strain and Selection of Potential Candidate Genes by Expression and Sequence Variation Analyses Cynthia Kanagaratham1*, Rafael Marino2, Pierre Camateros2, John Ren3, Daniel Houle4, Robert Sladek1,2,5, Silvia M. Vidal1,3, Danuta Radzioch1,2 1 Department of Human Genetics, McGill University, Montreal, Quebec, Canada, 2 Faculty of Medicine, Division of Experimental Medicine, McGill University, Montreal, Quebec, Canada, 3 Department of Microbiology and Immunology, McGill University, Montreal, Quebec, Canada, 4 Research Institute of the McGill University Health Center, Montreal, Quebec, Canada, 5 McGill University and Genome Quebec Innovation Centre, Montreal, Quebec, Canada Abstract In a previous study we determined that BcA86 mice, a strain belonging to a panel of AcB/BcA recombinant congenic strains, have an airway responsiveness phenotype resembling mice from the airway hyperresponsive A/J strain. The majority of the BcA86 genome is however from the hyporesponsive C57BL/6J strain. The aim of this study was to identify candidate regions and genes associated with airway hyperresponsiveness (AHR) by quantitative trait locus (QTL) analysis using the BcA86 strain. Airway responsiveness of 205 F2 mice generated from backcrossing BcA86 strain to C57BL/6J strain was measured and used for QTL analysis to identify genomic regions in linkage with AHR. Consomic mice for the QTL containing chromosomes were phenotyped to study the contribution of each chromosome to lung responsiveness. Candidate genes within the QTL were selected based on expression differences in mRNA from whole lungs, and the presence of coding non- synonymous mutations that were predicted to have a functional effect by amino acid substitution prediction tools. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Techniques for Storing and Processing Next-Generation DNA Sequencing Data
Techniques for Storing and Processing Next-Generation DNA Sequencing Data THESIS Presented in Partial Fulfillment of the Requirements for the Degree Master of Science in the Graduate School of The Ohio State University By Terry Camerlengo Graduate Program in Biophysics The Ohio State University 2014 Master's Examination Committee: Professor Kun Huang, PhD Professor Raghu Machiraju, PhD Professor Carlos Alvarez, PhD Copyright by Terry Camerlengo 2014 ABSTRACT Genomics is undergoing unprecedented transformation due to rapid improvements in genetic sequencing technology, which has lowered costs for genetic sequencing experiments while increasing the amount of data generated in a typical experiment (McKinsey Global Institute, May 2013, pp. 86-94). The increase in data has shifted the burden from analysis and research to expertise in IT hardware and network support for distributed and efficient processing. Bioinformaticians, in response to a data-rich environment, are challenged to develop better and faster algorithms to solve problems in genomics and molecular biology research. This thesis examines the storage and data processing issues inherent in next- generation DNA sequencing (NGS). This work details the design and implementation of a software prototype that exemplifies the current approaches as it relates to the efficient storage of NGS data. The software library is utilized within the context of a previous software project which accompanies the publication related to the HT_SOSA assay. The software for the HT_SOSA, called NGSPositionCounter, demonstrates a workflow that is common in a molecular biology research lab. In an effort to scale beyond the research institute, the software library‟s architecture takes into account scalability considerations ii for data storage and processing demands that are more likely to be encountered in a clinical or commercial enterprise. -

Receptor Signaling Through Osteoclast-Associated Monocyte
Downloaded from http://www.jimmunol.org/ by guest on September 29, 2021 is online at: average * The Journal of Immunology The Journal of Immunology , 20 of which you can access for free at: 2015; 194:3169-3179; Prepublished online 27 from submission to initial decision 4 weeks from acceptance to publication February 2015; doi: 10.4049/jimmunol.1402800 http://www.jimmunol.org/content/194/7/3169 Collagen Induces Maturation of Human Monocyte-Derived Dendritic Cells by Signaling through Osteoclast-Associated Receptor Heidi S. Schultz, Louise M. Nitze, Louise H. Zeuthen, Pernille Keller, Albrecht Gruhler, Jesper Pass, Jianhe Chen, Li Guo, Andrew J. Fleetwood, John A. Hamilton, Martin W. Berchtold and Svetlana Panina J Immunol cites 43 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Freely available online through http://www.jimmunol.org/content/suppl/2015/02/27/jimmunol.140280 0.DCSupplemental This article http://www.jimmunol.org/content/194/7/3169.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2015 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -
EAPP (1E4) Mouse Mab A
Revision 1 C 0 2 - t EAPP (1E4) Mouse mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] Support: 877-678-TECH (8324) 6 6 Web: [email protected] 1 www.cellsignal.com 5 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IF-IC H M R Mk Endogenous 45 Mouse IgG1 Q56P03 55837 Product Usage Information Application Dilution Western Blotting 1:1000 Immunofluorescence (Immunocytochemistry) 1:800 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity EAPP (1E4) Mouse mAb detects endogenous levels of total EAPP protein. Species Reactivity: Human, Mouse, Rat, Monkey Source / Purification Monoclonal antibody is produced by immunizing animals with a recombinant amino- terminal fragment of human EAPP protein. Background Regulation of the E2F family of transcription factors, primarily through the retinoblastoma protein (pRb), is vital for control of cell proliferation and cell death (reviewed in 1). E2F- associated phosphoprotein (EAPP) was identified as an E2F-family binding protein that modulates E2F-regulated transcription and may be required for S phase entry. EAPP is expressed at varied levels in all tissues and cell types examined and its expression is reduced in nocodazole-arrested cells (2). Mass spectrometry studies have identified multiple EAPP phosphorylation sites including Ser109 and Ser111, but biological consequences of EAPP phosphorylation have yet to be elucidated (3-5). -
SUPPLEMENTARY DATA Supplementary Table 1. Characteristics of the Organ Donors and Human Islet Preparations Used for RNA-Seq
SUPPLEMENTARY DATA Supplementary Table 1. Characteristics of the organ donors and human islet preparations used for RNA-seq and independent confirmation and mechanistic studies. Gender Age BMI Cause of death Purity (years) (kg/m2) (%) F 77 23.8 Trauma 45 M 36 26.3 CVD 51 M 77 25.2 CVD 62 F 46 22.5 CVD 60 M 40 26.2 Trauma 34 M 59 26.7 NA 58 M 51 26.2 Trauma 54 F 79 29.7 CH 21 M 68 27.5 CH 42 F 76 25.4 CH 30 F 75 29.4 CVD 24 F 73 30.0 CVD 16 M 63 NA NA 46 F 64 23.4 CH 76 M 69 25.1 CH 68 F 23 19.7 Trauma 70 M 47 27.7 CVD 48 F 65 24.6 CH 58 F 87 21.5 Trauma 61 F 72 23.9 CH 62 M 69 25 CVD 85 M 85 25.5 CH 39 M 59 27.7 Trauma 56 F 76 19.5 CH 35 F 50 20.2 CH 70 F 42 23 CVD 48 M 52 24.5 CH 60 F 79 27.5 CH 89 M 56 24.7 Cerebral ischemia 47 M 69 24.2 CVD 57 F 79 28.1 Trauma 61 M 79 23.7 NA 13 M 82 23 CH 61 M 32 NA NA 75 F 23 22.5 Cardiac arrest 46 M 51 NA Trauma 37 Abbreviations: F: Female; M: Male; BMI: Body mass index; CVD: Cardiovascular disease; CH: Cerebral hemorrhage. -

Datasheet Blank Template
SAN TA C RUZ BI OTEC HNOL OG Y, INC . EAPP (1E4): sc-130357 BACKGROUND PRODUCT E2F transcription factors play a major role in apoptosis and cell proliferation Each vial contains 200 µg IgG kappa light chain in 1.0 ml of PBS with < 0.1% and are found to be frequently deregulated in cancers. Through interactions sodium azide and 0.1% gelatin. with cell cycle regulators such as cyclins, cyclin-dependent kinases and retino- blastoma protein (Rb), E2F family members also integrate cell cycle progres - APPLICATIONS sion. EAPP (E2F-associated phosphoprotein) is a 285 amino acid highly phos - EAPP (1E4) is recommended for detection of EAPP of mouse, rat and human phorylated nuclear protein that fine-tunes E2F activities by interacting with origin by Western Blotting (starting dilution 1:200, dilution range 1:100- E2F-1, E2F-2 and E2F-3, but not E2F-4. By binding to the N-terminal domain 1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein (1 ml of these E2F family members, EAPP interferes with the binding of cyclin A, of cell lysate)] and immunofluorescence (starting dilution 1:50, dilution range Sp1 transcription factors, EBP1 and EBP2, therefore influencing E2F activity. 1:50-1:500). Interestingly, EAPP is expressed during the cell cycle, but disappears during mitosis, suggesting that this step is necessary to complete the cell cycle. Suitable for use as control antibody for EAPP siRNA (h): sc-92116, EAPP EAPP is ubiquitously expressed, with highest levels found in placenta, siRNA (m): sc-143265, EAPP shRNA Plasmid (h): sc-92116-SH, EAPP shRNA pan creas, skeletal muscle and heart. -

Identification of Genetic Factors in Atherosclerosis Using an Apoe Mouse Model
Identification of Genetic Factors in Atherosclerosis Using an Apoe Mouse Model Andrew Todd Grainger Los Gatos, California Bachelors of Science in Molecular Biology, University of California San Diego, San Diego, California, 2013 Masters of Science in Biology, University of California San Diego, San Diego, California, 2014 A Dissertation Presented to the Graduate Faculty of the University of Virginia in Candidacy for the Degree of Doctor of Philosophy Department of Biochemistry and Molecular Genetics University of Virginia December 2019 Dr. Weibin Shi Dr. Charles Farber Dr. Aakrosh Ratan Dr. Norbert Leitinger I Abstract Atherosclerosis is the primary cause of coronary artery disease (CAD), ischemic stroke and peripheral arterial disease. Despite major achievements made in the past few decades, CAD and atherosclerosis-related events remain the number one cause of death in the United States and other developed countries. Therefore, there is a critical medical need to develop novel and effective therapies. An effective way to find new targets for intervention is through conducting genetic studies in animal models. When deficient in Apoe, mouse strains BALB/cJ and SM/J exhibit distinct differences in atherosclerosis and its associated risk factors. We hypothesized that linkage analysis of progeny derived from these inbred strains would lead to the discovery of new genes and new pathways in atherosclerosis and its associated cardiometabolic phenotypes. F2 mice were generated from an intercross between the two Apoe-/- strains and fed 12 weeks of Western diet. Many QTL loci were mapped for plasma lipids and glucose, carotid lesion size, and aortic lesion size. This included a significant QTL for aortic atherosclerosis, Ath49, which was mapped to the major histocompatibility region. -

HBV DNA Integration and Clonal Hepatocyte Expansion in Chronic Hepatitis B Patients Considered Immune Tolerant
Accepted Manuscript HBV DNA Integration and Clonal Hepatocyte Expansion in Chronic Hepatitis B Patients Considered Immune Tolerant William S. Mason, Upkar S. Gill, Samuel Litwin, Yan Zhou, Suraj Peri, Oltin Pop, Michelle L.W. Hong, Sandhia Naik, Alberto Quaglia, Antonio Bertoletti, Patrick T.F. Kennedy PII: S0016-5085(16)34808-9 DOI: 10.1053/j.gastro.2016.07.012 Reference: YGAST 60585 To appear in: Gastroenterology Accepted Date: 7 July 2016 Please cite this article as: Mason WS, Gill US, Litwin S, Zhou Y, Peri S, Pop O, Hong MLW, Naik S, Quaglia A, Bertoletti A, Kennedy PTF, HBV DNA Integration and Clonal Hepatocyte Expansion in Chronic Hepatitis B Patients Considered Immune Tolerant, Gastroenterology (2016), doi: 10.1053/ j.gastro.2016.07.012. This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. ACCEPTED MANUSCRIPT TITLE: HBV DNA Integration and Clonal Hepatocyte Expansion in Chronic Hepatitis B Patients Considered Immune Tolerant SHORT TITLE : Immunopathology in immune tolerant CHB AUTHORS: William S. Mason 1, Upkar S. Gill 2, Samuel Litwin 1, Yan Zhou 1, Suraj Peri 1, Oltin Pop 3, Michelle L.W. Hong 4, Sandhia Naik 5, Alberto Quaglia 3, Antonio Bertoletti 4 & Patrick T.F. -

Homologue-Specific Chromosome Sequencing Characterizes Translocation Junctions and Permits Allelic Assignment Fumio Kasai1,2,*, Jorge C
DNA Research, 2018, 25(4), 353–360 doi: 10.1093/dnares/dsy007 Advance Access Publication Date: 6 March 2018 Full Paper Full Paper Homologue-specific chromosome sequencing characterizes translocation junctions and permits allelic assignment Fumio Kasai1,2,*, Jorge C. Pereira2, Arihiro Kohara1, and Malcolm A. Ferguson-Smith2 1Japanese Collection of Research Bioresources (JCRB) Cell Bank, Laboratory of Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, Ibaraki 567-0085, Osaka, Japan, and 2Cambridge Resource Centre for Comparative Genomics, Department of Veterinary Medicine, University of Cambridge, Cambridge CB3 0ES, UK *To whom correspondence should be addressed. Tel. þ81 72 641 9851. Fax. þ81 72 641 9859. Email: [email protected] Edited by Dr. Yuji Kohara Received 13 August 2017; Editorial decision 11 February 2018; Accepted 12 February 2018 Abstract Chromosome translocations can be detected by cytogenetic analysis, but it is hard to character- ize the breakpoints at the sequence level. Chromosome sorting by flow cytometry produces flow karyotypes that enable the isolation of abnormal chromosomes and the generation of chromosome-specific DNA. In this study, a derivative chromosome t(9; 14) and its homologous normal chromosomes 9 and 14 from the Ishikawa 3-H-12 cell line were sorted to collect homologue-specific samples. Chromosome sequencing identified the breakpoint junction in the der(9) at 9p24.3 and 14q13.1 and uncovered the formation of a fusion gene, WASH1–NPAS3. Amplicon sequencing targeted for neighbouring genes at the fusion breakpoint revealed that the variant frequencies correlate with the allelic copy number. Sequencing of sorted chromo- somes permits the assignment of allelic variants and can lead to the characterization of abnor- mal chromosomes.