14 Chromosome Chapter
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
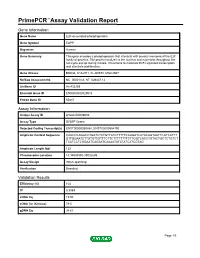
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name E2F-associated phosphoprotein Gene Symbol EAPP Organism Human Gene Summary This gene encodes a phosphoprotein that interacts with several members of the E2F family of proteins. The protein localizes to the nucleus and is present throughout the cell cycle except during mitosis. It functions to modulate E2F-regulated transcription and stimulate proliferation. Gene Aliases BM036, C14orf11, FLJ20578, MGC4957 RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.433269 Ensembl Gene ID ENSG00000129518 Entrez Gene ID 55837 Assay Information Unique Assay ID qHsaCID0008053 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000250454, ENST00000554792 Amplicon Context Sequence CAACCCAGGCCTGATCTCTGTTATCTTTTTCAGGATCATACAGTAATTCGTCATTT GTTGGAATCTTGTGTTGTTTCTTCTTTTTTTTCTTGGTCACCTGTACTGCTCTGTCT TCATCCTCGGAATCAGAATCAAAATATATATCATCGTAG Amplicon Length (bp) 122 Chromosome Location 14:34998580-35002699 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 104 R2 0.9989 cDNA Cq 19.93 cDNA Tm (Celsius) 79.5 gDNA Cq 34.63 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report EAPP, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Evidence for Differential Alternative Splicing in Blood of Young Boys With
Stamova et al. Molecular Autism 2013, 4:30 http://www.molecularautism.com/content/4/1/30 RESEARCH Open Access Evidence for differential alternative splicing in blood of young boys with autism spectrum disorders Boryana S Stamova1,2,5*, Yingfang Tian1,2,4, Christine W Nordahl1,3, Mark D Shen1,3, Sally Rogers1,3, David G Amaral1,3 and Frank R Sharp1,2 Abstract Background: Since RNA expression differences have been reported in autism spectrum disorder (ASD) for blood and brain, and differential alternative splicing (DAS) has been reported in ASD brains, we determined if there was DAS in blood mRNA of ASD subjects compared to typically developing (TD) controls, as well as in ASD subgroups related to cerebral volume. Methods: RNA from blood was processed on whole genome exon arrays for 2-4–year-old ASD and TD boys. An ANCOVA with age and batch as covariates was used to predict DAS for ALL ASD (n=30), ASD with normal total cerebral volumes (NTCV), and ASD with large total cerebral volumes (LTCV) compared to TD controls (n=20). Results: A total of 53 genes were predicted to have DAS for ALL ASD versus TD, 169 genes for ASD_NTCV versus TD, 1 gene for ASD_LTCV versus TD, and 27 genes for ASD_LTCV versus ASD_NTCV. These differences were significant at P <0.05 after false discovery rate corrections for multiple comparisons (FDR <5% false positives). A number of the genes predicted to have DAS in ASD are known to regulate DAS (SFPQ, SRPK1, SRSF11, SRSF2IP, FUS, LSM14A). In addition, a number of genes with predicted DAS are involved in pathways implicated in previous ASD studies, such as ROS monocyte/macrophage, Natural Killer Cell, mTOR, and NGF signaling. -

Construction of Stable Mouse Arti Cial Chromosome from Native Mouse
Construction of Stable Mouse Articial Chromosome from Native Mouse Chromosome 10 for Generation of Transchromosomic Mice Satoshi Abe Tottori University Kazuhisa Honma Trans Chromosomics, Inc Akane Okada Tottori University Kanako Kazuki Tottori University Hiroshi Tanaka Trans Chromosomics, Inc Takeshi Endo Trans Chromosomics, Inc Kayoko Morimoto Trans Chromosomics, Inc Takashi Moriwaki Tottori University Shusei Hamamichi Tottori University Yuji Nakayama Tottori University Teruhiko Suzuki Tokyo Metropolitan Institute of Medical Science Shoko Takehara Trans Chromosomics, Inc Mitsuo Oshimura Tottori University Yasuhiro Kazuki ( [email protected] ) Tottori University Research Article Page 1/21 Keywords: mouse articial chromosome (MAC), microcell-mediated chromosome transfer (MMCT), chromosome engineering, transchromosomic (Tc) mouse, humanized model mouse Posted Date: July 9th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-675300/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 2/21 Abstract Mammalian articial chromosomes derived from native chromosomes have been applied to biomedical research and development by generating cell sources and transchromosomic (Tc) animals. Human articial chromosome (HAC) is a precedent chromosomal vector which achieved generation of valuable humanized animal models for fully human antibody production and human pharmacokinetics. While humanized Tc animals created by HAC vector have attained signicant contributions, there was a potential issue to be addressed regarding stability in mouse tissues, especially highly proliferating hematopoietic cells. Mouse articial chromosome (MAC) vectors derived from native mouse chromosome 11 demonstrated improved stability, and they were utilized for humanized Tc mouse production as a standard vector. In mouse, however, stability of MAC vector derived from native mouse chromosome other than mouse chromosome 11 remains to be evaluated. -

IL21R Expressing CD14+CD16+ Monocytes Expand in Multiple
Plasma Cell Disorders SUPPLEMENTARY APPENDIX IL21R expressing CD14 +CD16 + monocytes expand in multiple myeloma patients leading to increased osteoclasts Marina Bolzoni, 1 Domenica Ronchetti, 2,3 Paola Storti, 1,4 Gaetano Donofrio, 5 Valentina Marchica, 1,4 Federica Costa, 1 Luca Agnelli, 2,3 Denise Toscani, 1 Rosanna Vescovini, 1 Katia Todoerti, 6 Sabrina Bonomini, 7 Gabriella Sammarelli, 1,7 Andrea Vecchi, 8 Daniela Guasco, 1 Fabrizio Accardi, 1,7 Benedetta Dalla Palma, 1,7 Barbara Gamberi, 9 Carlo Ferrari, 8 Antonino Neri, 2,3 Franco Aversa 1,4,7 and Nicola Giuliani 1,4,7 1Myeloma Unit, Dept. of Medicine and Surgery, University of Parma; 2Dept. of Oncology and Hemato-Oncology, University of Milan; 3Hematology Unit, “Fondazione IRCCS Ca’ Granda”, Ospedale Maggiore Policlinico, Milan; 4CoreLab, University Hospital of Parma; 5Dept. of Medical-Veterinary Science, University of Parma; 6Laboratory of Pre-clinical and Translational Research, IRCCS-CROB, Referral Cancer Center of Basilicata, Rionero in Vulture; 7Hematology and BMT Center, University Hospital of Parma; 8Infectious Disease Unit, University Hospital of Parma and 9“Dip. Oncologico e Tecnologie Avanzate”, IRCCS Arcispedale Santa Maria Nuova, Reggio Emilia, Italy ©2017 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2016.153841 Received: August 5, 2016. Accepted: December 23, 2016. Pre-published: January 5, 2017. Correspondence: [email protected] SUPPLEMENTAL METHODS Immunophenotype of BM CD14+ in patients with monoclonal gammopathies. Briefly, 100 μl of total BM aspirate was incubated in the dark with anti-human HLA-DR-PE (clone L243; BD), anti-human CD14-PerCP-Cy 5.5, anti-human CD16-PE-Cy7 (clone B73.1; BD) and anti-human CD45-APC-H 7 (clone 2D1; BD) for 20 min. -

14Q13 Deletions FTNW
14q13 deletions rarechromo.org 14q13 deletions A chromosome 14 deletion means that part of one of the body’s chromosomes (chromosome 14) has been lost or deleted. If the deleted material contains important genes, learning disability, developmental delay and health problems may occur. How serious these problems are depends on how much of the chromosome has been deleted, which genes have been lost and where precisely the deletion is. The features associated with 14q13 deletions vary from person to person, but are likely to include a degree of developmental delay, an unusually small or large head, a raised risk of medical problems and unusual facial features. Genes and chromosomes Our bodies are made up of billions of cells. Most of these cells contain a complete set of thousands of genes that act as instructions, controlling our growth, development and how our bodies work. Inside human cells there is a nucleus where the genes are carried on microscopically small, thread-like structures called chromosomes which are made up p arm p arm of DNA. p arm p arm Chromosomes come in pairs of different sizes and are numbered from largest to smallest, roughly according to their size, from number 1 to number 22. In addition to these so-called autosomal chromosomes there are the sex chromosomes, X and Y. So a human cell has 46 chromosomes: 23 inherited from the mother and 23 inherited from the father, making two sets of 23 chromosomes. A girl has two X chromosomes (XX) while a boy will have one X and one Y chromosome (XY). -

The Breakpoint of an Inversion of Chromosome 14 in a T-Cell
Proc. Nati. Acad. Sci. USA Vol. 84, pp. 9069-9073, December 1987 Genetics The breakpoint of an inversion of chromosome 14 in a T-cell leukemia: Sequences downstream of the immunoglobulin heavy chain locus are implicated in tumorigenesis (T-cell receptor/ataxia-telangiectasia) R. BAER*t, A. HEPPELLt, A. M. R. TAYLORt, P. H. RABBITTS§, B. BOULLIER§, AND T. H. RABBITTS* *Medical Research Council Laboratory of Molecular Biology and §Ludwig Institute for Cancer Research, Hills Road, Cambridge, CB2 2QH, England; and tUniversity of Birmingham, Cancer Research Laboratories, Department of Cancer Studies, the Medical School, Birmingham, B15 2TJ, England Communicated by C. Milstein, August 11, 1987 (received for review July 13, 1987) ABSTRACT T-cell tumors are characterized by inversions this alternative view. Cytogenetic studies of inv(14) chromo- or translocations of chromosome 14. The breakpoints of these some, by high resolution banding, have identified two differ- karyotypic abnormalities occur in chromosome bands 14qll ent break-reassociation points involved in inv(14) chromo- and 14q32-the same bands in which the T-cell receptor (TCR) somes (15). Notably, the 14q32 breakpoints of nonmalignant a-chain and immunoglobulin heavy chain genes have been clone inversions associated in ataxia-telangiectasia (A-T) are mapped, respectively. Patients with ataxia-telangiectasia are distinct from the 14q32 breakpoints of sporadic inversions particularly prone to development of T-cell chronic lympho- from normal subjects. A similar dichotomy of 14q32 cytic leukemia with such chromosomal abnormalities. We now breakpoints was found in clonal and sporadic t(14;14)(qll;- describe DNA rearrangements of the TCR a-chain gene in an q32) translocations (16). -

Amplified Fragments of an Autosome-Borne Gene
G C A T T A C G G C A T genes Article Amplified Fragments of an Autosome-Borne Gene Constitute a Significant Component of the W Sex Chromosome of Eremias velox (Reptilia, Lacertidae) Artem Lisachov 1,2,* , Daria Andreyushkova 3, Guzel Davletshina 2,3, Dmitry Prokopov 3 , Svetlana Romanenko 3 , Svetlana Galkina 4 , Alsu Saifitdinova 5 , Evgeniy Simonov 1, Pavel Borodin 2,6 and Vladimir Trifonov 3,6 1 Institute of Environmental and Agricultural Biology (X-BIO), University of Tyumen, Lenina str. 23, 625003 Tyumen, Russia; [email protected] 2 Institute of Cytology and Genetics SB RAS, Acad. Lavrentiev Ave. 10, 630090 Novosibirsk, Russia; [email protected] (G.D.); [email protected] (P.B.) 3 Institute of Molecular and Cellular Biology SB RAS, Acad. Lavrentiev Ave. 8/2, 630090 Novosibirsk, Russia; [email protected] (D.A.); [email protected] (D.P.); [email protected] (S.R.); [email protected] (V.T.) 4 Department of Genetics and Biotechnology, Saint Petersburg State University, Universitetskaya Emb. 7–9, 199034 Saint Petersburg, Russia; [email protected] 5 Department of Human and Animal Anatomy and Physiology, Herzen State Pedagogical University of Russia, Moyka Emb. 48, 191186 Saint Petersburg, Russia; saifi[email protected] 6 Novosibirsk State University, Pirogova str. 3, 630090 Novosibirsk, Russia Citation: Lisachov, A.; * Correspondence: [email protected] Andreyushkova, D.; Davletshina, G.; Prokopov, D.; Romanenko, S.; Abstract: Heteromorphic W and Y sex chromosomes often experience gene loss and heterochroma- Galkina, S.; Saifitdinova, A.; Simonov, tinization, which is frequently viewed as their “degeneration”. -

WNT16 Is a New Marker of Senescence
Table S1. A. Complete list of 177 genes overexpressed in replicative senescence Value Gene Description UniGene RefSeq 2.440 WNT16 wingless-type MMTV integration site family, member 16 (WNT16), transcript variant 2, mRNA. Hs.272375 NM_016087 2.355 MMP10 matrix metallopeptidase 10 (stromelysin 2) (MMP10), mRNA. Hs.2258 NM_002425 2.344 MMP3 matrix metallopeptidase 3 (stromelysin 1, progelatinase) (MMP3), mRNA. Hs.375129 NM_002422 2.300 HIST1H2AC Histone cluster 1, H2ac Hs.484950 2.134 CLDN1 claudin 1 (CLDN1), mRNA. Hs.439060 NM_021101 2.119 TSPAN13 tetraspanin 13 (TSPAN13), mRNA. Hs.364544 NM_014399 2.112 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.070 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.026 DCBLD2 discoidin, CUB and LCCL domain containing 2 (DCBLD2), mRNA. Hs.203691 NM_080927 2.007 SERPINB2 serpin peptidase inhibitor, clade B (ovalbumin), member 2 (SERPINB2), mRNA. Hs.594481 NM_002575 2.004 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.989 OBFC2A Oligonucleotide/oligosaccharide-binding fold containing 2A Hs.591610 1.962 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.947 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 2, mRNA. Hs.472101 NM_182797 1.934 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 1, mRNA. Hs.472101 NM_000933 1.933 KRTAP1-5 keratin associated protein 1-5 (KRTAP1-5), mRNA. Hs.534499 NM_031957 1.894 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.884 CYTL1 cytokine-like 1 (CYTL1), mRNA. Hs.13872 NM_018659 tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain (TNFRSF10D), 1.848 TNFRSF10D Hs.213467 NM_003840 mRNA. -

Somatic Rearrangement of Chromosome 14 in Human Lymphocytes (Leukemia/Lymphoproliferation/Ataxia-Telangiectasia) BARBARA KAISER Mccaw*, FREDERICK HECHT*, DAVID G
Proc. Nat. Acad. Sci. USA Vol. 72, No. 6, pp. 2071-2075, June 1975 Somatic Rearrangement of Chromosome 14 in Human Lymphocytes (leukemia/lymphoproliferation/ataxia-telangiectasia) BARBARA KAISER McCAW*, FREDERICK HECHT*, DAVID G. HARNDENt, AND RAYMOND L. TEPLITZ$ * Genetics Clinic, Child Development and Rehabilitation Center, University of Oregon Health Sciences Center, Portland, Oreg. 97201; t Department of Cancer Studies, The Medical School, The University of Birmingham, Birmingham B15 2TJ, England; and * Department of Cytogenetics and Cytology, City of Hope National Medical Center, Duarte, California 91010 Communicated by David M. Prescott, March 17, 1975 ABSTRACT Ataxia-telangiectasia is a rare genetic dis- tion to lymphoid malignancy (8). Previous longitudinal order associated with immune deficiency, chromosome studies of benign lymphocytes in a patient with A-T, showed instability, and a predisposition to lymphoid malignancy. We have detected chromosomally anomalous clones of a clone marked by a translocation involving both chromo- lymphocytes in eight patients with this disorder. Chromo- somes 14 (9). We have now detected similar clones of chro- some banding disclosed that the clones are consistently mosomally marked lymphocytes in seven other patients with marked by structural rearrangement of the long arm (q) of this disorder. chromosome 14. A translocation involving 14q was found in clones obtained from seven of the eight patients whereas The clones consistently show rearrangement of the long a ring 14 chromosome was found in a clone obtained from arm (q) of chromosome 14. The break points in this chromo- the other. These findings as well as data obtained by others some are within a specific region, and there is no obvious loss for patients with ataxia-telangiectasia suggest that struc- or gain of chromosome material. -

Mapping of a Chromosome 12 Region Associated with Airway Hyperresponsiveness in a Recombinant Congenic Mouse Strain and Selectio
Mapping of a Chromosome 12 Region Associated with Airway Hyperresponsiveness in a Recombinant Congenic Mouse Strain and Selection of Potential Candidate Genes by Expression and Sequence Variation Analyses Cynthia Kanagaratham1*, Rafael Marino2, Pierre Camateros2, John Ren3, Daniel Houle4, Robert Sladek1,2,5, Silvia M. Vidal1,3, Danuta Radzioch1,2 1 Department of Human Genetics, McGill University, Montreal, Quebec, Canada, 2 Faculty of Medicine, Division of Experimental Medicine, McGill University, Montreal, Quebec, Canada, 3 Department of Microbiology and Immunology, McGill University, Montreal, Quebec, Canada, 4 Research Institute of the McGill University Health Center, Montreal, Quebec, Canada, 5 McGill University and Genome Quebec Innovation Centre, Montreal, Quebec, Canada Abstract In a previous study we determined that BcA86 mice, a strain belonging to a panel of AcB/BcA recombinant congenic strains, have an airway responsiveness phenotype resembling mice from the airway hyperresponsive A/J strain. The majority of the BcA86 genome is however from the hyporesponsive C57BL/6J strain. The aim of this study was to identify candidate regions and genes associated with airway hyperresponsiveness (AHR) by quantitative trait locus (QTL) analysis using the BcA86 strain. Airway responsiveness of 205 F2 mice generated from backcrossing BcA86 strain to C57BL/6J strain was measured and used for QTL analysis to identify genomic regions in linkage with AHR. Consomic mice for the QTL containing chromosomes were phenotyped to study the contribution of each chromosome to lung responsiveness. Candidate genes within the QTL were selected based on expression differences in mRNA from whole lungs, and the presence of coding non- synonymous mutations that were predicted to have a functional effect by amino acid substitution prediction tools. -

Paracentric Inversion of Chromosome 14: a Case Report
Jpn J Human Genet 39, 353-356, 1994 Case Report PARACENTRIC INVERSION OF CHROMOSOME 14: A CASE REPORT Shigeki UEHARA, Shingo TANIGAWARA,Yoichi TAKEYAMA, Toshifumi TAKABAYASHI,Kunihiro OKAMURA,and Akira YAJIMA Department of Obstetrics and Gynecology, Tohoku University School of Medicine, Seiryo-machi, Aoba-ku, Sendai 980-77, Japan Summary A new case of familial heterozygous paracentric inversion in the long arm of chromosome 14 [inv(14)(q22q32)] is presented. The rearrangement was first ascertained in a fetus examined due to advanced maternal age, and then detected in.,the father. The phenotypes of the newborn and the father were completely normal. The parents had no history of spontaneous abortion. With reference to previous reports, the risk of clinical abnormalities are discussed for both de novo and fa- milial paracentric inversions of chromosome 14. Key Words paracentric inversion, chromosome 14, phenotype, chromo- some rearrangement INTRODUCTION We recently experienced a Japanese family having paracentric inversion of chromosome 14, inv(14)(q22q32), which is herein reported. With reference to previous reports, we discuss the possibility of clinical abnormalities in various types of inv(14). CASE REPORT The mother, a 39-year-old healthy Japanese, received a prenatal chromosomal examination at 15 weeks of gestation because of her advanced age. The father was a 41-year-old healthy Japanese. They were not consanguineous. This was the mother's second marriage. She had experienced one normal pregnancy with the present husband. The phenotype of the first child, a male, was normal. Pre- natal cytogenetic analysis revealed a heterozygous paracentric inversion in the long Received May 13, 1994; Revised version accepted June 28, 1994. -

NF1) Pseudogenes on Chromosomes 2, 14 and 22
European Journal of Human Genetics (2000) 8, 209–214 © 2000 Macmillan Publishers Ltd All rights reserved 1018–4813/00 $15.00 y www.nature.com/ejhg ARTICLE Mechanism of spreading of the highly related neurofibromatosis type 1 (NF1) pseudogenes on chromosomes 2, 14 and 22 Mirjam Luijten1, YingPing Wang2, Blaine T Smith2, Andries Westerveld1, Luc J Smink3, Ian Dunham3, Bruce A Roe2 and Theo JM Hulsebos1 1Department of Human Genetics, Academic Medical Center, University of Amsterdam, The Netherlands; 2Department of Chemistry and Biochemistry, University of Oklahoma, Norman, OK, USA; 3Sanger Centre, Wellcome Trust Genome Campus, Hinxton Hall, Cambridge, UK Neurofibromatosis type 1 (NF1) is a frequent hereditary disorder that involves tissues derived from the embryonic neural crest. Besides the functional gene on chromosome arm 17q, NF1-related sequences (pseudogenes) are present on a number of chromosomes including 2, 12, 14, 15, 18, 21, and 22. We elucidated the complete nucleotide sequence of the NF1 pseudogene on chromosome 22. Only the middle part of the functional gene but not exons 21–27a, encoding the functionally important GAP-related domain of the NF1 protein, is presented in this pseudogene. In addition to the two known NF1 pseudogenes on chromosome 14 we identified two novel variants. A phylogenetic tree was constructed, from which we concluded that the NF1 pseudogenes on chromosomes 2, 14, and 22 are closely related to each other. Clones containing one of these pseudogenes cross-hybridised with the other pseudogenes in this subset, but did not reveal any in situ hybridisation with the functional NF1 gene or with NF1 pseudogenes on other chromosomes.