Evaluation of the Detection of GBA Missense Mutations and Other Variants Using the Oxford Nanopore Minion
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

In Silico Haplotyping, Genotyping and Analysis of Resequencing Data Using Markov Models
IN SILICO HAPLOTYPING, GENOTYPING AND ANALYSIS OF RESEQUENCING DATA USING MARKOV MODELS by Yun Li A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Biostatistics) in The University of Michigan 2009 Doctoral Committee: Professor Gonçalo R. Abecasis, Co-Chair Professor Michael Lee Boehnke, Co-Chair Professor Margit Burmeister Professor Roderick J.A. Little Assistant Professor Noah A. Rosenberg © Yun Li 2009 To Mom, Dad, Iory and Duan. ii Acknowledgements I am grateful to the myriad of people who have made this work possible. First of all, I would like to thank Gonçalo and Mike for teaching me about science, for letting me discover how much I can enjoy research, for being encouraging and enthusiastic both because of and despite what we found, for guiding me to grow up as an independent researcher, for being exceptional role models both career-wise and family-wise, and for being approachable anytime I need help. I owe gratitude to my committee: Rod, Margit and Noah for their help and guidance beyond their duties as committee members. I sincerely appreciate their time, advice, and encouragement. Special thanks to Rod for brining in the missing data perspectives; to Margit for always asking questions that I have an answer in the next few slides; and to Noah for access to the HGDP data. I would like also to thank the CSG members: they make everyday research joyful and keep my spirits up even during my down times. I am also fortunate to be around a group of good friends including Qixuan, Jin, Rui, Jun, and Liming, who have made my 5-year graduate studies more colorful. -

Mafr De Enterococcus Faecalis Y Mgap De Streptococcus Pneumoniae
UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE CIENCIAS BIOLÓGICAS TESIS DOCTORAL Caracterización de dos activadores transcripcionales: MafR de Enterococcus faecalis y MgaP de Streptococcus pneumoniae Characterization of two transcriptional activators: MafR of Enterococcus faecalis and MgaP of Streptococcus pneumoniae MEMORIA PARA OPTAR AL GRADO DE DOCTORA PRESENTADA POR Ana Moreno Blanco DIRECTORA Alicia Bravo García Madrid © Ana Moreno Blanco, 2020 UNIVERSIDAD COMPLUTENSE DE MADRID Facultad de Ciencias Biológicas Departamento de Bioquímica y Biología Molecular TESIS DOCTORAL Caracterización de dos activadores transcripcionales: MafR de Enterococcus faecalis y MgaP de Streptococcus pneumoniae Characterization of two transcriptional activators: MafR of Enterococcus faecalis and MgaP of Streptococcus pneumoniae Ana Moreno Blanco Directora: Alicia Bravo García Madrid, 2020 UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE CIENCIAS BIOLÓGICAS TESIS DOCTORAL CARACTERIZACIÓN DE DOS ACTIVADORES TRANSCRIPCIONALES: MafR DE Enterococcus faecalis Y MgaP DE Streptococcus pneumoniae CHARACTERIZATION OF TWO TRANSCRIPTIONAL ACTIVATORS: MafR OF Enterococcus faecalis AND MgaP OF Streptococcus pneumoniae MEMORIA PARA OPTAR AL GRADO DE DOCTOR PRESENTADA POR ANA MORENO BLANCO DIRECTORA ALICIA BRAVO GARCÍA CENTRO DE INVESTIGACIONES BIOLÓGICAS MARGARITA SALAS CONSEJO SUPERIOR DE INVESTIGACIONES CIENTÍFICAS (CSIC) MADRID, 2020 A mi madre, mi padre y mis hermanos. “Que no sea inmortal puesto que es llama, pero que sea eterno mientras dure” Vinicius de Moraes Poco espacio para expresar todo lo que llevo dentro. Ciertas palabras se repiten una y otra vez a lo largo de estos agradecimientos, "orgullo", "confianza", "vida", "motivar"... No es falta de léxico, es ser afortunada por haberos encontrado en mi camino. A ti, Alicia, por tu infinita paciencia y buen hacer. Por darme la oportunidad y confiar en mí. -

Detecting Non-Allelic Homologous Recombination from High-Throughput Sequencing Data Matthew M Parks1, Charles E Lawrence1,2 and Benjamin J Raphael2,3*
Parks et al. Genome Biology (2015) 16:72 DOI 10.1186/s13059-015-0633-1 RESEARCH Open Access Detecting non-allelic homologous recombination from high-throughput sequencing data Matthew M Parks1, Charles E Lawrence1,2 and Benjamin J Raphael2,3* Abstract Non-allelic homologous recombination (NAHR) is a common mechanism for generating genome rearrangements and is implicated in numerous genetic disorders, but its detection in high-throughput sequencing data poses a serious challenge. We present a probabilistic model of NAHR and demonstrate its ability to find NAHR in low-coverage sequencing data from 44 individuals. We identify NAHR-mediated deletions or duplications in 109 of 324 potential NAHR loci in at least one of the individuals. These calls segregate by ancestry, are more common in closely spaced repeats, often result in duplicated genes or pseudogenes, and affect highly studied genes such as GBA and CYP2E1. Background detection and verification[16]. Experimental techniques Non-allelic homologous recombination (NAHR) is a bio- for discovering instances of structural variation, includ- logical mechanism for repairing broken chromosomes, ing array comparative genomic hybridization (aCGH), which results in gross genome rearrangements. A signif- SNP microarrays, and fluorescence in situ hybridiza- icant portion (approximately 10% to 22%) of all genome tion (FISH), are frustrated by repetitive regions and thus rearrangements in humans, also called structural varia- encounter considerable difficulty in detecting NAHR [16]. tions, is thought to be the result of NAHR [1-5]. Under- While validations of NAHR have been done using these standing and detecting NAHR in individuals provide techniques [3,11,17], they are not used on nearly the valuable insight for a wide variety of genomic disorders, same scale as high-throughput sequencing. -

Pseudogenes: Implications in Disease and Diagnostics
GeNeViSTA Pseudogenes: Implications in Disease and Diagnostics Divya Pasumarthi, Ashwin Dalal Diagnostics Division, Centre for DNA Fingerprinting and Diagnostics, Hyderabad, Telangana, India Correspondence to: Dr Ashwin Dalal Email: [email protected] The genome is the complete set of deoxy- of protein coding (1%) and non-coding regions. ribonucleic acid (DNA) in an organism. Human The protein coding part consists of exons and cells contain two copies of the haploid genome the non-coding part includes pseudogenes, gene consisting of base pairs of DNA. Only fragments, introns, untranslated regions, repetitive 9 1-2% of the mammalian3 × 10 genome codes for the DNA etc. (Figure 1). protein function and remaining codes for repeti- tive elements and non-coding regions. Repetitive What are Pseudogenes and why are elements include transposons and Alu elements. The gene is a hereditary unit of DNA. About 25,000 they important? genes are encoded in the DNA, which is organized Pseudogenes are also known as junk DNA. These into structures known as chromosomes. Unique are present in the noncoding part of the human non-coding sequences known as pseudogenes are genome. Pseudogenes have a lot of resemblance present in 15% of the human genome. to functional genes but have accumulated mu- tations which have inactivated them during the Organization of the human genome course of evolution. Pseudogenes can arise due to tandem duplication of normal genes and accumu- Human genome consists of nuclear and mito- lation of several mutations in them, which leads to chondrial genome. The nuclear genome consists loss of the ability of gene expression. Pseudogenes Figure 1 Organization of the human genome. -

The Genetics of Parkinson's Disease and Implications for Clinical Practice
G C A T T A C G G C A T genes Review The Genetics of Parkinson’s Disease and Implications for Clinical Practice Jacob Oliver Day 1 and Stephen Mullin 1,2,* 1 Faculty of Health, University of Plymouth, Plymouth PL4 8AA, UK; [email protected] 2 Department of Clinical and Movement Neurosciences, University College London Institute of Neurology, London WC1N 3BG, UK * Correspondence: [email protected] Abstract: The genetic landscape of Parkinson’s disease (PD) is characterised by rare high penetrance pathogenic variants causing familial disease, genetic risk factor variants driving PD risk in a signif- icant minority in PD cases and high frequency, low penetrance variants, which contribute a small increase of the risk of developing sporadic PD. This knowledge has the potential to have a major impact in the clinical care of people with PD. We summarise these genetic influences and discuss the implications for therapeutics and clinical trial design. Keywords: Parkinson’s disease; genetics; precision medicine; clinical trials; monogenic; polygenic 1. Introduction Parkinson’s disease (PD) is a neurodegenerative condition affecting over 6 million people worldwide that is expected to double in prevalence by 2040 [1]. It is characterised by a core set of movement (motor) abnormalities - slowness of movement, muscle rigidity Citation: Day, J.O.; Mullin, S. The and tremor – as well as a number of non-motor features such as constipation, anxiety and Genetics of Parkinson’s Disease and dementia [2]. There is often a prodromal phase of non-motor symptoms which precede Implications for Clinical Practice. motor symptoms by many years [3]. -
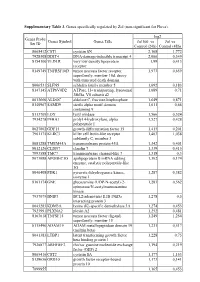
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -

Thesis Submitted in Conformity with the Requirements for the Degree of Doctor of Philosophy Institute of Medical Science University of Toronto
Expression of Parkinson’s disease and associated neurophenotypes in 22q11.2 deletion syndrome by Nancy Jean Butcher A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Institute of Medical Science University of Toronto © Copyright by Nancy Butcher 2015 Expression of Parkinson’s disease and associated neurophenotypes in 22q11.2 deletion syndrome Nancy Jean Butcher Doctor of Philosophy Institute of Medical Science University of Toronto 2015 Abstract The etiology of Parkinson’s disease (PD) remains largely unknown with the exception of a few genetic mutations that affect a small proportion of patients. Case reports suggest that individuals with 22q11.2 deletion syndrome (22q11.2DS), a multisystem genomic disorder associated with hemizygous 22q11.2 deletions, may be at increased risk of early-onset PD. The aim of this thesis was to investigate 22q11.2 deletions as a risk factor for early-onset PD. The prevalence of PD was assessed in a well-characterized cohort of adults with 22q11.2DS. Neuropathological studies were performed in cases with available post-mortem tissue. Whole- genome sequencing was used to investigate the possible contribution of genome-wide rare coding mutations to disease penetrance in neuropathologically confirmed PD cases. Assessment of putative pre-diagnostic clinical and neuroimaging markers of PD in a subset of the older adults (30 to 54 years) provided evidence of an elevated prevalence of pre-morbid motor and olfactory deficits, and of nigrostriatal dopaminergic dysfunction assessed using 11C-dihydrotetrabenazine and positron emission tomography. The clinical context of early-onset PD in this adult cohort of individuals with 22q11.2DS was also investigated. -

Annotation Landscape Analysis with Genecards Arye Harel*1, Aron Inger, Gil Stelzer1, Liora Strichman-Almashanu1, Irina Dalah1, Marilyn Safran1,2 and Doron Lancet1
BMC Bioinformatics BioMed Central Research article Open Access GIFtS: annotation landscape analysis with GeneCards Arye Harel*1, Aron Inger, Gil Stelzer1, Liora Strichman-Almashanu1, Irina Dalah1, Marilyn Safran1,2 and Doron Lancet1 Address: 1Department of Molecular Genetics, Weizmann Institute of Science Rehovot 76100, Israel and 2Department of Biological Services (Bioinformatics Unit), Weizmann Institute of Science, Rehovot 76100, Israel Email: Arye Harel* - [email protected]; Aron Inger - [email protected]; Gil Stelzer - [email protected]; Liora Strichman-Almashanu - [email protected]; Irina Dalah - [email protected]; Marilyn Safran - [email protected]; Doron Lancet - [email protected] * Corresponding author Published: 23 October 2009 Received: 22 February 2009 Accepted: 23 October 2009 BMC Bioinformatics 2009, 10:348 doi:10.1186/1471-2105-10-348 This article is available from: http://www.biomedcentral.com/1471-2105/10/348 © 2009 Harel et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Gene annotation is a pivotal component in computational genomics, encompassing prediction of gene function, expression analysis, and sequence scrutiny. Hence, quantitative measures of the annotation landscape constitute a pertinent bioinformatics tool. GeneCards® is a gene-centric compendium of rich annotative information for over 50,000 human gene entries, building upon 68 data sources, including Gene Ontology (GO), pathways, interactions, phenotypes, publications and many more. -

A Common and Two Novel GBA Mutations in Thai Patients with Gaucher Disease
Journal of Human Genetics (2013) 58, 594–599 & 2013 The Japan Society of Human Genetics All rights reserved 1434-5161/13 www.nature.com/jhg ORIGINAL ARTICLE A common and two novel GBA mutations in Thai patients with Gaucher disease Rachaneekorn Tammachote1,2, Siraprapa Tongkobpetch2, Chalurmpon Srichomthong3, Kampon Phipatthanananti2, Suthipong Pungkanon4, Duangrurdee Wattanasirichaigoon5, Kanya Suphapeetiporn2,3 and Vorasuk Shotelersuk2,3 Gaucher disease (GD) is an autosomal recessive disorder caused by mutations in the glucocerebrosidase (GBA) gene, leading to a deficiency of lysosomal b-glucosidase and accumulation of glycosphingolipids in macrophages. We studied five Thai families with GD (four with GD type 1 and one with GD type 2). Using long-template PCR, PCR using specific primers for the functional gene, direct sequencing of all coding regions of GBA and restriction enzyme digestions, all 10 mutant alleles were successfully identified. The common c.1448T4C (p.L483P or L444P) mutation was identified in 60% of mutant alleles. Of the two patients homozygous for the p.L483P (L444P) mutation, one died from hepatic failure at age 16 years and the other died from sepsis at age 12 years. This p.L483P (L444P) mutation was found in four different haplotypes, suggesting that it was a recurrent mutation, not caused by a founder effect. Two novel mutations, a missense (c.1204T4C, p.Y402H), and a termination codon mutation (c.1609T4C, p.X537A) were found. Studies to determine the molecular pathomechanism of the p.X537A mutation, the first of its kind in this gene, showed that it decreased the amount of protein being expressed and the enzymatic activity, while it was still correctly localized. -

Table of Contents
Table of Contents Comparison Page Numbers RF vs Static 2-231 HSS vs Static 232-507 LSS vs Static 508-715 RF vs HSS 716-746 RF vs LSS 747-751 HSS vs LSS 752-764 Fold change Regulation ([RF] vs ([RF] vs Probe Name [Static]) [Static]) Common name Gene Symbol Description Genbank Accession Homo sapiens apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3B (APOBEC3B), A_24_P66027 2.741304 down NM_004900 APOBEC3B mRNA [NM_004900] NM_004900 Homo sapiens vasohibin 1 (VASH1), mRNA A_23_P77000 3.932838 down NM_014909 VASH1 [NM_014909] NM_014909 Homo sapiens transmembrane protein 97 A_24_P151920 2.292691 down NM_014573 TMEM97 (TMEM97), mRNA [NM_014573] NM_014573 CN479126 UI-CF-FN0-afv-i-22-0-UI.s1 UI-CF-FN0 Homo sapiens cDNA clone UI-CF-FN0-afv-i-22-0-UI A_32_P149011 3.661964 up CN479126 CN479126 3', mRNA sequence [CN479126] CN479126 Homo sapiens notchless homolog 1 (Drosophila) (NLE1), transcript variant 2, mRNA A_23_P141315 1.785076 down NM_001014445 NLE1 [NM_001014445] NM_001014445 Homo sapiens ribonucleotide reductase M1 A_23_P87351 1.519893 down NM_001033 RRM1 polypeptide (RRM1), mRNA [NM_001033] NM_001033 AT_ssH_PC_3 2.387579 down AT_ssH_PC_3 AT_ssH_PC_3 Homo sapiens huntingtin interacting protein 2 A_23_P155677 1.640733 up NM_005339 HIP2 (HIP2), mRNA [NM_005339] NM_005339 Homo sapiens chromosome 1 open reading frame A_24_P131173 3.527582 down NM_024709 C1orf115 115 (C1orf115), mRNA [NM_024709] NM_024709 Homo sapiens CCR4-NOT transcription complex, A_23_P110846 1.616577 up NM_004779 CNOT8 subunit 8 (CNOT8), mRNA [NM_004779] NM_004779 Homo sapiens protein kinase, AMP-activated, alpha A_23_P60811 3.629454 down NM_006252 PRKAA2 2 catalytic subunit (PRKAA2), mRNA [NM_006252] NM_006252 Homo sapiens pannexin 1 (PANX1), mRNA A_23_P47155 2.378446 up NM_015368 PANX1 [NM_015368] NM_015368 Homo sapiens chemokine (C-X-C motif) ligand 2 A_23_P315364 11.12551 down NM_002089 CXCL2 (CXCL2), mRNA [NM_002089] NM_002089 Centromere protein P (CENP-P). -

High-Throughput Carrier Screening Using Taqman Allelic Discrimination
High-Throughput Carrier Screening Using TaqMan Allelic Discrimination Anastasia Fedick1,2*, Jing Su2, Chaim Jalas3, Lesley Northrop2, Batsal Devkota2, Josef Ekstein4, Nathan R. Treff1,2 1 Department of Microbiology and Molecular Genetics, University of Medicine and Dentistry of New Jersey-Robert Wood Johnson Medical School, Piscataway, New Jersey, United States of America, 2 Reproductive Medicine Associates of New Jersey, Basking Ridge, New Jersey, United States of America, 3 Bonei Olam, Center for Rare Jewish Genetic Disorders, Brooklyn, New York, United States of America, 4 Dor Yeshorim, The Committee for Prevention of Jewish Diseases, Brooklyn, New York, United States of America Abstract Members of the Ashkenazi Jewish community are at an increased risk for inheritance of numerous genetic diseases such that carrier screening is medically recommended. This paper describes the development and evaluation of 30 TaqMan allelic discrimination qPCR assays for 29 mutations on 2 different high-throughput platforms. Four of these mutations are in the GBA gene and are successfully examined using short amplicons due to the qualitative nature of TaqMan allelic discrimination. Two systems were tested for their reliability (call rate) and consistency with previous diagnoses (diagnostic accuracy) indicating a call rate of 99.04% and a diagnostic accuracy of 100% (+/20.00%) from one platform, and a call rate of 94.66% and a diagnostic accuracy of 93.35% (+/20.29%) from a second for 9,216 genotypes. Results for mutations tested at the expected carrier frequency indicated a call rate of 97.87% and a diagnostic accuracy of 99.96% (+/20.05%). This study demonstrated the ability of a high throughput qPCR methodology to accurately and reliably genotype 29 mutations in parallel. -

Effects of Inhibitors of FGFR3 on Gene Transcription
(19) TZZ 54¥9A_T (11) EP 2 546 359 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 16.01.2013 Bulletin 2013/03 C12Q 1/68 (2006.01) (21) Application number: 12163693.0 (22) Date of filing: 07.12.2006 (84) Designated Contracting States: •Moler,Edward AT BE BG CH CY CZ DE DK EE ES FI FR GB GR Emeryville, CA 94662-8097 (US) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Rowe, Michael SK TR Emeryville, CA 94662-8097 (US) • Stewart, Keith (30) Priority: 08.12.2005 US 748944 P Toronto, Ontario M5G 2M9 (CA) • Trudel, Suzanne (62) Document number(s) of the earlier application(s) in Toronto, Ontario M5G 2M9 (CA) accordance with Art. 76 EPC: 06840156.1 / 1 960 547 (74) Representative: UEXKÜLL & STOLBERG Patentanwälte (71) Applicant: Novartis AG Beselerstrasse 4 4056 Basel (CH) 22607 Hamburg (DE) (72) Inventors: Remarks: •Heise,Carla •This application was filed on 11-04-2012 as a Emeryville, CA 94662-8097 (US) divisional application to the application mentioned •Masih-Khan, Esther under INID code 62. Toronto, Ontario M5G 2M9 (CA) •Claims filed after the date of filing of the application (Rule 68(4) EPC). (54) Effects of inhibitors of FGFR3 on gene transcription (57) Methods of utilizing biomarkers to identify pa- hibition, are measured and identifications or adjustments tients for treatment or to monitor response to treatment may be made accordingly. are taught herein. Alterations in levels of gene expression of the biomarkers, particularly in response to FGFR3 in- EP 2 546 359 A1 Printed by Jouve, 75001 PARIS (FR) EP 2 546 359 A1 Description BACKGROUND OF THE INVENTION 5 [0001] The present invention relates generally to the field of pharmacogenomics and in particular to the use of biomar- kers for identifying patients suitable for treatment as well as to methods of following their response to methods of treatment.