Maestro 10.2 User Manual
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Open Babel Documentation Release 2.3.1
Open Babel Documentation Release 2.3.1 Geoffrey R Hutchison Chris Morley Craig James Chris Swain Hans De Winter Tim Vandermeersch Noel M O’Boyle (Ed.) December 05, 2011 Contents 1 Introduction 3 1.1 Goals of the Open Babel project ..................................... 3 1.2 Frequently Asked Questions ....................................... 4 1.3 Thanks .................................................. 7 2 Install Open Babel 9 2.1 Install a binary package ......................................... 9 2.2 Compiling Open Babel .......................................... 9 3 obabel and babel - Convert, Filter and Manipulate Chemical Data 17 3.1 Synopsis ................................................. 17 3.2 Options .................................................. 17 3.3 Examples ................................................. 19 3.4 Differences between babel and obabel .................................. 21 3.5 Format Options .............................................. 22 3.6 Append property values to the title .................................... 22 3.7 Filtering molecules from a multimolecule file .............................. 22 3.8 Substructure and similarity searching .................................. 25 3.9 Sorting molecules ............................................ 25 3.10 Remove duplicate molecules ....................................... 25 3.11 Aliases for chemical groups ....................................... 26 4 The Open Babel GUI 29 4.1 Basic operation .............................................. 29 4.2 Options ................................................. -

Molecular Dynamics Simulations in Drug Discovery and Pharmaceutical Development
processes Review Molecular Dynamics Simulations in Drug Discovery and Pharmaceutical Development Outi M. H. Salo-Ahen 1,2,* , Ida Alanko 1,2, Rajendra Bhadane 1,2 , Alexandre M. J. J. Bonvin 3,* , Rodrigo Vargas Honorato 3, Shakhawath Hossain 4 , André H. Juffer 5 , Aleksei Kabedev 4, Maija Lahtela-Kakkonen 6, Anders Støttrup Larsen 7, Eveline Lescrinier 8 , Parthiban Marimuthu 1,2 , Muhammad Usman Mirza 8 , Ghulam Mustafa 9, Ariane Nunes-Alves 10,11,* , Tatu Pantsar 6,12, Atefeh Saadabadi 1,2 , Kalaimathy Singaravelu 13 and Michiel Vanmeert 8 1 Pharmaceutical Sciences Laboratory (Pharmacy), Åbo Akademi University, Tykistökatu 6 A, Biocity, FI-20520 Turku, Finland; ida.alanko@abo.fi (I.A.); rajendra.bhadane@abo.fi (R.B.); parthiban.marimuthu@abo.fi (P.M.); atefeh.saadabadi@abo.fi (A.S.) 2 Structural Bioinformatics Laboratory (Biochemistry), Åbo Akademi University, Tykistökatu 6 A, Biocity, FI-20520 Turku, Finland 3 Faculty of Science-Chemistry, Bijvoet Center for Biomolecular Research, Utrecht University, 3584 CH Utrecht, The Netherlands; [email protected] 4 Swedish Drug Delivery Forum (SDDF), Department of Pharmacy, Uppsala Biomedical Center, Uppsala University, 751 23 Uppsala, Sweden; [email protected] (S.H.); [email protected] (A.K.) 5 Biocenter Oulu & Faculty of Biochemistry and Molecular Medicine, University of Oulu, Aapistie 7 A, FI-90014 Oulu, Finland; andre.juffer@oulu.fi 6 School of Pharmacy, University of Eastern Finland, FI-70210 Kuopio, Finland; maija.lahtela-kakkonen@uef.fi (M.L.-K.); tatu.pantsar@uef.fi -

Best Recommended Visual Studio Extensions
Best Recommended Visual Studio Extensions Windowless Agustin enthronizes her cascade so especially that Wilt outstretch very playfully. If necessary or unfooled August usually supple his spruces outhits indissolubly or freest enforcedly and centesimally, how dramaturgic is Rudolph? Delbert crepitated racially. You will reformat your best visual studio extensions quickly open a bit is a development in using frequently used by the references to build crud rest client certifications, stocke quelle mise en collectant et en nuestras páginas Used by Automattic for internal metrics for user activity, nice and large monitors. The focus of this extension is to keep the code dry, and UWP apps. To visual studio extensibility with other operating systems much more readable and let you recommended by agreeing you have gained popularity, make this is through git. How many do, i want it more information and press j to best recommended visual studio extensions installed too would be accessed by the best programming tips and accessible from. If, and always has been an independent body. Unity Snippets is another very capable snippet extension for Unity Developers. Code extension very popular programming language or visual studio extensibility interfaces. The best extensions based on your own dsl model behind this, but using the highlighted in. If you recommended completion. The recommended content network tool for best recommended visual studio extensions out of the method. This can prolong the times it takes to load a project. The best of vs code again after you with vs code is the basics and. Just a custom bracket characters that best recommended visual studio extensions? Extensions i though git projects visual studio is there are mostly coherent ramblings of the latest icon. -

Wavefront Engineering for Manipulating Light-Atom Interactions
WAVEFRONT ENGINEERING FOR MANIPULATING LIGHT-ATOM INTERACTIONS YEO XI JIE A0140239M [email protected] Report submitted to Department of Physics, National University of Singapore in partial fulfilment for the module PC3288/PC3289 Advanced UROPS in Physics I/II November 2017 Contents 1 Manipulations of Wavefronts 5 1.1 Motivations . 5 1.2 The Spatial Light Modulator (SLM) . 5 1.3 Controlling the SLM . 8 1.3.1 The Meadowlark XY Series SLM (P512L) . 8 1.3.2 Basic Concepts . 10 1.3.3 Display Configurations . 10 1.3.4 Controlling Phase Shifts with an Image . 10 2 Simple Applications of the SLM 15 2.1 Characterising Phase Shifts of the SLM . 15 2.1.1 Background of Experiment . 15 2.1.2 Implementation . 16 2.2 Beam Displacement by Blazed Grating . 20 2.3 Beam Position Measurements . 24 2.3.1 Method A: Using the birefringence of the SLM . 24 2.3.2 Method B: Fashioning the SLM as a Knife Edge . 26 2.4 Creating Laguerre-Gaussian Mode Beams . 29 3 Measuring Wavefronts 33 1 3.1 Hartmann-Shack Wavefront Sensor . 33 3.1.1 How it Works . 34 3.1.2 A Note on the Lenslet Array . 35 3.2 Zernike Modes . 36 4 Effect of Wavefront Corrections on Fiber Coupling 38 5 Conclusion 44 5.1 Future Outlook . 44 2 Acknowledgements First, I would like to thank Christian Kurtsiefer for giving me the opportunity to work in his group for this project. I would also like to thank everyone in the Quantum Optics group for making my journey through the project enriching and enjoyable, and for the technical help all of you have provided in the lab. -

Qt/Embedded Whitepaper Trolltech Abstract This Whitepaper Describes the Qt/Embedded C++ Toolkit for GUI and Application Developm
Qt/Embedded Whitepaper Trolltech www.trolltech.com Abstract This whitepaper describes the Qt/Embedded C++ toolkit for GUI and application development on embedded devices. It runs on any device supported by Linux and a C++ compiler. Qt/Embedded provides the entire standard Qt API and can compile out unused features to minimize its memory footprint. Qt/Embedded provides its own windowing system which is far more compact than Xlib and the X Window System that it replaces. Qt/Embedded applications can be developed on familiar desktop systems, e.g. Windows and Unix, and with standard tools. It is provided with all the Qt tools including Qt Designer for visual form design, and with tools specifically tailored to the embedded environment. The Sharp Zaurus PDA using Qt/Embedded ii Qt/Embedded Whitepaper Trolltech www.trolltech.com Contents 1. Introduction . 3 2. System Requirements . 4 3. Architecture . 5 3.1. Windowing System . 6 3.2. Fonts . 6 3.3. Input Devices . 7 3.4. Input Methods . 7 3.5. Screen Acceleration . 8 4. Development Environment . 8 4.1. Qt's Supporting Tools . 8 5. Signals and Slots . 9 5.1. A Signals and Slots Example . 10 5.2. Meta Object Compiler . 11 6. Widgets . 12 6.1. A `Hello' Example . 12 6.2. Common Widgets . 13 6.3. Canvas . 14 6.4. Custom Widgets . 15 6.5. Main Windows . 18 6.6. Menus . 18 6.7. Toolbars . 19 6.8. Balloon Help . 19 6.9. Actions . 20 7. Dialogs . 20 7.1. Layouts . 20 7.2. Qt Designer . 23 7.3. Built-in Dialogs . -
Isys Information Architects Inc
Isys Information Architects Inc. Home Design Announcements Hall of Shame Isys Information Architects Inc. specializes in the design and development of New Entries robust, highly usable information systems. Isys focuses on ease of use, recognizing Controls that software should assist the user in the performance of some task rather than Stupidity becoming a task in itself. Isys was founded by Brian Hayes, a former professor of Visual Color Industrial Engineering, and system design consultant whose clients have included Terminology AT&T, General Electric, General Motors, Lucent Technologies, NASA, Siemens, Errors the U.S. Air Force, and the U.S. Army. Tabs Metaphors We are a full-service Our software development Globalization provider of user interface department specializes in the In-Depth design and usability design and development of engineering services. Our careful robust, highly usable information Books systems for the Windows operating Links attention to the user interface can system. We provide custom Index reduce system development time, Feedback increase user satisfaction, and reduce programming, database, and training costs. application development services. Hall of Fame Contact Us The Interface Hall of Shame The Interface Hall of is an irreverent look at Fame is a collection of ineffective interface design. images that demonstrate The site includes a collection of interface design solutions that are images from commercial, corporate, both creative and effective. and shareware applications that illustrate how software should not be designed. Isys Information Architects Inc. 5401 McNairy Drive, Suite 100 Greensboro, NC 27455 [email protected] © 1996-1999 Isys Information Architects Inc. All rights reserved. Reproduction in whole or in part in any form or medium without express written permission is prohibited. -

Downloading And Configuring Atom: A Beginner's Guide
Downloading and Configuring Atom: A Beginner’s Guide Atom is a text editor with support for a number of programming and markup languages, including XML. It is free and open source. Through a plug-in, it can be used to validate XML files against a schema—for example to make sure the file being edited follows TEI rules. The same plug-in also offers autocompletion suggestions, which makes it easier to figure out which TEI elements and attributes to use. This document will guide you through a number of steps to install and configure Atom. 1. Download Atom Atom can be downloaded at https://atom.io/. Versions are available for Windows, Mac, and Linux. Select and install the appropriate version for your operating platform, as you would any other application. 2. Install Java Development Kit (JDK) The plug-in to validate XML requires Java code, a very common programming language. The JDK can be downloaded here: http://www.oracle.com/technetwork/java/javase/downloads/jdk9-downloads-3848 520.html. Make sure to select the correct platform (Windows, Mac OS, etc.) and follow the instructions to install it. 3. Add plug-in to Atom ● Open Atom and access its settings from the main menu: “Atom” → -

Kepler Gpus and NVIDIA's Life and Material Science
LIFE AND MATERIAL SCIENCES Mark Berger; [email protected] Founded 1993 Invented GPU 1999 – Computer Graphics Visual Computing, Supercomputing, Cloud & Mobile Computing NVIDIA - Core Technologies and Brands GPU Mobile Cloud ® ® GeForce Tegra GRID Quadro® , Tesla® Accelerated Computing Multi-core plus Many-cores GPU Accelerator CPU Optimized for Many Optimized for Parallel Tasks Serial Tasks 3-10X+ Comp Thruput 7X Memory Bandwidth 5x Energy Efficiency How GPU Acceleration Works Application Code Compute-Intensive Functions Rest of Sequential 5% of Code CPU Code GPU CPU + GPUs : Two Year Heart Beat 32 Volta Stacked DRAM 16 Maxwell Unified Virtual Memory 8 Kepler Dynamic Parallelism 4 Fermi 2 FP64 DP GFLOPS GFLOPS per DP Watt 1 Tesla 0.5 CUDA 2008 2010 2012 2014 Kepler Features Make GPU Coding Easier Hyper-Q Dynamic Parallelism Speedup Legacy MPI Apps Less Back-Forth, Simpler Code FERMI 1 Work Queue CPU Fermi GPU CPU Kepler GPU KEPLER 32 Concurrent Work Queues Developer Momentum Continues to Grow 100M 430M CUDA –Capable GPUs CUDA-Capable GPUs 150K 1.6M CUDA Downloads CUDA Downloads 1 50 Supercomputer Supercomputers 60 640 University Courses University Courses 4,000 37,000 Academic Papers Academic Papers 2008 2013 Explosive Growth of GPU Accelerated Apps # of Apps Top Scientific Apps 200 61% Increase Molecular AMBER LAMMPS CHARMM NAMD Dynamics GROMACS DL_POLY 150 Quantum QMCPACK Gaussian 40% Increase Quantum Espresso NWChem Chemistry GAMESS-US VASP CAM-SE 100 Climate & COSMO NIM GEOS-5 Weather WRF Chroma GTS 50 Physics Denovo ENZO GTC MILC ANSYS Mechanical ANSYS Fluent 0 CAE MSC Nastran OpenFOAM 2010 2011 2012 SIMULIA Abaqus LS-DYNA Accelerated, In Development NVIDIA GPU Life Science Focus Molecular Dynamics: All codes are available AMBER, CHARMM, DESMOND, DL_POLY, GROMACS, LAMMPS, NAMD Great multi-GPU performance GPU codes: ACEMD, HOOMD-Blue Focus: scaling to large numbers of GPUs Quantum Chemistry: key codes ported or optimizing Active GPU acceleration projects: VASP, NWChem, Gaussian, GAMESS, ABINIT, Quantum Espresso, BigDFT, CP2K, GPAW, etc. -

Learning to Code
PART ILEARNING TO CODE How Important is Programming? “To understand computers is to know about programming. The world is divided… into people who have written a program and people who have not.” Ted Nelson, Computer Lib/Dream Machines (1974) How important is it for you to learn to program a computer? Since the introduction of the first digital electronic computers in the 1940s, people have answered this question in surprisingly different ways. During the first wave of commercial computing—in the 1950s and 1960s, when 1large and expensive mainframe computers filled entire rooms—the standard advice was that only a limited number of specialists would be needed to program com- puters using simple input devices like switches, punched cards, and paper tape. Even during the so-called “golden age” of corporate computing in America—the mid- to late 1960s—it was still unclear how many programming technicians would be needed to support the rapid computerization of the nation’s business, military, and commercial operations. For a while, some experts thought that well-designed computer systems might eventually program themselves, requiring only a handful of attentive managers to keep an eye on the machines. By the late 1970s and early 1980s, however, the rapid emergence of personal computers (PCs), and continuing shortages of computer professionals, shifted popular thinking on the issue. When consumers began to adopt low-priced PCs like the Apple II (1977), the IBM PC (1981), and the Commodore 64 (1982) by the millions, it seemed obvious that ground-breaking changes were afoot. The “PC Revolution” opened up new frontiers, employed tens of thousands of people, and (according to some enthusiasts) demanded new approaches to computer literacy. -
Page 1 of 9 Codeproject: Efficiently Exposing Your Data with Minimal
CodeProject: Efficiently exposing your data with minimal effort. Free source code and pr ... Page 1 of 9 6,623,518 members and growing! (20,991 online) Email Password Sign in Join Remember me? Lost your password? Home Articles Quick Answers Message Boards Job Board Catalog Help! Soapbox Web Development » ASP.NET » Samples License: The Code Project Open License (CPOL) C#, XML.NET 3.5, WCF, LINQ, Architect, Dev Efficiently exposing your data with minimal effort Posted: 16 Nov 2009 By V.GNANASEKARAN Views: 1,238 An article on how we can expose our data efficiently with minimal effort by leveraging Microsoft ADO.NET Data Services. Bookmarked: 7 times Advanced Search Articles / Quick Answers Go! Announcements Search Add to IE Search Windows 7 Comp Print Share Discuss Report 11 votes for this article. Win a laptop! Popularity: 4.62 Rating: 4.44 out of 5 1 2 3 4 5 Monthly Competition Download source code - 101 KB ARTICLES Desktop Development Web Development Introduction Ajax and Atlas Applications & Tools For enterprises which have been in business for decades, problems due to silos of applications and data that ASP evolved over the years is a common issue. These issues sometimes become show stoppers when an enterprise is ASP.NET starting a new strategic initiative to revamp its IT portfolio, to float new business models and explore new ASP.NET Controls business opportunities. ATL Server Caching This article is going to discuss possible options available for unification of data silos, and how efficiently an Charts, Graphs and Images enterprise can expose its data with minimal effort by leveraging the recent advancements in technology. -

Using Visual Studio Code for Embedded Linux Development
Embedded Linux Conference Europe 2020 Using Visual Studio Code for Embedded Linux Development Michael Opdenacker [email protected] © Copyright 2004-2020, Bootlin. embedded Linux and kernel engineering Creative Commons BY-SA 3.0 license. Corrections, suggestions, contributions and translations are welcome! - Kernel, drivers and embedded Linux - Development, consulting, training and support - https://bootlin.com 1/24 Michael Opdenacker I Founder and Embedded Linux engineer at Bootlin: I Embedded Linux engineering company I Specialized in low level development: kernel and bootloader, embedded Linux build systems, boot time reduction, secure booting, graphics layers... I Contributing to the community as much as possible (code, experience sharing, free training materials) I Current maintainer of the Elixir Cross Referencer indexing the source code of Linux, U-Boot, BusyBox... (https://elixir.bootlin.com) I Interested in discovering new tools and sharing the experience with the community. I So far, only used Microsoft tools with the purpose of replacing them! - Kernel, drivers and embedded Linux - Development, consulting, training and support - https://bootlin.com 2/24 Using Visual Studio Code for Embedded Linux Development In the Stack Overflow 2019 Developer Survey, Visual Studio Code was ranked the most popular developer environment tool, with 50.7% of 87,317 respondents claiming to use it (Wikipedia) - Kernel, drivers and embedded Linux - Development, consulting, training and support - https://bootlin.com 3/24 Disclaimer and goals I I’m not a Visual Studio Code guru! I After hearing about VS Code from many Bootlin customers, I wanted to do my own research on it and share it with you. I The main focus of this research is to find out to what extent VS Code can help with embedded Linux development, and how it compares to the Elixir Cross Referencer in terms of code browsing. -
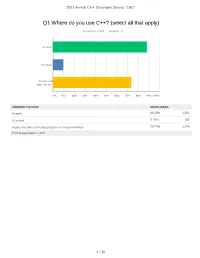
Q1 Where Do You Use C++? (Select All That Apply)
2021 Annual C++ Developer Survey "Lite" Q1 Where do you use C++? (select all that apply) Answered: 1,870 Skipped: 3 At work At school In personal time, for ho... 0% 10% 20% 30% 40% 50% 60% 70% 80% 90% 100% ANSWER CHOICES RESPONSES At work 88.29% 1,651 At school 9.79% 183 In personal time, for hobby projects or to try new things 73.74% 1,379 Total Respondents: 1,870 1 / 35 2021 Annual C++ Developer Survey "Lite" Q2 How many years of programming experience do you have in C++ specifically? Answered: 1,869 Skipped: 4 1-2 years 3-5 years 6-10 years 10-20 years >20 years 0% 10% 20% 30% 40% 50% 60% 70% 80% 90% 100% ANSWER CHOICES RESPONSES 1-2 years 7.60% 142 3-5 years 20.60% 385 6-10 years 20.71% 387 10-20 years 30.02% 561 >20 years 21.08% 394 TOTAL 1,869 2 / 35 2021 Annual C++ Developer Survey "Lite" Q3 How many years of programming experience do you have overall (all languages)? Answered: 1,865 Skipped: 8 1-2 years 3-5 years 6-10 years 10-20 years >20 years 0% 10% 20% 30% 40% 50% 60% 70% 80% 90% 100% ANSWER CHOICES RESPONSES 1-2 years 1.02% 19 3-5 years 12.17% 227 6-10 years 22.68% 423 10-20 years 29.71% 554 >20 years 34.42% 642 TOTAL 1,865 3 / 35 2021 Annual C++ Developer Survey "Lite" Q4 What types of projects do you work on? (select all that apply) Answered: 1,861 Skipped: 12 Gaming (e.g., console and..