Early Growth Response 1 Acts As a Tumor Suppressor in Vivo and in Vitro Via Regulation of P53
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

New Approaches to Functional Process Discovery in HPV 16-Associated Cervical Cancer Cells by Gene Ontology
Cancer Research and Treatment 2003;35(4):304-313 New Approaches to Functional Process Discovery in HPV 16-Associated Cervical Cancer Cells by Gene Ontology Yong-Wan Kim, Ph.D.1, Min-Je Suh, M.S.1, Jin-Sik Bae, M.S.1, Su Mi Bae, M.S.1, Joo Hee Yoon, M.D.2, Soo Young Hur, M.D.2, Jae Hoon Kim, M.D.2, Duck Young Ro, M.D.2, Joon Mo Lee, M.D.2, Sung Eun Namkoong, M.D.2, Chong Kook Kim, Ph.D.3 and Woong Shick Ahn, M.D.2 1Catholic Research Institutes of Medical Science, 2Department of Obstetrics and Gynecology, College of Medicine, The Catholic University of Korea, Seoul; 3College of Pharmacy, Seoul National University, Seoul, Korea Purpose: This study utilized both mRNA differential significant genes of unknown function affected by the display and the Gene Ontology (GO) analysis to char- HPV-16-derived pathway. The GO analysis suggested that acterize the multiple interactions of a number of genes the cervical cancer cells underwent repression of the with gene expression profiles involved in the HPV-16- cancer-specific cell adhesive properties. Also, genes induced cervical carcinogenesis. belonging to DNA metabolism, such as DNA repair and Materials and Methods: mRNA differential displays, replication, were strongly down-regulated, whereas sig- with HPV-16 positive cervical cancer cell line (SiHa), and nificant increases were shown in the protein degradation normal human keratinocyte cell line (HaCaT) as a con- and synthesis. trol, were used. Each human gene has several biological Conclusion: The GO analysis can overcome the com- functions in the Gene Ontology; therefore, several func- plexity of the gene expression profile of the HPV-16- tions of each gene were chosen to establish a powerful associated pathway, identify several cancer-specific cel- cervical carcinogenesis pathway. -

Datasheet: VMA00210 Product Details
Datasheet: VMA00210 Description: MOUSE ANTI NAB1 Specificity: NAB1 Format: Purified Product Type: PrecisionAb™ Monoclonal Clone: OTI1D2 Isotype: IgG1 Quantity: 100 µl Product Details Applications This product has been reported to work in the following applications. This information is derived from testing within our laboratories, peer-reviewed publications or personal communications from the originators. Please refer to references indicated for further information. For general protocol recommendations, please visit www.bio-rad-antibodies.com/protocols. Yes No Not Determined Suggested Dilution Western Blotting 1/1000 PrecisionAb antibodies have been extensively validated for the western blot application. The antibody has been validated at the suggested dilution. Where this product has not been tested for use in a particular technique this does not necessarily exclude its use in such procedures. Further optimization may be required dependant on sample type. Target Species Human Product Form Purified IgG - liquid Preparation Mouse monoclonal antibody purified by affinity chromatography from ascites Buffer Solution Phosphate buffered saline Preservative 0.09% Sodium Azide (NaN3) Stabilisers 1% Bovine Serum Albumin 50% Glycerol Immunogen Recombinant protein corresponding to amino acids 237-487 of human NAB1 (NP_005957) produced in E. coli External Database UniProt: Links Q13506 Related reagents Entrez Gene: 4664 NAB1 Related reagents Specificity Mouse anti Human NAB1 antibody recognizes the NGFI-A-binding protein 1, also known as Page 1 of 2 EGR-1-binding protein 1, EGR1 binding protein 1 and transcriptional regulatory protein p54. Mouse anti Human NAB1 antibody detects a band of 54 kDa. The antibody has been extensively validated for western blotting using whole cell lysates. Western Blotting Anti NAB1 detects a band of approximately 54 kDa in Jurkat cell lysates Instructions For Use Please refer to the PrecisionAb western blotting protocol. -

The Transcription Factor Egr1 Is a Direct Regulator of Multiple Tumor
Cancer Gene Therapy (2006) 13, 115–124 r 2006 Nature Publishing Group All rights reserved 0929-1903/06 $30.00 www.nature.com/cgt REVIEW The transcription factor Egr1 is a direct regulator of multiple tumor suppressors including TGFb1, PTEN, p53, and fibronectin V Baron1, ED Adamson2, A Calogero3, G Ragona3 and D Mercola1,4,5 1Sidney Kimmel Cancer Center, San Diego, CA, USA; 2The Burnham Institute, La Jolla, CA, USA; 3The University of Rome, Rome, Italy; 4The Rebecca and John Moores Cancer Center, University of California at San Diego, La Jolla, CA, USA and 5The Department of Pathology, University of California at Irvine, Irvine, CA, USA Recent studies are reviewed indicating that the transcription factor early growth response-1 (Egr1) is a direct regulator of multiple tumor suppressors including TGFb1, PTEN, p53, and fibronectin. The downstream pathways of these factors display multiple nodes of interaction with each other, suggesting the existence of a functional network of suppressor factors that serve to maintain normal growth regulation and resist the emergence of transformed variants. Paradoxically, Egr1 is oncogenic in prostate cancer. In the majority of these cancers, PTEN or p53 is inactive. It is suggested that these defects in the suppressor network allow for the unopposed induction of TGFb1 and fibronectin, which favor transformation and survival of prostate tumor epithelial cells, and explain the role of Egr1 in prostate cancer. Egr1 is a novel and logical target for intervention by gene therapy methods, and targeting methods are discussed. Cancer Gene Therapy (2006) 13, 115–124. doi:10.1038/sj.cgt.7700896; published online 2 September 2005 Keywords: suppressor network; anoikis; systems biology; prostate cancer; tumor progression Introduction products. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

A Dissertation Entitled the Androgen Receptor
A Dissertation entitled The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit Submitted to the Graduate Faculty as partial fulfillment of the requirements for the PhD Degree in Biomedical science Dr. Manohar Ratnam, Committee Chair Dr. Lirim Shemshedini, Committee Member Dr. Robert Trumbly, Committee Member Dr. Edwin Sanchez, Committee Member Dr. Beata Lecka -Czernik, Committee Member Dr. Patricia R. Komuniecki, Dean College of Graduate Studies The University of Toledo August 2011 Copyright 2011, Mesfin Gonit This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. An Abstract of The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit As partial fulfillment of the requirements for the PhD Degree in Biomedical science The University of Toledo August 2011 Prostate cancer depends on the androgen receptor (AR) for growth and survival even in the absence of androgen. In the classical models of gene activation by AR, ligand activated AR signals through binding to the androgen response elements (AREs) in the target gene promoter/enhancer. In the present study the role of AREs in the androgen- independent transcriptional signaling was investigated using LP50 cells, derived from parental LNCaP cells through extended passage in vitro. LP50 cells reflected the signature gene overexpression profile of advanced clinical prostate tumors. The growth of LP50 cells was profoundly dependent on nuclear localized AR but was independent of androgen. Nevertheless, in these cells AR was unable to bind to AREs in the absence of androgen. -

Quantitative SUMO Proteomics Reveals the Modulation of Several
www.nature.com/scientificreports OPEN Quantitative SUMO proteomics reveals the modulation of several PML nuclear body associated Received: 10 October 2017 Accepted: 28 March 2018 proteins and an anti-senescence Published: xx xx xxxx function of UBC9 Francis P. McManus1, Véronique Bourdeau2, Mariana Acevedo2, Stéphane Lopes-Paciencia2, Lian Mignacca2, Frédéric Lamoliatte1,3, John W. Rojas Pino2, Gerardo Ferbeyre2 & Pierre Thibault1,3 Several regulators of SUMOylation have been previously linked to senescence but most targets of this modifcation in senescent cells remain unidentifed. Using a two-step purifcation of a modifed SUMO3, we profled the SUMO proteome of senescent cells in a site-specifc manner. We identifed 25 SUMO sites on 23 proteins that were signifcantly regulated during senescence. Of note, most of these proteins were PML nuclear body (PML-NB) associated, which correlates with the increased number and size of PML-NBs observed in senescent cells. Interestingly, the sole SUMO E2 enzyme, UBC9, was more SUMOylated during senescence on its Lys-49. Functional studies of a UBC9 mutant at Lys-49 showed a decreased association to PML-NBs and the loss of UBC9’s ability to delay senescence. We thus propose both pro- and anti-senescence functions of protein SUMOylation. Many cellular mechanisms of defense have evolved to reduce the onset of tumors and potential cancer develop- ment. One such mechanism is cellular senescence where cells undergo cell cycle arrest in response to various stressors1,2. Multiple triggers for the onset of senescence have been documented. While replicative senescence is primarily caused in response to telomere shortening3,4, senescence can also be triggered early by a number of exogenous factors including DNA damage, elevated levels of reactive oxygen species (ROS), high cytokine signa- ling, and constitutively-active oncogenes (such as H-RAS-G12V)5,6. -

Genomics Analysis of Potassium Channel Genes in Songbirds Reveals
Lovell et al. BMC Genomics 2013, 14:470 http://www.biomedcentral.com/1471-2164/14/470 RESEARCH ARTICLE Open Access Genomics analysis of potassium channel genes in songbirds reveals molecular specializations of brain circuits for the maintenance and production of learned vocalizations Peter V Lovell, Julia B Carleton and Claudio V Mello* Abstract Background: A fundamental question in molecular neurobiology is how genes that determine basic neuronal properties shape the functional organization of brain circuits underlying complex learned behaviors. Given the growing availability of complete vertebrate genomes, comparative genomics represents a promising approach to address this question. Here we used genomics and molecular approaches to study how ion channel genes influence the properties of the brain circuitry that regulates birdsong, a learned vocal behavior with important similarities to human speech acquisition. We focused on potassium (K-)Channels, which are major determinants of neuronal cell excitability. Starting with the human gene set of K-Channels, we used cross-species mRNA/protein alignments, and syntenic analysis to define the full complement of orthologs, paralogs, allelic variants, as well as novel loci not previously predicted in the genome of zebra finch (Taeniopygia guttata). We also compared protein coding domains in chicken and zebra finch orthologs to identify genes under positive selective pressure, and those that contained lineage-specific insertions/deletions in functional domains. Finally, we conducted comprehensive in situ hybridizations to determine the extent of brain expression, and identify K-Channel gene enrichments in nuclei of the avian song system. Results: We identified 107 K-Channel finch genes, including 6 novel genes common to non-mammalian vertebrate lineages. -
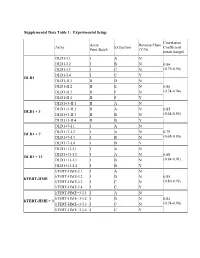
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Egr1 (NM 007913) Mouse Tagged ORF Clone – MG227136 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for MG227136 Egr1 (NM_007913) Mouse Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Egr1 (NM_007913) Mouse Tagged ORF Clone Tag: TurboGFP Symbol: Egr1 Synonyms: A530045N19Rik; egr; Egr-1; ETR103; Krox-1; Krox-24; Krox24; NGF1-A; NGFI-A; NGFIA; TIS8; Zenk; Zfp-6 Vector: pCMV6-AC-GFP (PS100010) E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: Neomycin This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 4 Egr1 (NM_007913) Mouse Tagged ORF Clone – MG227136 ORF Nucleotide >MG227136 representing NM_007913 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) GACGTTGTATACGACTCCTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGGCGCGCCC ATGGCAGCGGCCAAGGCCGAGATGCAATTGATGTCTCCGCTGCAGATCTCTGACCCGTTCGGCTCCTTTC CTCACTCACCCACCATGGACAACTACCCCAAACTGGAGGAGATGATGCTGCTGAGCAACGGGGCTCCCCA GTTCCTCGGTGCTGCCGGAACCCCAGAGGGCAGCGGCGGTAATAGCAGCAGCAGCACCAGCAGCGGGGGC GGTGGTGGGGGCGGCAGCAACAGCGGCAGCAGCGCCTTCAATCCTCAAGGGGAGCCGAGCGAACAACCCT ATGAGCACCTGACCACAGAGTCCTTTTCTGACATCGCTCTGAATAATGAGAAGGCGATGGTGGAGACGAG TTATCCCAGCCAAACGACTCGGTTGCCTCCCATCACCTATACTGGCCGCTTCTCCCTGGAGCCCGCACCC AACAGTGGCAACACTTTGTGGCCTGAACCCCTTTTCAGCCTAGTCAGTGGCCTCGTGAGCATGACCAATC CTCCGACCTCTTCATCCTCGGCGCCTTCTCCAGCTGCTTCATCGTCTTCCTCTGCCTCCCAGAGCCCGCC -

Budesonide Epimer R Or Dexamethasone Selectively
Proc. Natl. Acad. Sci. USA Vol. 95, pp. 3914–3919, March 1998 Medical Sciences Budesonide epimer R or dexamethasone selectively inhibit platelet-activating factor-induced or interleukin 1b-induced DNA binding activity of cis-acting transcription factors and cyclooxygenase-2 gene expression in human epidermal keratinocytes WALTER J. LUKIW*, RICARDO PALACIOS PELAEZ†,JORGE MARTINEZ†, AND NICOLAS G. BAZAN*‡ *Louisiana State University Medical Center, School of Medicine, Neuroscience Center of Excellence and Department of Ophthalmology, 2020 Gravier Street, Suite D, New Orleans, LA 70112-2272; and †Research and Development Department, IFIDESA-ARISTEGUI, Industrial Farmaceutica y de Especialidades SA, Maria de Molina 37, Madrid 28006 Spain Communicated by Ralph T. Holman, University of Minnesota, Austin, MN, January 2, 1998 (received for review July 23, 1997) ABSTRACT To further understand the molecular mecha- cytokines, matrix metalloproteinases, and cell adhesion molecules nism of glucocorticoid action on gene expression, DNA-binding known to be critical to both inflammation and the immune activities of the cis-acting transcription factors activator protein response (1, 2). Besides a direct “type 1” interaction with a 1 (AP1), AP2, Egr1 (zif268), NF-kB, the signal transducers and palindromic GC responsive element in GC-sensitive gene pro- activators of transcription proteins gamma interferon activation moters (3, 4), functions of the GC receptor also include a “type site (GAS), Sis-inducible element, and the TATA binding protein 2” interaction with chromatin-associated cis-acting transcription transcription factor II D (TFIID) were examined in human factors. For example, the GC receptor can physically interact with epidermal keratinocytes. The cytokine interleukin 1b (IL-1b) the Fos and Jun components of dimeric activator protein 1 (AP1) and platelet-activating factor (PAF), both potent mediators of (5, 6), the p65 subunit of NF-kB (7, 8), and the STAT5 family of inflammation, were used as triggers for gene expression. -

Additional Table 3 Transcription Factors (Tfs) Information Total Tfs
Additional Table 3 Transcription factors (TFs) information Total TFs 24 h TFs 48 h TFs TFs Target Genes 18 5 17 Cebpa Cebpa Cebpa Egr2 Cebpa Cebpa Cdkn1a Cebpb Stat3 Cebpb Cebpa Afp Egr1 Wt1 Egr2 Cebpa Gnrh1 Egr2 Tp53 Ets1 Cebpa Got1 Ets1 Egr1 Pax8 Cebpa Cpb2 Hif1a Pparg Cebpa Cyp2c12 Hoxa1 Rela Cebpa Klf5 Junb Sp4 Cebpa A2m Pax8 Stat1 Cebpa Thrsp Pparg Stat3 Cebpa Orm1 Rela Tp53 Cebpa Pla2g2a Sp4 Usf1 Cebpa Arg1 Stat1 Usf2 Cebpa Mmp2 Stat3 Hif1a Cebpa Tat Tp53 Hoxa1 Cebpa Xdh Usf1 Junb Cebpa Spin2a Usf2 Wt1 Cebpa Sst Wt1 Cebpa Mt1a Cebpa Ptgs2 Cebpa Slc2a2 Cebpa Fabp2 Cebpa Adh1 Cebpa Ucp1 Cebpb Cebpa Cebpb Adh1 Cebpb Cyp2c12 Cebpb Cgm3 Cebpb Arg1 Cebpb A2m Cebpb Ccl2 Cebpb Pla2g2a Cebpb Col1a1 Cebpb Cyp7a1 Cebpb Sst Cebpb Xdh Cebpb Ptgs2 Cebpb Ddit3 Cebpb Pdgfra Cebpb Ucp1 Cebpb Afp Cebpb Cpb2 Cebpb Star Cebpb Prl Cebpb Slc2a2 Egr1 Lhb Egr1 Adrb1 Egr1 Pgy1 Egr1 Th Egr1 Dlk1 Egr1 Chrna7 Egr1 Cntn1 Egr1 Slc9a3 Egr1 Pdgfb Egr1 Aldoc Egr1 Me1 Egr1 Pdgfa Egr1 Ptp4a1 Egr1 Serpine1 Egr1 Atp2a2 Egr1 Lhcgr Egr1 Mmp14 Egr1 Myh6 Egr1 Nudt6 Egr1 F3 Egr1 Id1 Egr2 Aldoc Ets1 Cited2 Ets1 Thy1 Ets1 Lhb Ets1 Runx2 Ets1 Ccnd1 Ets1 Spp1 Ets1 Casp3 Ets1 Prl Ets1 Mmp2 Ets1 Tat Ets1 Ucp1 Ets1 Ccnb1 Ets1 Cdkn1a Ets1 Mmp9 Hif1a Th Hif1a Wt1 Hif1a Vhl Hif1a Nos2 Hif1a Slc2a1 Hif1a Tfrc Hif1a Nppa Hif1a Serpine1 Hif1a Gck Hif1a Adra1a Hif1a Plat Hif1a Hmox1 Hif1a Mt1a Hif1a Pklr Hif1a Cdkn1a Hoxa1 Col1a1 Junb Th Junb Vgf Junb Timp1 Pax8 Wbp2 Pax8 Slc5a5 Pax8 Tg Pparg Hadha Pparg Ucp2 Pparg Aldh3a1 Pparg Nr1d1 Pparg Mapk1 Pparg Atp2a2 Pparg Ptgs2 Pparg Rxrg Pparg -

POGLUT1, the Putative Effector Gene Driven by Rs2293370 in Primary
www.nature.com/scientificreports OPEN POGLUT1, the putative efector gene driven by rs2293370 in primary biliary cholangitis susceptibility Received: 6 June 2018 Accepted: 13 November 2018 locus chromosome 3q13.33 Published: xx xx xxxx Yuki Hitomi 1, Kazuko Ueno2,3, Yosuke Kawai1, Nao Nishida4, Kaname Kojima2,3, Minae Kawashima5, Yoshihiro Aiba6, Hitomi Nakamura6, Hiroshi Kouno7, Hirotaka Kouno7, Hajime Ohta7, Kazuhiro Sugi7, Toshiki Nikami7, Tsutomu Yamashita7, Shinji Katsushima 7, Toshiki Komeda7, Keisuke Ario7, Atsushi Naganuma7, Masaaki Shimada7, Noboru Hirashima7, Kaname Yoshizawa7, Fujio Makita7, Kiyoshi Furuta7, Masahiro Kikuchi7, Noriaki Naeshiro7, Hironao Takahashi7, Yutaka Mano7, Haruhiro Yamashita7, Kouki Matsushita7, Seiji Tsunematsu7, Iwao Yabuuchi7, Hideo Nishimura7, Yusuke Shimada7, Kazuhiko Yamauchi7, Tatsuji Komatsu7, Rie Sugimoto7, Hironori Sakai7, Eiji Mita7, Masaharu Koda7, Yoko Nakamura7, Hiroshi Kamitsukasa7, Takeaki Sato7, Makoto Nakamuta7, Naohiko Masaki 7, Hajime Takikawa8, Atsushi Tanaka 8, Hiromasa Ohira9, Mikio Zeniya10, Masanori Abe11, Shuichi Kaneko12, Masao Honda12, Kuniaki Arai12, Teruko Arinaga-Hino13, Etsuko Hashimoto14, Makiko Taniai14, Takeji Umemura 15, Satoru Joshita 15, Kazuhiko Nakao16, Tatsuki Ichikawa16, Hidetaka Shibata16, Akinobu Takaki17, Satoshi Yamagiwa18, Masataka Seike19, Shotaro Sakisaka20, Yasuaki Takeyama 20, Masaru Harada21, Michio Senju21, Osamu Yokosuka22, Tatsuo Kanda 22, Yoshiyuki Ueno 23, Hirotoshi Ebinuma24, Takashi Himoto25, Kazumoto Murata4, Shinji Shimoda26, Shinya Nagaoka6, Seigo Abiru6, Atsumasa Komori6,27, Kiyoshi Migita6,27, Masahiro Ito6,27, Hiroshi Yatsuhashi6,27, Yoshihiko Maehara28, Shinji Uemoto29, Norihiro Kokudo30, Masao Nagasaki2,3,31, Katsushi Tokunaga1 & Minoru Nakamura6,7,27,32 Primary biliary cholangitis (PBC) is a chronic and cholestatic autoimmune liver disease caused by the destruction of intrahepatic small bile ducts. Our previous genome-wide association study (GWAS) identifed six susceptibility loci for PBC.