Qt6506g5sk.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

UNIVERSITY of CALIFORNIA Los Angeles
UNIVERSITY OF CALIFORNIA Los Angeles Dissecting transcriptional control by Klf4 in somatic cell reprogramming A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Biological Chemistry by Huajun Zhou 2017 ABSTRACT OF THE DISSERTATION Dissecting transcriptional control by Klf4 in somatic cell reprogramming by Huajun Zhou Doctor of Philosophy in Biological Chemistry University of California, Los Angeles, 2017 Professor Gregory S. Payne, Chair Ectopic expression of four transcription factors, Oct4, Sox2, Klf4, and c-Myc, coverts somatic cells directly into induced pluripotent stem cells (iPSCs), which are functionally equivalent to embryonic stem cells (ESCs). The discovery of iPSC has been reshaping the methodology of disease modeling and drug screening in the past decade, and provides tremendous promise for regenerative medicine. However, the mechanism underlying this conversion process, reprogramming, is not yet fully understood. I aimed to dissect the reprogramming process, by characterizing the functional domains of one reprogramming factor Klf4. The transcriptional activation domain (TAD) of Klf4 was revealed to be critical for reprogramming. To search for the factors that mediates the functionality of Klf4 TAD, I identified transcriptional coactivators CBP/p300 and Mediator complex as the physical interaction partners of Klf4 TAD, and further showed that this interaction is functionally required ii for Klf4 mediated transcriptional activation in reprograming. Clathrin heavy chain, initially identified as a physical interaction partner of Klf4 TAD, was shown to be not required for Klf4 transcriptional activation. Clathrin heavy chain was furthered characterized for its potential transcriptional activation activity in CHC-TFE3, a chromosomal fusion discovered in renal cell carcinoma. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The GATA2 Transcription Factor Negatively Regulates the Proliferation of Neuronal Progenitors
RESEARCH ARTICLE 2155 Development 133, 2155-2165 (2006) doi:10.1242/dev.02377 The GATA2 transcription factor negatively regulates the proliferation of neuronal progenitors Abeer El Wakil*, Cédric Francius*,†, Annie Wolff, Jocelyne Pleau-Varet† and Jeannette Nardelli†,§ Postmitotic neurons are produced from a pool of cycling progenitors in an orderly fashion that requires proper spatial and temporal coordination of proliferation, fate determination, differentiation and morphogenesis. This probably relies on complex interplay between mechanisms that control cell cycle, specification and differentiation. In this respect, we have studied the possible implication of GATA2, a transcription factor that is involved in several neuronal specification pathways, in the control of the proliferation of neural progenitors in the embryonic spinal cord. Using gain- and loss-of-function manipulations, we have shown that Gata2 can drive neural progenitors out of the cycle and, to some extent, into differentiation. This correlates with the control of cyclin D1 transcription and of the expression of the p27/Kip1 protein. Interestingly, this functional aspect is not only associated with silencing of the Notch pathway but also appears to be independent of proneural function. Consistently, GATA2 also controls the proliferation capacity of mouse embryonic neuroepithelial cells in culture. Indeed, Gata2 inactivation enhances the proliferation rate in these cells. By contrast, GATA2 overexpression is sufficient to force such cells and neuroblastoma cells to stop dividing but not to drive either type of cell into differentiation. Furthermore, a non-cell autonomous effect of Gata2 expression was observed in vivo as well as in vitro. Hence, our data have provided evidence for the ability of Gata2 to inhibit the proliferation of neural progenitors, and they further suggest that, in this regard, Gata2 can operate independently of neuronal differentiation. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Supplementary Material
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Neurol Neurosurg Psychiatry Page 1 / 45 SUPPLEMENTARY MATERIAL Appendix A1: Neuropsychological protocol. Appendix A2: Description of the four cases at the transitional stage. Table A1: Clinical status and center proportion in each batch. Table A2: Complete output from EdgeR. Table A3: List of the putative target genes. Table A4: Complete output from DIANA-miRPath v.3. Table A5: Comparison of studies investigating miRNAs from brain samples. Figure A1: Stratified nested cross-validation. Figure A2: Expression heatmap of miRNA signature. Figure A3: Bootstrapped ROC AUC scores. Figure A4: ROC AUC scores with 100 different fold splits. Figure A5: Presymptomatic subjects probability scores. Figure A6: Heatmap of the level of enrichment in KEGG pathways. Kmetzsch V, et al. J Neurol Neurosurg Psychiatry 2021; 92:485–493. doi: 10.1136/jnnp-2020-324647 BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Neurol Neurosurg Psychiatry Appendix A1. Neuropsychological protocol The PREV-DEMALS cognitive evaluation included standardized neuropsychological tests to investigate all cognitive domains, and in particular frontal lobe functions. The scores were provided previously (Bertrand et al., 2018). Briefly, global cognitive efficiency was evaluated by means of Mini-Mental State Examination (MMSE) and Mattis Dementia Rating Scale (MDRS). Frontal executive functions were assessed with Frontal Assessment Battery (FAB), forward and backward digit spans, Trail Making Test part A and B (TMT-A and TMT-B), Wisconsin Card Sorting Test (WCST), and Symbol-Digit Modalities test. -

Identifying Isl1 Genetic Lineage in the Developing Olfactory System and in Gnrh-1 Neurons 2 3 Ed Zandro M
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.31.276360; this version posted August 31, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Identifying Isl1 genetic lineage in the developing olfactory system and in GnRH-1 neurons 2 3 Ed Zandro M. Taroc1, Raghu Ram Katreddi1 and Paolo E. Forni1*. 4 5 1Department of Biological Sciences; The RNA Institute, and the Center for Neuroscience 6 Research; University at Albany, State University of New York, Albany, NY 12222, USA. 7 8 * Corresponding Author 9 [email protected] 10 11 12 Keywords: Olfactory neurons, vomeronasal sensory neurons, GnRH neurons, Islet-1/Isl1, Isl1 13 Conditional knock-out; genetic lineage tracing, olfactory placode, neural crest. 14 15 Abstract (299, at 245 after editing) 16 During embryonic development, symmetric ectodermal thickenings (olfactory placodes) give rise 17 to several cell types that comprise the olfactory system, such as those that form the terminal nerve 18 ganglion (TN), gonadotropin releasing hormone-1 (GnRH-1) neurons and other migratory 19 neurons in rodents. Even though the genetic heterogeneity among these cell types are 20 documented, unidentified cell populations arising from the olfactory placode remain. One 21 candidate to identify placodal derived neurons in the developing nasal area is the transcription 22 factor Isl1, which was recently identified in GnRH-3 neurons of the terminal nerve in fish, as well 23 as expression in neurons of the nasal migratory mass. Here, we analyzed the Isl1 genetic lineage 24 in chemosensory neuronal populations in the nasal area and migratory GnRH-1 neurons in mice 25 using in-situ hybridization, immunolabeling a Tamoxifen inducible Isl1CreERT and a constitutive 26 Isl1Cre knock-in mouse lines. -

"Description"" ""Generatio"
TABLE S4 "ID ""Description"" ""GeneRatio"" ""BgRatio"" ""pvalue"" ""p.adjust"" ""qvalue"" ""geneID"" ""Count""" "GO:0003712 ""GO:0003712"" ""transcription coregulator activity"" ""84/1859"" ""454/22744"" 9.49597175224444e-13 9.80933882006851e-10 8.20651874588704e-10 ""Ncoa2/Zfp451/Dhx9/Hnrnpu/Cited2/Ncoa7/Ccar1/ Sirt1/Arid5b/Sirt6/Med1/Rara/Atxn7l3/Ddx5/Wbp2/Hdac9/Zmynd11/Cdyl/ Mier3/Sfmbt1/Gata4/Med4/Basp1/Zfpm2/Zhx2/Ddx17/Mkl2/Hes1/Nrip1/Usp16/ Elob/Rrp1b/Rxrb/Kat2b/Mta3/Hsbp1l1/Tle4/Sfr1/Eid1/Cops2/Sox12/Raly/ Ncoa6/Rbm39/Lpin3/Skil/Jade1/Maml3/Supt20/Med12l/Hdgf/Glmp/Nfib/Jun/ Pex14/Rere/Psmd9/Ncor2/Trim24/Ruvbl1/Rybp/Bhlhe40/Atf7ip/Ube3a/Mef2a/ Nrg1/Rbpms/Cnot7/Sin3b/Pou4f2/Pkn1/Cdyl2/Taf5l/Irf2bp2/Birc2/Yap1/ Skor1/Tfdp2/Rad54l2/Ctnnb1/Limd1/Med14/Rap2c/Tbl1x"" 84" "GO:0003779 ""GO:0003779"" ""actin binding"" ""65/1859"" ""414/22744"" 2.57546466939442e-07 8.86818334494813e-05 7.41914559148359e-05 ""Actr3/Cxcr4/Hnrnpu/Enah/Utrn/Epb41l2/Marcks/Ctnna3/Eef2/Pawr/ Ccdc88a/Anxa6/Gas7/Lasp1/Tns4/Syne2/Sipa1l1/Syne3/Phactr1/Enc1/Pxk/ Vcl/Ang/Myo10/Mtss1/Triobp/Mkl2/Afdn/Daam2/Svil/Ctnna1/Synpo/Myo5b/ Nrap/Ablim1/Shtn1/Fmnl2/Itprid2/Ino80/Pfn2/Myoz2/Pdlim5/Cap1/Macf1/ Epb41/Wasf2/Myom3/Ywhah/Coro1c/Ssh1/Hip1/Ppp1r9a/Wasl/Ctnna2/Mical3/ Eps8/Tlnrd1/Myom2/Klhl2/Sntb2/Spire2/Coro2b/Clasp2/Hdac6/Diaph2"" 65" "GO:0046332 ""GO:0046332"" ""SMAD binding"" ""22/1859"" ""84/22744"" 6.87941766027971e-07 0.000177660961076723 0.000148631628923412 ""Bmpr2/Cited2/Usp15/Ddx5/Axin2/Ppm1a/Yy1/Gata4/Tgif1/Ldlrad4/Smad7/ Acvr2a/Pmepa1/Skil/Trim33/Jun/Mef2a/Ipo7/Skor1/Rnf111/Tcf12/Ctnnb1"" -

REVIEWS Transcriptional Regulation Through Mediator-Like Coactivators
TIBS 25 – JUNE 2000 REVIEWS termed Mediator, that was necessary to support activated transcription in vitro in conjunction with general initiation Transcriptional regulation factors8, but apparently distinct from the TAF (Ref. 5) and USA (Ref. 7) coacti- through Mediator-like vators identified in metazoans (see below). Concurrent genetic analysis also implicated two broad groups of pro- coactivators in yeast and teins in transcriptional regulation in vivo. The SRB (suppressor of RNA metazoan cells polymerase B) proteins first emerged from genetic screens for suppressors of partial truncations in the C-terminal domain (CTD) of the largest subunit of Sohail Malik and Robert G. Roeder Pol II (Ref. 9). Other gene products, including GAL11, RGR1 and SIN4, emerged from genetic screens for regulatory A novel multiprotein complex has recently been identified as a coactivator for factors that function in other path- transcriptional control of protein-encoding genes by RNA polymerase II in ways10 (see below). Subsequent studies higher eukaryotic cells. This complex is evolutionarily related to the Mediator aimed at purifying the Mediator activity complex from yeast and, on the basis of its structural and functional charac- and the SRB proteins resulted in the teristics, promises to be a key target of diverse regulatory circuits. identification of a Pol II holoenzyme that contained the 12-subunit core Pol II, SRBs, other genetically identified TRANSCRIPTIONAL REGULATION IN factors that mediate, and might integrate, polypeptides (including GAL11, RGR1, eukaryotes is achieved through multi- the effects of transcriptional activators SIN4) and novel Mediator polypeptides protein complexes assembled at the on the Pol II basal machinery2. -

1 Nitrogen Cost Minimization Is Promoted by Structural Changes in the Transcriptome of N 1 Deprived Prochlorococcus Cells 2 3 Ro
bioRxiv preprint doi: https://doi.org/10.1101/087643; this version posted November 14, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Nitrogen cost minimization is promoted by structural changes in the transcriptome of N 2 deprived Prochlorococcus cells 3 4 Robert W. Read1, Paul M. Berube2, Steven J. Biller2, Iva Neveux1, Andres Cubillos- 5 Ruiz2,3, Sallie W. Chisholm2,4, and Joseph J. Grzymski1* 6 7 1. Division of Earth and Ecosystem Sciences, Desert Research Institute, Reno, NV, 8 USA 9 10 2. Department of Civil and Environmental Engineering, Massachusetts Institute of 11 Technology, Cambridge, MA, USA 12 13 3. Microbiology Graduate Program, Massachusetts Institute of Technology, 14 Cambridge, MA, USA 15 16 4. Department of Biology, Massachusetts Institute of Technology, Cambridge, MA, 17 USA 18 Correspondence: Joseph J. Grzymski, Department of Earth and Ecosystem Sciences, 19 Desert Research Institute, 2215 Raggio Parkway, Reno, NV 89509, USA. 20 *E-mail: [email protected] 21 22 Conflict of interest. 23 The authors declare no conflict of interest 1 bioRxiv preprint doi: https://doi.org/10.1101/087643; this version posted November 14, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 24 Abstract 25 Prochlorococcus is a globally abundant marine cyanobacterium with many adaptations 26 that reduce cellular nutrient requirements, facilitating growth in its nutrient-poor 27 environment. One such genomic adaptation is the preferential utilization of amino acids 28 containing fewer N-atoms, which minimizes cellular nitrogen requirements. -

GATA Factors and Hindbrain Development 5525
Development 126, 5523-5531 (1999) 5523 Printed in Great Britain © The Company of Biologists Limited 1999 DEV2473 The transcription factor GATA3 is a downstream effector of Hoxb1 specification in rhombomere 4 Illar Pata1,‡, Michèle Studer2,‡, J. Hikke van Doorninck3,*, James Briscoe4, Sulev Kuuse1, J. Douglas Engel5, Frank Grosveld3 and Alar Karis1,3,¶ 1Institute of Molecular and Cell Biology, University of Tartu, 23 Riia St, 51010 Tartu, Estonia 2Department of Developmental Neurobiology, King’s College London, 4th Floor, New Hunts House, Guy’s Hospital, London SE1 9RT, UK 3Department of Cell Biology and Genetics, Erasmus University Rotterdam, PO Box 1738, 3000 DR Rotterdam, The Netherlands 4Department of Biochemistry and Molecular Biophysics, Howard Hughes Medical Institute, Columbia University, New York, NY10032, USA 5Department of Biochemistry, Molecular Biology and Cell Biology, North Western University, Evanston, IL 60208 USA *Present address: Rudolf Magnus Institute for Neurosciences, Utrecht University, The Netherlands ‡These authors contributed equally to this work ¶Author for correspondence at address 1 (e-mail: [email protected]) Accepted 15 September; published on WWW 9 November 1999 SUMMARY In this paper, we show that the transcription factor GATA3 Hoxb1-deficient mice. Ubiquitous expression of Hoxb1 in is dynamically expressed during hindbrain development. the hindbrain induces ectopic expression of GATA2 and Function of GATA3 in ventral rhombomere (r) 4 is GATA3 in ventral r2 and r3. These findings demonstrate dependent on functional GATA2, which in turn is under the that GATA2 and GATA3 lie downstream of Hoxb1 and control of Hoxb1. In particular, the absence of Hoxb1 provide the first example of Hox pathway transcription results in the loss of GATA2 expression in r4 and the factors within a defined population of vertebrate motor absence of GATA2 results in the loss of GATA3 expression. -
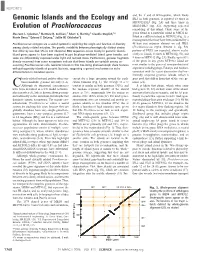
Genomic Islands and the Ecology and Evolution of Prochlorococcus
REPORTS ond, the 3¶ end of tRNA-proline, which flanks Genomic Islands and the Ecology and ISL3 in both genomes, is repeated 13 times in MIT9312-ISL3 (Fig. 2A) and three times in MED4-ISL3 (fig. S2), suggesting repeated Evolution of Prochlorococcus remodeling of this island. Third, some of the Maureen L. Coleman,1 Matthew B. Sullivan,1 Adam C. Martiny,1 Claudia Steglich,1* genes found in a particular island in MED4 are Kerrie Barry,2 Edward F. DeLong,1 Sallie W. Chisholm1† found in a different island in MIT9312 (Fig. 1), a rearrangement that may have been mediated by a Prochlorococcus ecotypes are a useful system for exploring the origin and function of diversity 48–base pair sequence element we call PRE1 among closely related microbes. The genetic variability between phenotypically distinct strains (Prochlorococcus repeat element 1; fig. S3); that differ by less that 1% in 16S ribosomal RNA sequences occurs mostly in genomic islands. portions of PRE1 are repeated, almost exclu- Island genes appear to have been acquired in part by phage-mediated lateral gene transfer, and sively in islands, 13 times in MED4 (fig. S2), and some are differentially expressed under light and nutrient stress. Furthermore, genome fragments 9 times in MIT9312 (Fig. 2A). Finally, up to 80% directly recovered from ocean ecosystems indicate that these islands are variable among co- of the genes in any given MIT9312 island are occurring Prochlorococcus cells. Genomic islands in this free-living photoautotroph share features most similar to the genes of noncyanobacterial with pathogenicity islands of parasitic bacteria, suggesting a general mechanism for niche organisms including phage, Eukarya, and Archaea, differentiation in microbial species. -

Recombinant Human MED4 Protein Catalog Number: ATGP0552
Recombinant human MED4 protein Catalog Number: ATGP0552 PRODUCT INPORMATION Expression system E.coli Domain 1-270aa UniProt No. Q9NPJ6 NCBI Accession No. AAH05189 Alternative Names Mediator complex subunit 4, ARC36, DRIP36, VDRIP, Mediator complex subunit 4 Activator-recruited cofactor 36 kDa component, HSPC126, Mediator of RNA polymerase II transcription subunit 4, TRAP/SMCC/PC2 subunit p36 subunit, Vitamin D3 receptor-interacting protein complex 36 kDa component. PRODUCT SPECIFICATION Molecular Weight 30.7 kDa (278aa) confirmed by MALDI-TOF Concentration 0.5mg/ml (determined by Bradford assay) Formulation Liquid in. 20mM Tris-HCl buffer (pH 8.0) containing 20% glycerol, 1mM DTT, 100mM NaCl Purity > 80% by SDS-PAGE Tag His-Tag Application SDS-PAGE Storage Condition Can be stored at +2C to +8C for 1 week. For long term storage, aliquot and store at -20C to -80C. Avoid repeated freezing and thawing cycles. BACKGROUND Description Mediator complex subunit 4 (MED4) is also known as mediator of RNA polymerase II transcription subunit 4 or vitamin D3 receptor-interacting protein complex 36 kDa component (DRIP36). This protein is a component of the vitamin D receptor-interacting protein (DRIP) complex which functions as a nuclear receptor coactivator. The DRIP complex is capable of activating nuclear receptors in a ligand-dependent manner. Recombinant human 1 Recombinant human MED4 protein Catalog Number: ATGP0552 MED4, fused to His-tag at C-terminus, was expressed in E. coli and purified by using conventional chromatography techniques. Amino acid Sequence MAASSSGEKE KERLGGGLGV AGGNSTRERL LSALEDLEVL SRELIEMLAI SRNQKLLQAG EENQVLELLI HRDGEFQELM KLALNQGKIH HEMQVLEKEV EKRDGDIQQL QKQLKEAEQI LATAVYQAKE KLKSIEKARK GAISSEEIIK YAHRISASNA VCAPLTWVPG DPRRPYPTDL EMRSGLLGQM NNPSTNGVNG HLPGDALAAG RLPDVLAPQY PWQSNDMSMN MLPPNHSSDF LLEPPGHNKE DEDDVEIMST DSSSSSSESD <LEHHHHHH> General References Rachez C., et al.