Defensin Beta 5 Human Protein – AR31146PU-N | Origene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Manual Annotation and Analysis of the Defensin Gene Cluster in the C57BL
BMC Genomics BioMed Central Research article Open Access Manual annotation and analysis of the defensin gene cluster in the C57BL/6J mouse reference genome Clara Amid*†1, Linda M Rehaume*†2, Kelly L Brown2,3, James GR Gilbert1, Gordon Dougan1, Robert EW Hancock2 and Jennifer L Harrow1 Address: 1Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, Cambridgeshire CB10 1SA, UK, 2University of British Columbia, Centre for Microbial Disease & Immunity Research, 2259 Lower Mall, Vancouver, BC, V6T 1Z4, Canada and 3Department of Rheumatology and Inflammation Research, Göteborg University, Guldhedsgatan 10, S-413 46 Göteborg, Sweden Email: Clara Amid* - [email protected]; Linda M Rehaume* - [email protected]; Kelly L Brown - [email protected]; James GR Gilbert - [email protected]; Gordon Dougan - [email protected]; Robert EW Hancock - [email protected]; Jennifer L Harrow - [email protected] * Corresponding authors †Equal contributors Published: 15 December 2009 Received: 15 May 2009 Accepted: 15 December 2009 BMC Genomics 2009, 10:606 doi:10.1186/1471-2164-10-606 This article is available from: http://www.biomedcentral.com/1471-2164/10/606 © 2009 Amid et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Host defense peptides are a critical component of the innate immune system. Human alpha- and beta-defensin genes are subject to copy number variation (CNV) and historically the organization of mouse alpha-defensin genes has been poorly defined. -

Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected]
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School July 2017 Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected] Follow this and additional works at: http://scholarcommons.usf.edu/etd Part of the Pathology Commons Scholar Commons Citation Mohamed, Mai, "Role of Amylase in Ovarian Cancer" (2017). Graduate Theses and Dissertations. http://scholarcommons.usf.edu/etd/6907 This Dissertation is brought to you for free and open access by the Graduate School at Scholar Commons. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Scholar Commons. For more information, please contact [email protected]. Role of Amylase in Ovarian Cancer by Mai Mohamed A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Pathology and Cell Biology Morsani College of Medicine University of South Florida Major Professor: Patricia Kruk, Ph.D. Paula C. Bickford, Ph.D. Meera Nanjundan, Ph.D. Marzenna Wiranowska, Ph.D. Lauri Wright, Ph.D. Date of Approval: June 29, 2017 Keywords: ovarian cancer, amylase, computational analyses, glycocalyx, cellular invasion Copyright © 2017, Mai Mohamed Dedication This dissertation is dedicated to my parents, Ahmed and Fatma, who have always stressed the importance of education, and, throughout my education, have been my strongest source of encouragement and support. They always believed in me and I am eternally grateful to them. I would also like to thank my brothers, Mohamed and Hussien, and my sister, Mariam. I would also like to thank my husband, Ahmed. -

Looking for Missing Proteins in the Proteome Of
Looking for Missing Proteins in the Proteome of Human Spermatozoa: An Update Yves Vandenbrouck, Lydie Lane, Christine Carapito, Paula Duek, Karine Rondel, Christophe Bruley, Charlotte Macron, Anne Gonzalez de Peredo, Yohann Coute, Karima Chaoui, et al. To cite this version: Yves Vandenbrouck, Lydie Lane, Christine Carapito, Paula Duek, Karine Rondel, et al.. Looking for Missing Proteins in the Proteome of Human Spermatozoa: An Update. Journal of Proteome Research, American Chemical Society, 2016, 15 (11), pp.3998-4019. 10.1021/acs.jproteome.6b00400. hal-02191502 HAL Id: hal-02191502 https://hal.archives-ouvertes.fr/hal-02191502 Submitted on 19 Mar 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Journal of Proteome Research 1 2 3 Looking for missing proteins in the proteome of human spermatozoa: an 4 update 5 6 Yves Vandenbrouck1,2,3,#,§, Lydie Lane4,5,#, Christine Carapito6, Paula Duek5, Karine Rondel7, 7 Christophe Bruley1,2,3, Charlotte Macron6, Anne Gonzalez de Peredo8, Yohann Couté1,2,3, 8 Karima Chaoui8, Emmanuelle Com7, Alain Gateau5, AnneMarie Hesse1,2,3, Marlene 9 Marcellin8, Loren Méar7, Emmanuelle MoutonBarbosa8, Thibault Robin9, Odile Burlet- 10 Schiltz8, Sarah Cianferani6, Myriam Ferro1,2,3, Thomas Fréour10,11, Cecilia Lindskog12,Jérôme 11 1,2,3 7,§ 12 Garin , Charles Pineau . -

Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications
Stem Cell Rev and Rep DOI 10.1007/s12015-016-9662-8 Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications Behnam Ahmadian Baghbaderani 1 & Adhikarla Syama2 & Renuka Sivapatham3 & Ying Pei4 & Odity Mukherjee2 & Thomas Fellner1 & Xianmin Zeng3,4 & Mahendra S. Rao5,6 # The Author(s) 2016. This article is published with open access at Springerlink.com Abstract We have recently described manufacturing of hu- help determine which set of tests will be most useful in mon- man induced pluripotent stem cells (iPSC) master cell banks itoring the cells and establishing criteria for discarding a line. (MCB) generated by a clinically compliant process using cord blood as a starting material (Baghbaderani et al. in Stem Cell Keywords Induced pluripotent stem cells . Embryonic stem Reports, 5(4), 647–659, 2015). In this manuscript, we de- cells . Manufacturing . cGMP . Consent . Markers scribe the detailed characterization of the two iPSC clones generated using this process, including whole genome se- quencing (WGS), microarray, and comparative genomic hy- Introduction bridization (aCGH) single nucleotide polymorphism (SNP) analysis. We compare their profiles with a proposed calibra- Induced pluripotent stem cells (iPSCs) are akin to embryonic tion material and with a reporter subclone and lines made by a stem cells (ESC) [2] in their developmental potential, but dif- similar process from different donors. We believe that iPSCs fer from ESC in the starting cell used and the requirement of a are likely to be used to make multiple clinical products. We set of proteins to induce pluripotency [3]. Although function- further believe that the lines used as input material will be used ally identical, iPSCs may differ from ESC in subtle ways, at different sites and, given their immortal status, will be used including in their epigenetic profile, exposure to the environ- for many years or even decades. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

High-Resolution Analysis of Chromosomal Alterations in Adult Acute Lymphoblastic Leukemia
Elmer Press Original Article J Hematol. 2014;3(3):65-71 High-Resolution Analysis of Chromosomal Alterations in Adult Acute Lymphoblastic Leukemia Lam Kah Yuena, c, Zakaria Zubaidaha, Ivyna Bong Pau Nia, Megat Baharuddin Puteri Jamilatul Noora, Esa Ezaliaa, Chin Yuet Menga, Ong Tee Chuanb, Vegappan Subramanianb, Chang Kiang Mengb Abstract Introduction Background: Chromosomal alterations occur frequently in acute Acute lymphoblastic leukemia (ALL) is a heterogeneous lymphoblastic leukemia (ALL), affecting either the chromosome disease, resulting from the accumulation of chromosomal al- number or structural changes. These alterations can lead to inacti- terations either in the form of numerical or structural chang- vation of tumor suppressor genes and/or activation of oncogenes. es such as amplification, deletion, inversion or translocation. The objective of this study was to identify recurrent and/or novel The frequency of chromosomal alterations in adult ALL is chromosomal alterations in adult ALL using single nucleotide poly- 64-85% [1], compared to 60-69% in childhood ALL. Trans- morphism (SNP) array analysis. location t(9.22), one of the most common recurring chromo- Methods: We studied 41 cases of adult ALL compared with healthy somal alterations, is found in 20-40% adult ALL patients [2], normal controls using SNP array. and its incidence increases with age. Some of the chromo- somal alterations are significantly associated with remission Results: Our analysis revealed 43 copy number variant regions, duration, complete remission rate and disease-free survival of which 44% were amplifications and 56% were deletions. The [1]. Increasing age is also associated with lower remission most common amplifications were on chromosome regions 8p23.1 rates, shorter remissions and poor outcomes in adult ALL (71%), 1q44 (66%), 1q23.3 (54%), 11q23.3 (54%), 12p13.33 [1]. -

Genomic Alterations on 8P21-P23 Are the Most Frequent Genetic Events in Stage I Squamous Cell Carcinoma of the Lung
EXPERIMENTAL AND THERAPEUTIC MEDICINE 9: 345-350, 2015 Genomic alterations on 8p21-p23 are the most frequent genetic events in stage I squamous cell carcinoma of the lung JIUN KANG Department of Biomedical Laboratory Science, Korea Nazarene University, Cheonan 330-718, Republic of Korea Received April 23, 2014; Accepted October 31, 2014 DOI: 10.3892/etm.2014.2123 Abstract. Genetic alterations in the early stages of cancer genetic changes, and chromosomal aberrations are a hallmark have a close correlation with tumor initiation and poten- of cancer cells, occurring at a high prevalence in SCC. Although tially activate downstream pathways implicated in tumor a number of studies have been performed to evaluate genetic progression; however, the method of initiation in sporadic events associated with the development and progression of neoplasias is largely unknown. In this study, whole-genome SCC (2,3), the molecular mechanism remains to be uncovered, microarray-comparative genomic hybridization was and the identification of predictive markers is crucial. performed to identify the early genetic alterations that define Genetic alterations in the early stages of cancer have a the prognosis of patients with stage I squamous cell carcinoma close correlation with tumor initiation, and potentially activate (SCC) of the lung. The most striking finding was the high downstream pathways that are implicated in tumor progres- frequency of copy number losses and hemizygous deletions on sion. With the recent advances of computed tomographic chromosome 8p, which occurred in 94.7% (18/19) and 63.2% technology, the number of patients diagnosed with stage I (12/19) of the cases, respectively, with a delineated minimal lung SCC has been increasing (3). -

Caractérisation De Nouveaux Gènes Et Polymorphismes Potentiellement Impliqués Dans Les Interactions Hôtes-Pathogènes
Aix-Marseille Université, Faculté de Médecine de Marseille Ecole Doctorale des Sciences de la Vie et de la Santé THÈSE DE DOCTORAT Présentée par Charbel ABOU-KHATER Date et lieu de naissance: 08-Juilllet-1990, Zahlé, LIBAN En vue de l’obtention du grade de Docteur de l’Université d’Aix-Marseille Mention: Biologie, Spécialité: Microbiologie Caractérisation de nouveaux gènes et polymorphismes potentiellement impliqués dans les interactions hôtes-pathogènes Publiquement soutenue le 5 Juillet 2017 devant le jury composé de : Pr. Daniel OLIVE Directeur de Thèse Pr. Brigitte CROUAU-ROY Rapporteur Dr. Benoît FAVIER Rapporteur Dr. Pierre PONTAROTTI Examinateur Thèse codirigée par Pr. Daniel OLIVE et Dr Laurent ABI-RACHED Laboratoires d’accueil URMITE Research Unit on Emerging Infectious and Tropical Diseases, UMR 6236, Faculty of Medicine, 27, Boulevard Jean Moulin, 13385 Marseille, France CRCM, Centre de Recherche en Cancérologie de Marseille,Inserm 1068, 27 Boulevard Leï Roure, BP 30059, 13273 Marseille Cedex 09, France 2 Acknowledgements First and foremost, praises and thanks to God, Holy Mighty, Holy Immortal, All-Holy Trinity, for His showers of blessings throughout my whole life and to whom I owe my very existence. Glory to the Father, and to the Son, and to the Holy Spirit: now and ever and unto ages of ages. I would like to express my sincere gratitude to my advisors Prof. Daniel Olive and Dr. Laurent Abi-Rached, for the continuous support, for their patience, motivation, and immense knowledge. Someday, I hope to be just like you. A special thanks to my “Godfather” who perfectly fulfilled his role, Dr. -

Francine Blumental De Abreu
VARIAÇÃO NO NÚMERO DE CÓPIAS GENÔMICAS NA AVALIAÇÃO DE GENES PRINCIPAIS DE PREDISPOSIÇÃO EM PACIENTES COM SÍNDROME DE MAMA-CÓLON TRIADOS PARA MUTAÇÕES NOS GENES BRCA1, BRCA2, TP53, CHEK2, MLH1 E MSH2 FRANCINE BLUMENTAL DE ABREU Tese apresentada à Fundação Antônio Prudente para obtenção do Título de Doutor em Ciências Área de concentração: Oncologia Orientadora: Profª Dra. Silvia Regina Rogatto São Paulo 2012 FICHA CATALOGRÁFICA Preparada pela Biblioteca da Fundação Antônio Prudente Abreu, Francine Blumental de Variação no número de cópias genômicas na avaliação de genes principais de predisposição em pacientes com síndrome de mama- cólon triados para mutações nos genes BRCA1, BRCA2, TP53, CHEK2, MLH1 E MSH2 / Francine Blumental de Abreu – São Paulo, 2012. 245p. Tese (Doutorado)-Fundação Antônio Prudente. Curso de Pós-Graduação em Ciências - Área de concentração: Oncologia. Orientadora: Silvia Regina Rogatto Descritores: 1. SÍNDROME DE LYNCH. 2. NEOPLASIAS DA MAMA. 3. VARIAÇÃO DO NÚMERO DE CÓPIA DO DNA. 4. HIBRIDIZAÇÃO GENÔMICA COMPARATIVA. DEDICATÓRIA A minha família!!!! AGRADECIMENTOS À minha família... Edilucia (II:5) e Antonio (II:6) por serem mais do que pais, por serem mais do que amigos, por serem os meus super heróis! Obrigada pelos abraços, pelas palavras de apoio e de ensinamento, pelo carinho, pelo amor incondicional! Obrigada por estarem sempre disponíveis, por estenderem as mãos depois de uma “rasteira” da vida, por enxugarem as minhas lágrimas! Obrigada principalmente por me fazerem acreditar que momentos difíceis existem, -
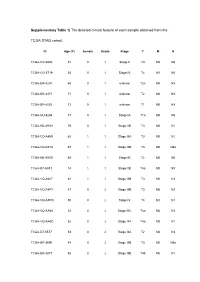
Supplementary Table 1| the Detailed Clinical Feature of Each Sample Obtained from The
Supplementary Table 1| The detailed clinical feature of each sample obtained from the TCGA STAD cohort. ID Age (Y) Gender Grade Stage T M N TCGA-CD-5800 51 0 1 Stage II T3 M0 N0 TCGA-CG-5719 54 0 1 Stage IV T4 M1 N0 TCGA-BR-4201 66 0 1 unknow T2a M0 NX TCGA-BR-4371 71 0 1 unknow T2 M0 NX TCGA-BR-4292 73 0 1 unknow T1 M0 NX TCGA-HU-8244 77 0 1 Stage IA T1a M0 N0 TCGA-KB-A93H 79 0 1 Stage IIB T3 M0 N1 TCGA-CD-A4MI 62 1 1 Stage IIIA T3 M0 N1 TCGA-VQ-A91A 67 1 1 Stage IIIB T3 M0 N3a TCGA-KB-A93G 68 1 1 Stage IB T2 M0 N0 TCGA-B7-A5TJ 74 1 1 Stage IIB T4a M0 NX TCGA-VQ-A927 81 1 1 Stage IIIB T3 M0 N3 TCGA-VQ-A8PY 47 0 2 Stage IIIB T3 M0 N2 TCGA-VQ-A8PQ 50 0 2 Stage IV T4 M1 N1 TCGA-VQ-AA68 52 0 2 Stage IIIC T4a M0 N3 TCGA-VQ-AA6D 52 0 2 Stage IIIA T4a M0 N1 TCGA-D7-5577 53 0 2 Stage IIIA T2 M0 N3 TCGA-BR-8690 54 0 2 Stage IIIB T3 M0 N3a TCGA-BR-8077 58 0 2 Stage IIIB T4b M0 N1 TCGA-HU-A4H2 58 0 2 Stage IIIB T3 M0 N3b TCGA-VQ-A91N 59 0 2 Stage IV T4 M0 N3 TCGA-D7-A74A 61 0 2 Stage IIIA T3 M0 N2 TCGA-HU-A4GP 62 0 2 Stage IIA T2 M0 N1 TCGA-BR-7716 62 0 2 Stage IIB T3 M0 N1 TCGA-BR-8679 63 0 2 Stage IB T2 M0 N0 TCGA-BR-8373 65 0 2 Stage IIIA T4a M0 N1 TCGA-VQ-A8PB 65 0 2 Stage II T3 M0 N0 TCGA-R5-A7ZF 65 0 2 Stage IV T4a M1 N1 TCGA-CG-4460 66 0 2 Stage IV T4 M1 N1 TCGA-D7-8579 66 0 2 Stage IIB T2 M0 N2 TCGA-R5-A7ZE 66 0 2 Stage IIIA T3 M0 N2 TCGA-VQ-A91Z 67 0 2 Stage IIIA T3 M0 N1 TCGA-HU-A4G9 67 0 2 Stage IA T1a M0 N0 TCGA-D7-6526 67 0 2 Stage IIIA T3 M0 N2 TCGA-BR-A4PE 68 0 2 Stage IB T2 M0 N0 TCGA-VQ-A8DL 69 0 2 Stage IIA T3 M0 N0 TCGA-BR-A4J6 -

Genome‑Wide Copy Number Analysis of Circulating Tumor Cells in Breast Cancer Patients with Liver Metastasis
ONCOLOGY REPORTS 44: 1075-1093, 2020 Genome‑wide copy number analysis of circulating tumor cells in breast cancer patients with liver metastasis LINGLIN ZOU1*, SABER IMANI1*, MAZAHER MAGHSOUDLOO2,3, MARZIEH DEHGHAN SHASALTANEH4, LANYANG GAO5, JIA ZHOU6, QINGLIAN WEN1, SHUYA LIU1, LEISHENG ZHANG7 and GANG CHEN8 1Department of Oncology, The Affiliated Hospital of Southwest Medical University, Southwest Medical University, Luzhou, Sichuan 646000, P.R. China; 2Laboratory of Systems Biology and Bioinformatics (LBB), Institute of Biochemistry and Biophysics, University of Tehran, Tehran 1417614411; 3Department of Genetics, Faculty of Advanced Science and Technology, Tehran Medical Sciences, Islamic Azad University, Tehran 1916893813; 4Department of Biology, Faculty of Science, University of Zanjan, Zanjan 4537138791, Iran; 5Sichuan Provincial Center for Gynaecology and Breast Disease, The Affiliated Hospital of Southwest Medical University;6 School of Humanities and Management Science, Southwest Medical University, Luzhou, Sichuan 646000; 7The Postdoctoral Research Station, School of Medicine, Nankai University, Tianjin 300071; 8Department of Medical Equipment, The Affiliated Hospital of Southwest Medical University, Southwest Medical University, Luzhou, Sichuan 646000, P.R. China Received December 6, 2019; Accepted May 12, 2020 DOI: 10.3892/or.2020.7650 Abstract. The genome-wide copy number analysis of circu- lating tumor cells (CTCs) provides a promising prognostic biomarker for survival in breast cancer liver metastasis Correspondence to: -

Chromosomal Breakpoints in Primary Colon Cancer Cluster at Sites of Structural Variants in the Genome
Research Article Chromosomal Breakpoints in Primary Colon Cancer Cluster at Sites of Structural Variants in the Genome Jordi Camps,1 Marian Grade,1,3 Quang Tri Nguyen,1 Patrick Ho¨rmann,1 Sandra Becker,1 Amanda B. Hummon,1 Virginia Rodriguez,2 Settara Chandrasekharappa,2 Yidong Chen,1 Michael J. Difilippantonio,1 Heinz Becker,3 B. Michael Ghadimi,3 and Thomas Ried1 1Genetics Branch, Center for Cancer Research, National Cancer Institute/NIH; 2Genome Technology Branch, National Human Genome Research Institute/NIH, Bethesda, Maryland; and 3Department of General and Visceral Surgery, University Medicine Go¨ttingen, Go¨ttingen, Germany Abstract 8q, 13, and 20q as well as losses of chromosomes 4q, 8p, 17p, and 18q (2). Genomic aberrations on chromosome 8 are common in colon cancer, and are associated with lymph node and distant Within the last decade, microarray technology has been metastases as well as with disease susceptibility. This extensively applied to survey the cellular transcriptome of common prompted us to generate a high-resolution map of genomic solid tumors, including colorectal cancer, and for colon cancers, imbalances of chromosome 8 in 51 primary colon carcinomas gene expression signatures were subsequently correlated with using a custom-designed genomic array consisting of a tiling clinical outcome (for reviews, see refs. 3–5). However, high- path of BAC clones. This analysis confirmed the dominant role resolution mapping of chromosomal copy number changes has of this chromosome. Unexpectedly, the position of the break- only recently been achieved using BAC or cDNA clone-based arrays points suggested colocalization with structural variants in the (6–10).