Three Domains of Life
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Basal Body Structure and Composition in the Apicomplexans Toxoplasma and Plasmodium Maria E
Francia et al. Cilia (2016) 5:3 DOI 10.1186/s13630-016-0025-5 Cilia REVIEW Open Access Basal body structure and composition in the apicomplexans Toxoplasma and Plasmodium Maria E. Francia1* , Jean‑Francois Dubremetz2 and Naomi S. Morrissette3 Abstract The phylum Apicomplexa encompasses numerous important human and animal disease-causing parasites, includ‑ ing the Plasmodium species, and Toxoplasma gondii, causative agents of malaria and toxoplasmosis, respectively. Apicomplexans proliferate by asexual replication and can also undergo sexual recombination. Most life cycle stages of the parasite lack flagella; these structures only appear on male gametes. Although male gametes (microgametes) assemble a typical 9 2 axoneme, the structure of the templating basal body is poorly defined. Moreover, the rela‑ tionship between asexual+ stage centrioles and microgamete basal bodies remains unclear. While asexual stages of Plasmodium lack defined centriole structures, the asexual stages of Toxoplasma and closely related coccidian api‑ complexans contain centrioles that consist of nine singlet microtubules and a central tubule. There are relatively few ultra-structural images of Toxoplasma microgametes, which only develop in cat intestinal epithelium. Only a subset of these include sections through the basal body: to date, none have unambiguously captured organization of the basal body structure. Moreover, it is unclear whether this basal body is derived from pre-existing asexual stage centrioles or is synthesized de novo. Basal bodies in Plasmodium microgametes are thought to be synthesized de novo, and their assembly remains ill-defined. Apicomplexan genomes harbor genes encoding δ- and ε-tubulin homologs, potentially enabling these parasites to assemble a typical triplet basal body structure. -

Anoxygenic Photosynthesis in Photolithotrophic Sulfur Bacteria and Their Role in Detoxication of Hydrogen Sulfide
antioxidants Review Anoxygenic Photosynthesis in Photolithotrophic Sulfur Bacteria and Their Role in Detoxication of Hydrogen Sulfide Ivan Kushkevych 1,* , Veronika Bosáková 1,2 , Monika Vítˇezová 1 and Simon K.-M. R. Rittmann 3,* 1 Department of Experimental Biology, Faculty of Science, Masaryk University, 62500 Brno, Czech Republic; [email protected] (V.B.); [email protected] (M.V.) 2 Department of Biology, Faculty of Medicine, Masaryk University, 62500 Brno, Czech Republic 3 Archaea Physiology & Biotechnology Group, Department of Functional and Evolutionary Ecology, Universität Wien, 1090 Vienna, Austria * Correspondence: [email protected] (I.K.); [email protected] (S.K.-M.R.R.); Tel.: +420-549-495-315 (I.K.); +431-427-776-513 (S.K.-M.R.R.) Abstract: Hydrogen sulfide is a toxic compound that can affect various groups of water microorgan- isms. Photolithotrophic sulfur bacteria including Chromatiaceae and Chlorobiaceae are able to convert inorganic substrate (hydrogen sulfide and carbon dioxide) into organic matter deriving energy from photosynthesis. This process takes place in the absence of molecular oxygen and is referred to as anoxygenic photosynthesis, in which exogenous electron donors are needed. These donors may be reduced sulfur compounds such as hydrogen sulfide. This paper deals with the description of this metabolic process, representatives of the above-mentioned families, and discusses the possibility using anoxygenic phototrophic microorganisms for the detoxification of toxic hydrogen sulfide. Moreover, their general characteristics, morphology, metabolism, and taxonomy are described as Citation: Kushkevych, I.; Bosáková, well as the conditions for isolation and cultivation of these microorganisms will be presented. V.; Vítˇezová,M.; Rittmann, S.K.-M.R. -

Revised Glossary for AQA GCSE Biology Student Book
Biology Glossary amino acids small molecules from which proteins are A built abiotic factor physical or non-living conditions amylase a digestive enzyme (carbohydrase) that that affect the distribution of a population in an breaks down starch ecosystem, such as light, temperature, soil pH anaerobic respiration respiration without using absorption the process by which soluble products oxygen of digestion move into the blood from the small intestine antibacterial chemicals chemicals produced by plants as a defence mechanism; the amount abstinence method of contraception whereby the produced will increase if the plant is under attack couple refrains from intercourse, particularly when an egg might be in the oviduct antibiotic e.g. penicillin; medicines that work inside the body to kill bacterial pathogens accommodation ability of the eyes to change focus antibody protein normally present in the body acid rain rain water which is made more acidic by or produced in response to an antigen, which it pollutant gases neutralises, thus producing an immune response active site the place on an enzyme where the antimicrobial resistance (AMR) an increasing substrate molecule binds problem in the twenty-first century whereby active transport in active transport, cells use energy bacteria have evolved to develop resistance against to transport substances through cell membranes antibiotics due to their overuse against a concentration gradient antiretroviral drugs drugs used to treat HIV adaptation features that organisms have to help infections; they -

Limits of Life on Earth Some Archaea and Bacteria
Limits of life on Earth Thermophiles Temperatures up to ~55C are common, but T > 55C is Some archaea and bacteria (extremophiles) can live in associated usually with geothermal features (hot springs, environments that we would consider inhospitable to volcanic activity etc) life (heat, cold, acidity, high pressure etc) Thermophiles are organisms that can successfully live Distinguish between growth and survival: many organisms can survive intervals of harsh conditions but could not at high temperatures live permanently in such conditions (e.g. seeds, spores) Best studied extremophiles: may be relevant to the Interest: origin of life. Very hot environments tolerable for life do not seem to exist elsewhere in the Solar System • analogs for extraterrestrial environments • `extreme’ conditions may have been more common on the early Earth - origin of life? • some unusual environments (e.g. subterranean) are very widespread Extraterrestrial Life: Spring 2008 Extraterrestrial Life: Spring 2008 Grand Prismatic Spring, Yellowstone National Park Hydrothermal vents: high pressure in the deep ocean allows liquid water Colors on the edge of the at T >> 100C spring are caused by different colonies of thermophilic Vents emit superheated water (300C or cyanobacteria and algae more) that is rich in minerals Hottest water is lifeless, but `cooler’ ~50 species of such thermophiles - mostly archae with some margins support array of thermophiles: cyanobacteria and anaerobic photosynthetic bacteria oxidize sulphur, manganese, grow on methane + carbon monoxide etc… Sulfolobus: optimum T ~ 80C, minimum 60C, maximum 90C, also prefer a moderately acidic pH. Live by oxidizing sulfur Known examples can grow (i.e. multiply) at temperatures which is abundant near hot springs. -

Supplementary Table S2: New Taxonomic Assignment of Sequences of Basal Fungal Lineages
Supplementary Table S2: New taxonomic assignment of sequences of basal fungal lineages. Fungal sequences were subjected to BLAST-N analysis and checked for their taxonomic placement in the eukaryotic guide-tree of the SILVA release 111. Sequences were classified depending on combined results from the methods mentioned above as well as literature searches. Accession Name New classification Clustering of the sequence in the Best BLAST-N hit number based on combined results eukaryotic guide tree of SILVA Name Accession number E.value Identity AB191431 Uncultured fungus Chytridiomycota Chytridiomycota Basidiobolus haptosporus AF113413.1 0.0 91 AB191432 Unculltured eukaryote Blastocladiomycota Blastocladiomycota Rhizophlyctis rosea NG_017175.1 0.0 91 AB252775 Uncultured eukaryote Chytridiomycota Chytridiomycota Blastocladiales sp. EF565163.1 0.0 91 AB252776 Uncultured eukaryote Fungi Nucletmycea_Fonticula Rhizophydium sp. AF164270.2 0.0 87 AB252777 Uncultured eukaryote Chytridiomycota Chytridiomycota Basidiobolus haptosporus AF113413.1 0.0 91 AB275063 Uncultured fungus Chytridiomycota Chytridiomycota Catenomyces sp. AY635830.1 0.0 90 AB275064 Uncultured fungus Chytridiomycota Chytridiomycota Endogone lactiflua DQ536471.1 0.0 91 AB433328 Nuclearia thermophila Nuclearia Nucletmycea_Nuclearia Nuclearia thermophila AB433328.1 0.0 100 AB468592 Uncultured fungus Basal clone group I Chytridiomycota Physoderma dulichii DQ536472.1 0.0 90 AB468593 Uncultured fungus Basal clone group I Chytridiomycota Physoderma dulichii DQ536472.1 0.0 91 AB468594 Uncultured -

James A. Mccloskey, Jr
CHEMICAL HERITAGE FOUNDATION JAMES A. MCCLOSKEY, JR. Transcript of Interviews Conducted by Michael A. Grayson at the McCloskeys’ Home Helotes, Texas on 19 and 20 March 2012 (With Subsequent Corrections and Additions) James A. McCloskey, Jr. ACKNOWLEDGMENT This oral history is one in a series initiated by the Chemical Heritage Foundation on behalf of the American Society for Mass Spectrometry. The series documents the personal perspectives of individuals related to the advancement of mass spectrometric instrumentation, and records the human dimensions of the growth of mass spectrometry in academic, industrial, and governmental laboratories during the twentieth century. This project is made possible through the generous support of the American Society for Mass Spectrometry. This oral history is designated Free Access. Please note: Users citing this interview for purposes of publication are obliged under the terms of the Chemical Heritage Foundation (CHF) Center for Oral History to credit CHF using the format below: James A. McCloskey, Jr., interview by Michael A. Grayson at the McCloskeys’ home, Helotes, Texas, 19-20 March 2012 (Philadelphia: Chemical Heritage Foundation, Oral History Transcript # 0702). Chemical Heritage Foundation Center for Oral History 315 Chestnut Street Philadelphia, Pennsylvania 19106 The Chemical Heritage Foundation (CHF) serves the community of the chemical and molecular sciences, and the wider public, by treasuring the past, educating the present, and inspiring the future. CHF maintains a world-class collection of materials that document the history and heritage of the chemical and molecular sciences, technologies, and industries; encourages research in CHF collections; and carries out a program of outreach and interpretation in order to advance an understanding of the role of the chemical and molecular sciences, technologies, and industries in shaping society. -
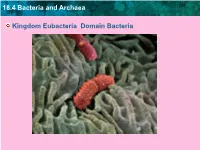
18.4 Bacteria and Archaea Kingdom Eubacteria Domain Bacteria
18.4 Bacteria and Archaea Kingdom Eubacteria Domain Bacteria 18.4 Bacteria and Archaea Description Bacteria are single-celled prokaryotes. 18.4 Bacteria and Archaea Where do they live? Prokaryotes are widespread on Earth. ( Est. over 1 billion types of bacteria, and over 1030 individual prokaryote cells on earth.) Found in all land and ocean environments, even inside other organisms! 18.4 Bacteria and Archaea Common Examples • E. Coli • Tetanus bacteria • Salmonella bacteria • Tuberculosis bacteria • Staphylococcus • Streptococcus 18.4 Bacteria and Archaea Modes Of Nutrition • Bacteria may be heterotrophs or autotrophs 18.4 Bacteria and Archaea Bacteria Reproduce How? • by binary fission. • exchange genes during conjugation= conjugation bridge increases diversity. • May survive by forming endospores = specialized cell with thick protective cell wall. TEM; magnification 6000x • Can survive for centuries until environment improves. Have been found in mummies! 18.4 Bacteria and Archaea • Bacteria Diagram – plasmid = small piece of genetic material, can replicate independently of the chromosome – flagellum = different than in eukaryotes, but for movement – pili = used to stick the bacteria to eachpili other or surfaces plasma membrance flagellum chromosome cell wall plasmid This diagram shows the typical structure of a prokaryote. Archaea and bacteria look very similar, although they have important molecular differences. 18.4 Bacteria and Archaea • Classified by: their need for oxygen, how they gram stain, and their shapes 18.4 Bacteria and Archaea Main Groups by Shapes – rod-shaped, called bacilli – spiral, called spirilla or spirochetes – spherical, called cocci Spirochaeta: spiral Lactobacilli: rod-shaped Enterococci: spherical 18.4 Bacteria and Archaea • Main Groups by their need for oxygen. -

The Great-Grandmother of LUCA (Last Universal Common Ancestor)
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 4 June 2018 doi:10.20944/preprints201806.0035.v1 Be introduced to the First Universal Common Ancestor (FUCA): the great-grandmother of LUCA (Last Universal Common Ancestor) Francisco Prosdocimi1*, Marco V José2 and Sávio Torres de Farias3* 1 Laboratório de Biologia Teórica e de Sistemas, Instituto de Bioquímica Médica Leopoldo de Meis, Universidade Federal do Rio de Janeiro, Rio de Janeiro, Brasil. 2 Theoretical Biology Group, Instituto de Investigaciones Biomédicas, Universidad Nacional Autónoma de México, Ciudad Universitaria, 04510 CDMX, Mexico. 3 Laboratório de Genética Evolutiva Paulo Leminsk, Departamento de Biologia Molecular, Universidade Federal da Paraíba, João Pessoa, Paraíba, Brasil. * Correspondence: [email protected]; [email protected] Abstract The existence of a common ancestor to all living organisms in Earth is a necessary corollary of Darwin idea of common ancestry. The Last Universal Common Ancestor (LUCA) has been normally considered as the ancestor of cellular organisms that originated the three domains of life: Bacteria, Archaea and Eukarya. Recent studies about the nature of LUCA indicate that this first organism should present hundreds of genes and a complex metabolism. Trying to bring another of Darwin ideas into the origins of life discussion, we went back into the prebiotic chemistry trying to understand how LUCA could be originated 1 © 2018 by the author(s). Distributed under a Creative Commons CC BY license. Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 4 June 2018 doi:10.20944/preprints201806.0035.v1 under gradualist assumptions. Along this line of reasoning, it became clear to us that the definition of another ancestral should be of particular relevance to the understanding about the emergence of biological systems. -

Repurposing of Conserved Autophagy-Related Protein ATG8 in a Divergent Eukaryote Maude Lévêque, Hoa Mai Nguyen, Sébastien Besteiro
Repurposing of conserved autophagy-related protein ATG8 in a divergent eukaryote Maude Lévêque, Hoa Mai Nguyen, Sébastien Besteiro To cite this version: Maude Lévêque, Hoa Mai Nguyen, Sébastien Besteiro. Repurposing of conserved autophagy-related protein ATG8 in a divergent eukaryote. Communicative and Integrative Biology, Taylor & Francis Open, 2016, 9 (4), pp.e1197447. 10.1080/19420889.2016.1197447. hal-01824938 HAL Id: hal-01824938 https://hal.archives-ouvertes.fr/hal-01824938 Submitted on 1 Jun 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution - NonCommercial| 4.0 International License COMMUNICATIVE & INTEGRATIVE BIOLOGY 2016, VOL. 9, NO. 4, e1197447 (4 pages) http://dx.doi.org/10.1080/19420889.2016.1197447 ARTICLE ADDENDUM Repurposing of conserved autophagy-related protein ATG8 in a divergent eukaryote Maude F. Lev eque,^ Hoa Mai Nguyen, and Sebastien Besteiro DIMNP- UMR5235, CNRS, Universite de Montpellier, Montpellier, France ABSTRACT ARTICLE HISTORY Toxoplasma gondii and other apicomplexan parasites contain a peculiar non-photosynthetic plastid Received 18 May 2016 called the apicoplast, which is essential for their survival. The localization of autophagy-related Accepted 30 May 2016 protein ATG8 to the apicoplast in several apicomplexan species and life stages has recently been KEYWORDS described, and we have shown this protein is essential for proper inheritance of this complex plastid apicomplexa; apicoplast; into daughter cells during cell division. -

Beyond the Big Bang • the Amazon's Lost Civilizations • the Truth
SFI Bulletin winter 2006, vol. 21 #1 Beyond the Big Bang • The Amazon’s Lost Civilizations • The Truth Behind Lying The Bulletin of the Santa Fe Institute is published by SFI to keep its friends and supporters informed about its work. The Santa Fe Institute is a private, independent, multidiscipli- nary research and education center founded in 1984. Since its founding, SFI has devoted itself to creating a new kind of sci- entific research community, pursuing emerging synthesis in science. Operating as a visiting institution, SFI seeks to cat- alyze new collaborative, multidisciplinary research; to break down the barriers between the traditional disciplines; to spread its ideas and methodologies to other institutions; and to encourage the practical application of its results. Published by the Santa Fe Institute 1399 Hyde Park Road Santa Fe, New Mexico 87501, USA phone (505) 984-8800 fax (505) 982-0565 home page: http://www.santafe.edu Note: The SFI Bulletin may be read at the website: www.santafe.edu/sfi/publications/Bulletin/. If you would prefer to read the Bulletin on your computer rather than receive a printed version, contact Patrisia Brunello at 505/984-8800, Ext. 2700 or [email protected]. EDITORIAL STAFF: Ginger Richardson Lesley S. King Andi Sutherland CONTRIBUTORS: Brooke Harrington Janet Yagoda Shagam Julian Smith Janet Stites James Trefil DESIGN & PRODUCTION: Paula Eastwood PHOTO: ROBERT BUELTEMAN ©2004 BUELTEMAN PHOTO: ROBERT SFI Bulletin Winter 2006 TOCtable of contents 3 A Deceptively Simple Formula 2 How Life Began 3 From -

Eukaryote Cell Biology - Michelle Gehringer
FUNDAMENTALS OF BIOCHEMISTRY, CELL BIOLOGY AND BIOPHYSICS – Vol. II - Eukaryote Cell Biology - Michelle Gehringer EUKARYOTE CELL BIOLOGY Michelle Gehringer Department of Biochemistry and Microbiology, University of Port Elizabeth, South Africa Keywords: cell theory, cell diversity, eukaryote cell structure, nucleus, chromatin, DNA, organelles, mitochondria, chloroplasts, transcription, RNA, translation, ribosomes, cell cycle, interphase, mitosis, meiosis, signal transduction, growth regulation, cancer, oncogenesis. Contents 1. Introduction 1.1. The first cell 2. Origin of Eukaryotes 3. Cellular differentiation in multicellular organisms 3.1. Plants 3.2. Animals 4. Eukaryotic cell structure 5. Organization of eukaryotic cells 5.1. Plasma membrane 5.2. Extracellular matrices 5.3. Protein synthesis and transport 5.4. Cytoskeleton and movement 5.5. Nucleus 5.5.1 Genomes 5.5.2 Gene expression 5.5.3 Maintaining the genome 5.6. Organelles 6. The cell cycle 6.1. Mitosis 6.2. Meiosis 7. Regulation of cell growth 7.1. Signal transduction 7.2. Programmed cell death 7.3. CancerUNESCO – EOLSS 8. Experimental Models 8.1. Yeast SAMPLE CHAPTERS 8.2. Arabidopsis 8.3. Drosophila 8.4. The mouse 8.5. Cell culture 8.6. Separation of cellular contents 8.7. Tracing biochemical pathways 9. Future Investigations Glossary Bibliography ©Encyclopedia of Life Support Systems (EOLSS) FUNDAMENTALS OF BIOCHEMISTRY, CELL BIOLOGY AND BIOPHYSICS – Vol. II - Eukaryote Cell Biology - Michelle Gehringer Biographical Sketch Summary Cells form the basic unit of life on our planet. They are well organized systems which perform all the essential tasks of eating, respiring, replicating and excreting waste products. The first cells, which are thought to have evolved about 3.8 billion years ago, much resembled present day prokaryotes. -

Mixotrophic Protists Among Marine Ciliates and Dinoflagellates: Distribution, Physiology and Ecology
FACULTY OF SCIENCE UNIVERSITY OF COPENHAGEN PhD thesis Woraporn Tarangkoon Mixotrophic Protists among Marine Ciliates and Dinoflagellates: Distribution, Physiology and Ecology Academic advisor: Associate Professor Per Juel Hansen Submitted: 29/04/10 Contents List of publications 3 Preface 4 Summary 6 Sammenfating (Danish summary) 8 สรุป (Thai summary) 10 The sections and objectives of the thesis 12 Introduction 14 1) Mixotrophy among marine planktonic protists 14 1.1) The role of light, food concentration and nutrients for 17 the growth of marine mixotrophic planktonic protists 1.2) Importance of marine mixotrophic protists in the 20 planktonic food web 2) Marine symbiont-bearing dinoflagellates 24 2.1) Occurrence of symbionts in the order Dinophysiales 24 2.2) The spatial distribution of symbiont-bearing dinoflagellates in 27 marine waters 2.3) The role of symbionts and phagotrophy in dinoflagellates with symbionts 28 3) Symbiosis and mixotrophy in the marine ciliate genus Mesodinium 30 3.1) Occurrence of symbiosis in Mesodinium spp. 30 3.2) The distribution of marine Mesodinium spp. 30 3.3) The role of symbionts and phagotrophy in marine Mesodinium rubrum 33 and Mesodinium pulex Conclusion and future perspectives 36 References 38 Paper I Paper II Paper III Appendix-Paper IV Appendix-I Lists of publications The thesis consists of the following papers, referred to in the synthesis by their roman numerals. Co-author statements are attached to the thesis (Appendix-I). Paper I Tarangkoon W, Hansen G Hansen PJ (2010) Spatial distribution of symbiont-bearing dinoflagellates in the Indian Ocean in relation to oceanographic regimes. Aquat Microb Ecol 58:197-213.