TMEM9 CRISPR/Cas9 KO Plasmid (H): Sc-407509
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

TMEM9 CRISPR/Cas9 KO Plasmid (M): Sc-425907
SANTA CRUZ BIOTECHNOLOGY, INC. TMEM9 CRISPR/Cas9 KO Plasmid (m): sc-425907 BACKGROUND APPLICATIONS The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and TMEM9 CRISPR/Cas9 KO Plasmid (m) is recommended for the disruption of CRISPR-associated protein (Cas9) system is an adaptive immune response gene expression in mouse cells. defense mechanism used by archea and bacteria for the degradation of foreign genetic material (4,6). This mechanism can be repurposed for other 20 nt non-coding RNA sequence: guides Cas9 functions, including genomic engineering for mammalian systems, such as to a specific target location in the genomic DNA gene knockout (KO) (1,2,3,5). CRISPR/Cas9 KO Plasmid products enable the U6 promoter: drives gRNA scaffold: helps Cas9 identification and cleavage of specific genes by utilizing guide RNA (gRNA) expression of gRNA bind to target DNA sequences derived from the Genome-scale CRISPR Knock-Out (GeCKO) v2 library developed in the Zhang Laboratory at the Broad Institute (3,5). Termination signal Green Fluorescent Protein: to visually REFERENCES verify transfection CRISPR/Cas9 Knockout Plasmid CBh (chicken β-Actin 1. Cong, L., et al. 2013. Multiplex genome engineering using CRISPR/Cas hybrid) promoter: drives systems. Science 339: 819-823. 2A peptide: expression of Cas9 allows production of both Cas9 and GFP from the 2. Mali, P., et al. 2013. RNA-guided human genome engineering via Cas9. same CBh promoter Science 339: 823-826. Nuclear localization signal 3. Ran, F.A., et al. 2013. Genome engineering using the CRISPR-Cas9 system. Nuclear localization signal SpCas9 ribonuclease Nat. Protoc. 8: 2281-2308. -

Comparison of Methylation Capture Sequencing and Infinium Methylationepic Array in Peripheral Blood Mononuclear Cells
Shu et al. Epigenetics & Chromatin (2020) 13:51 https://doi.org/10.1186/s13072-020-00372-6 Epigenetics & Chromatin RESEARCH Open Access Comparison of methylation capture sequencing and Infnium MethylationEPIC array in peripheral blood mononuclear cells Chang Shu1,2, Xinyu Zhang1,2, Bradley E. Aouizerat3,4 and Ke Xu1,2* Abstract Background: Epigenome-wide association studies (EWAS) have been widely applied to identify methylation CpG sites associated with human disease. To date, the Infnium MethylationEPIC array (EPIC) is commonly used for high- throughput DNA methylation profling. However, the EPIC array covers only 30% of the human methylome. Methyla- tion Capture bisulfte sequencing (MC-seq) captures target regions of methylome and has advantages of extensive coverage in the methylome at an afordable price. Methods: Epigenome-wide DNA methylation in four peripheral blood mononuclear cell samples was profled by using SureSelectXT Methyl-Seq for MC-seq and EPIC platforms separately. CpG site-based reproducibility of MC-seq was assessed with DNA sample inputs ranging in quantity of high (> 1000 ng), medium (300–1000 ng), and low (150 ng–300 ng). To compare the performance of MC-seq and the EPIC arrays, we conducted a Pearson correlation and methylation value diference at each CpG site that was detected by both MC-seq and EPIC. We compared the percentage and counts in each CpG island and gene annotation between MC-seq and the EPIC array. Results: After quality control, an average of 3,708,550 CpG sites per sample were detected by MC-seq with DNA quantity > 1000 ng. Reproducibility of DNA methylation in MC-seq-detected CpG sites was high among samples with high, medium, and low DNA inputs (r > 0.96). -

Supplementary Table 2 Targets Description Uniprot NM 001001182 Bromodomain Adjacent to Zinc Finger Domain, 2B Q8CFP4 NM 001001309 Integrin Alpha 8 (Itga8), Mrna
Supplementary Table 2 targets description UniProt NM_001001182 bromodomain adjacent to zinc finger domain, 2B Q8CFP4 NM_001001309 integrin alpha 8 (Itga8), mRNA. A2ARA8 NM_001001321 solute carrier family 35, member D2 (Slc35d2), mRNA. Q762D5 NM_001003920 BR serine/threonine kinase 1 (Brsk1), mRNA. Q5RJI5 NM_001004468 transforming, acidic coiled-coil containing protein 2 Q9JJG0 NM_001005508 Rho GTPase activating protein 30 (Arhgap30), mRNA. Q640N3 NM_001008785 kelch repeat and BTB (POZ) domain containing 8 Q3UQV5 NM_001013022 outer dense fiber of sperm tails 3B (Odf3b), mRNA. Q5M8M2 NM_001013609 testis expressed gene 24 (Tex24), mRNA. Q5DP50 NM_001015681 RIKEN cDNA E130308A19 gene (E130308A19Rik), transcript Q8C4P0 NM_001017426 KDM1 lysine (K)-specific demethylase 6B (Kdm6b), mRNA. Q8K0Z1 NM_001024945 quiescin Q6 sulfhydryl oxidase 1 (Qsox1), transcript Q8BND5 NM_001025296 DNA fragmentation factor, alpha subunit (Dffa), Q8CA98 NM_001029983 mannosidase, alpha, class 1B, member 1 (Man1b1), mRNA. Q923C1 NM_001033269 eukaryotic translation initiation factor 4E family - NM_001033536 regulatory factor X, 7 (Rfx7), mRNA. Q8CB07 NM_001034037 RIKEN cDNA 1700024G13 gene (1700024G13Rik), mRNA. - NM_001034863 transmembrane protein 136 (Tmem136), mRNA. Q3TYE7 NM_001039088 SEH1-like (S. cerevisiae (Seh1l), transcript variant Q8R2U0 NM_001039644 ER degradation enhancer, mannosidase alpha-like 3 Q8R1X5 NM_001042592 arrestin domain containing 4 (Arrdc4), transcript Q9D6S6 NM_001045536 zinc finger, ZZ-type with EF hand domain 1 (Zzef1), Q5SSH7 NM_001081048 solute carrier family 25 (mitochondrial carrier), Q9DB41 NM_001081088 low density lipoprotein receptor-related protein 2 Q3V346 NM_001081152 nuclear protein in the AT region (Npat), mRNA. Q8BPV1 NM_001081205 NIPA-like domain containing 1 (Nipal1), mRNA. Q8BMW7 NM_001081206 diacylglycerol kinase, iota (Dgki), mRNA. - NM_001081366 vacuolar protein sorting 8 homolog (S. cerevisiae) Q8CIG5 NM_001081417 chromodomain helicase DNA binding protein 7 (Chd7), A2AJK6 NM_001081433 ankyrin repeat domain 44 (Ankrd44), mRNA. -

Molecular Targeting and Enhancing Anticancer Efficacy of Oncolytic HSV-1 to Midkine Expressing Tumors
University of Cincinnati Date: 12/20/2010 I, Arturo R Maldonado , hereby submit this original work as part of the requirements for the degree of Doctor of Philosophy in Developmental Biology. It is entitled: Molecular Targeting and Enhancing Anticancer Efficacy of Oncolytic HSV-1 to Midkine Expressing Tumors Student's name: Arturo R Maldonado This work and its defense approved by: Committee chair: Jeffrey Whitsett Committee member: Timothy Crombleholme, MD Committee member: Dan Wiginton, PhD Committee member: Rhonda Cardin, PhD Committee member: Tim Cripe 1297 Last Printed:1/11/2011 Document Of Defense Form Molecular Targeting and Enhancing Anticancer Efficacy of Oncolytic HSV-1 to Midkine Expressing Tumors A dissertation submitted to the Graduate School of the University of Cincinnati College of Medicine in partial fulfillment of the requirements for the degree of DOCTORATE OF PHILOSOPHY (PH.D.) in the Division of Molecular & Developmental Biology 2010 By Arturo Rafael Maldonado B.A., University of Miami, Coral Gables, Florida June 1993 M.D., New Jersey Medical School, Newark, New Jersey June 1999 Committee Chair: Jeffrey A. Whitsett, M.D. Advisor: Timothy M. Crombleholme, M.D. Timothy P. Cripe, M.D. Ph.D. Dan Wiginton, Ph.D. Rhonda D. Cardin, Ph.D. ABSTRACT Since 1999, cancer has surpassed heart disease as the number one cause of death in the US for people under the age of 85. Malignant Peripheral Nerve Sheath Tumor (MPNST), a common malignancy in patients with Neurofibromatosis, and colorectal cancer are midkine- producing tumors with high mortality rates. In vitro and preclinical xenograft models of MPNST were utilized in this dissertation to study the role of midkine (MDK), a tumor-specific gene over- expressed in these tumors and to test the efficacy of a MDK-transcriptionally targeted oncolytic HSV-1 (oHSV). -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

TMEM9 (NM 001288571) Human Tagged ORF Clone – RC236583
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC236583 TMEM9 (NM_001288571) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: TMEM9 (NM_001288571) Human Tagged ORF Clone Tag: Myc-DDK Symbol: TMEM9 Synonyms: DERM4; TMEM9A Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC236583 representing NM_001288571 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGAAGCTCTTATCTTTGGTGGCTGTGGTCGGGTGTTTGCTGGTGCCCCCAGCTGAAGCCAACAAGGTGA GGGAGGTGAGCCTGCAGCACCTGGTCACGACCACAGTGCACGGCCACCCTGTCTACAGGGCTGACTCAGA GAGTTCTGAAGATATCCGGTGCAAATGCATCTGTCCACCTTATAGAAACATCAGTGGGCACATTTACAAC CAGAATGTATCCCAGAAGGACTGCAACTGCCTGCACGTGGTGGAGCCCATGCCAGTGCCTGGCCATGACG TGGAGGCCTACTGCCTGCTGTGCGAGTGCAGGTACGAGGAGCGCAGCACCACCACCATCAAGGTCATCAT TGTCATCTACCTGTCCGTGGTGGGTGCCCTGTTGCTCTACATGGCCTTCCTGATGCTGGTGGACCCTCTG ATCCGAAAGCCGGATGCATATACTGAGCAACTGCACAATGAGGAGGAGAATGAGGATGCTCGCTCTATGG CAGCAGCTGCTGCATCCCTCGGGGGACCCCGAGCAAACACAGTCCTGGAGCGTGTGGAAGGTGCCCAGCA GCGGTGGAAGCTGCAGGTGCAGGAGCAGCGGAAGACAGTCTTCGATCGGCACAAGATGCTCAGC ACGCGTACGCGGCCGCTCGAGCAGAAACTCATCTCAGAAGAGGATCTGGCAGCAAATGATATCCTGGATT ACAAGGATGACGACGATAAGGTTTAA Protein Sequence: >RC236583 representing NM_001288571 Red=Cloning site Green=Tags(s) MKLLSLVAVVGCLLVPPAEANKVREVSLQHLVTTTVHGHPVYRADSESSEDIRCKCICPPYRNISGHIYN -

Supplementary Material
Supplementary material Gene Expression (mRNA) Markers for Differentiating between Malignant and Benign Follicular Thyroid Tumours Bartosz Wojtas 1,2,†, Aleksandra Pfeifer 1,3,†, Malgorzata Oczko-Wojciechowska 1, Jolanta Krajewska 1, Agnieszka Czarniecka 4, Aleksandra Kukulska 1, Markus Eszlinger 5, Thomas Musholt 6, Tomasz Stokowy 1,3,7, Michal Swierniak 1,8, Ewa Stobiecka 9, Ewa Chmielik 9, Dagmara Rusinek 1, Tomasz Tyszkiewicz 1, Monika Halczok 1, Steffen Hauptmann 10, Dariusz Lange 9, Michal Jarzab 11, Ralf Paschke 12 and Barbara Jarzab 1,* 1 Department of Nuclear Medicine and Endocrine Oncology, Maria Sklodowska-Curie Institute— Oncology Center, Gliwice Branch, Wybrzeze Armii Krajowej 15, 44-101 Gliwice, Poland; 2 Laboratory of Molecular Neurobiology, Neurobiology Center, Nencki Institute of Experimental Biology, Pasteura 3, 02-093 Warsaw, Poland 3 Faculty of Automatic Control, Electronics and Computer Science, Silesian University of Technology, Akademicka 2A, 44-100 Gliwice, Poland 4 The Oncologic and Reconstructive Surgery Clinic, Maria Sklodowska-Curie Institute—Oncology Center, Gliwice Branch, Wybrzeze Armii Krajowej 15, 44-101 Gliwice, Poland; 5 Department of Oncology & Arnie Charbonneau Cancer Institute, Cumming School of Medicine, University of Calgary, Calgary, Alberta T2N 4N1 6 Department of General, Visceral, and Transplantation Surgery, University Medical Center of the Johannes Gutenberg University, D55099 Mainz, Germany 7 Department of Clinical Science, University of Bergen, 5020 Bergen, Norway 8 Genomic Medicine, Department -

Interaction Mapping of Endoplasmic Reticulum Ubiquitin Ligases Identifies Modulators of Innate Immune Signalling
RESEARCH ARTICLE Interaction mapping of endoplasmic reticulum ubiquitin ligases identifies modulators of innate immune signalling Emma J Fenech1†, Federica Lari1, Philip D Charles2, Roman Fischer2, Marie Lae´ titia-The´ ze´ nas2, Katrin Bagola1‡, Adrienne W Paton3, James C Paton3, Mads Gyrd-Hansen1, Benedikt M Kessler2,4, John C Christianson1,5,6* 1Ludwig Institute for Cancer Research, Nuffield Department of Medicine, University of Oxford, Oxford, United Kingdom; 2TDI Mass Spectrometry Laboratory, Target Discovery Institute, University of Oxford, Oxford, United Kingdom; 3Research Centre for Infectious Diseases, Department of Molecular and Biomedical Science, University of Adelaide, Adelaide, Australia; 4Chinese Academy of Medical Sciences Oxford Institute, Nuffield Department of Medicine, University of Oxford, Oxford, United Kingdom; 5Nuffield Department of Orthopaedics, Rheumatology, and Musculoskeletal Sciences, University of Oxford, Botnar Research Centre, Oxford, United Kingdom; 6Oxford Centre for Translational Myeloma Research, University of Oxford, Botnar Research Centre, Oxford, United Kingdom *For correspondence: [email protected]. Abstract Ubiquitin ligases (E3s) embedded in the endoplasmic reticulum (ER) membrane uk regulate essential cellular activities including protein quality control, calcium flux, and sterol Present address: †Department homeostasis. At least 25 different, transmembrane domain (TMD)-containing E3s are predicted to of Molecular Genetics, be ER-localised, but for most their organisation and cellular -
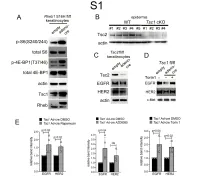
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test). -

Sheet1 Page 1 Gene Symbol Gene Description Entrez Gene ID
Sheet1 RefSeq ID ProbeSets Gene Symbol Gene Description Entrez Gene ID Sequence annotation Seed matches location(s) Ago-2 binding specific enrichment (replicate 1) Ago-2 binding specific enrichment (replicate 2) OE lysate log2 fold change (replicate 1) OE lysate log2 fold change (replicate 2) Probability NM_022823 218843_at FNDC4 Homo sapiens fibronectin type III domain containing 4 (FNDC4), mRNA. 64838 TR(1..1649)CDS(367..1071) 1523..1530 3.73 1.77 -1.91 -0.39 1 NM_003919 204688_at SGCE Homo sapiens sarcoglycan, epsilon (SGCE), transcript variant 2, mRNA. 8910 TR(1..1709)CDS(112..1425) 1495..1501 3.09 1.56 -1.02 -0.27 1 NM_006982 206837_at ALX1 Homo sapiens ALX homeobox 1 (ALX1), mRNA. 8092 TR(1..1320)CDS(5..985) 916..923 2.99 1.93 -0.19 -0.33 1 NM_019024 233642_s_at HEATR5B Homo sapiens HEAT repeat containing 5B (HEATR5B), mRNA. 54497 TR(1..6792)CDS(97..6312) 5827..5834,4309..4315 3.28 1.51 -0.92 -0.23 1 NM_018366 223431_at CNO Homo sapiens cappuccino homolog (mouse) (CNO), mRNA. 55330 TR(1..1546)CDS(96..749) 1062..1069,925..932 2.89 1.51 -1.2 -0.41 1 NM_032436 226194_at C13orf8 Homo sapiens chromosome 13 open reading frame 8 (C13orf8), mRNA. 283489 TR(1..3782)CDS(283..2721) 1756..1762,3587..3594,1725..1731,3395..3402 2.75 1.72 -1.38 -0.34 1 NM_031450 221534_at C11orf68 Homo sapiens chromosome 11 open reading frame 68 (C11orf68), mRNA. 83638 TR(1..1568)CDS(153..908) 967..973 3.07 1.35 -0.72 -0.06 1 NM_033318 225795_at,225794_s_at C22orf32 Homo sapiens chromosome 22 open reading frame 32 (C22orf32), mRNA. -

Supplementary Material Peptide-Conjugated Oligonucleotides Evoke Long-Lasting Myotonic Dystrophy Correction in Patient-Derived C
Supplementary material Peptide-conjugated oligonucleotides evoke long-lasting myotonic dystrophy correction in patient-derived cells and mice Arnaud F. Klein1†, Miguel A. Varela2,3,4†, Ludovic Arandel1, Ashling Holland2,3,4, Naira Naouar1, Andrey Arzumanov2,5, David Seoane2,3,4, Lucile Revillod1, Guillaume Bassez1, Arnaud Ferry1,6, Dominic Jauvin7, Genevieve Gourdon1, Jack Puymirat7, Michael J. Gait5, Denis Furling1#* & Matthew J. A. Wood2,3,4#* 1Sorbonne Université, Inserm, Association Institut de Myologie, Centre de Recherche en Myologie, CRM, F-75013 Paris, France 2Department of Physiology, Anatomy and Genetics, University of Oxford, South Parks Road, Oxford, UK 3Department of Paediatrics, John Radcliffe Hospital, University of Oxford, Oxford, UK 4MDUK Oxford Neuromuscular Centre, University of Oxford, Oxford, UK 5Medical Research Council, Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, UK 6Sorbonne Paris Cité, Université Paris Descartes, F-75005 Paris, France 7Unit of Human Genetics, Hôpital de l'Enfant-Jésus, CHU Research Center, QC, Canada † These authors contributed equally to the work # These authors shared co-last authorship Methods Synthesis of Peptide-PMO Conjugates. Pip6a Ac-(RXRRBRRXRYQFLIRXRBRXRB)-CO OH was synthesized and conjugated to PMO as described previously (1). The PMO sequence targeting CUG expanded repeats (5′-CAGCAGCAGCAGCAGCAGCAG-3′) and PMO control reverse (5′-GACGACGACGACGACGACGAC-3′) were purchased from Gene Tools LLC. Animal model and ASO injections. Experiments were carried out in the “Centre d’études fonctionnelles” (Faculté de Médecine Sorbonne University) according to French legislation and Ethics committee approval (#1760-2015091512001083v6). HSA-LR mice are gift from Pr. Thornton. The intravenous injections were performed by single or multiple administrations via the tail vein in mice of 5 to 8 weeks of age.