Mouse Wasf1 Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification of Novel E2F1 Target Genes Regulated in Cell Cycle
Oncogene (2006) 25, 1786–1798 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc ORIGINAL ARTICLE Identification of novel E2F1 target genes regulated in cell cycle-dependent and independent manners R Iwanaga1,3, H Komori1, S Ishida2, N Okamura3, K Nakayama4, KI Nakayama5 and K Ohtani1 1Human Gene Sciences Center, Tokyo Medical and Dental University, Bunkyo-ku, Tokyo, Japan; 2Division of Pharmacology, National Institute of Health Sciences, Setagaya-ku, Tokyo, Japan; 3Laboratory of Microbiology and Immunology, Graduate School of Health Sciences, Tokyo Medical and Dental University, Bunkyo-ku, Tokyo, Japan; 4Department of Developmental Biology, Center for Translational and Advanced Animal Research on Human Disease, Graduate School of Medicine, Tohoku University, Aoba-ku, Sendai, Japan and 5Department of Molecular and Cellular Biology, Medical Institute of Bioregulation, Kyushu University, Fukuoka, Japan The transcription factor E2F mediates cell cycle-depen- cell cycle progression (Nevins et al., 1997; Dyson, 1998; dent expression of genes important for cell proliferation in Trimarchi and Lees, 2002). The transcriptional ability of response to growth stimulation. To further understand the E2F is cell cycle-regulated mainly through association role of E2F, we utilized a sensitive subtraction method to with the retinoblastoma tumor suppressor family of explore new E2F1 targets, which are expressed at low proteins pRb, p107 and p130. During the progression of levels and might have been unrecognized in previous cells from G1 to S phase, G1 cyclin-dependent kinases studies. We identified 33 new E2F1-inducible genes, (cdks)phosphorylate and dissociate the pRb family including checkpoint genes Claspin and Rad51ap1, and proteins from E2F, resulting in the activation of a group four genes with unknown function required for cell cycle of genes required for progression into the S phase. -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -
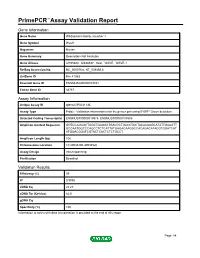
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name WAS protein family, member 1 Gene Symbol Wasf1 Organism Mouse Gene Summary Description Not Available Gene Aliases AI195380, AI838537, Scar, WAVE, WAVE-1 RefSeq Accession No. NC_000076.6, NT_039492.8 UniGene ID Mm.41353 Ensembl Gene ID ENSMUSG00000019831 Entrez Gene ID 83767 Assay Information Unique Assay ID qMmuCIP0031135 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSMUST00000019975, ENSMUST00000105509 Amplicon Context Sequence GGTCCAGAGCTGGCTGAGGATGACGCTGACCTCCTACACAAGCATATTGAAGTT GCCAATGGCCCAGCCTCTCATTATGAGACAAGGCCACAGACATACGTGGATCAT ATGGACGGATCGTACTCACTCTCTGCCT Amplicon Length (bp) 106 Chromosome Location 10:40933200-40934526 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 99 R2 0.9998 cDNA Cq 22.23 cDNA Tm (Celsius) 82.5 gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Wasf1, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

(SCD) Among Indian Patients Using Gene Expression Data Analysis
www.bioinformation.net Hypothesis Volume 14(7) Identification of therapeutic targets for inflammation in sickle cell disease (SCD) among Indian patients using gene expression data analysis Ipsita Das1, Hrishikesh Mishra2, Prafulla K. Khodiar1, Pradeep K. Patra1* 1Pt. J.N.M. Medical College, Raipur, India; 2Sickle Cell Institute Chhattisgarh, Raipur, India; Pradeep K Patra – E-mail: [email protected]; Phone: 91-771-2890012; *Corresponding author Received July 24, 2018; Revised July 29, 2018; Accepted July 29, 2018; Published July 31, 2018 doi: 10.6026/97320630014408 Abstract: Sickle cell disease (SCD) is life-threatening hemoglobinopathy prevalent in India, Sub-Saharan Africa and Middle East. Inflammation plays a pivotal role in disease process and involves intricate interaction among leukocytes, platelets, sickle erythrocytes and vascular endothelium. Available disease modifying therapies are hydroxyl-urea and blood transfusion. Therefore, it is of interest to develop improved pharmacological agents for SCD. We report up-regulated genes in steady state and vaso-occlusive crisis using analysis of gene expression data obtained by microarray experiment for SCD as potential targets. The association of these targets with inflammation in pathway analysis is also documented. Keywords: Sickle cell disease, vaso-occlusive crisis, inflammation, gene expression, pathophysiology, drug targets. Background: vascular endothelium. Leukocytosis and activation of neutrophils Sickle cell disease (SCD) is a life threatening hemoglobin disorder and monocytes further increases vascular inflammation and affecting about 5% of world population and is prevalent in India endothelial damage and plays as a trigger for VOC [3]. In and other parts of the world including Sub-Saharan Africa and addition to vaso-occlusion, inflammation plays a pivotal role in Middle East. -

The Human Gene Connectome As a Map of Short Cuts for Morbid Allele Discovery
The human gene connectome as a map of short cuts for morbid allele discovery Yuval Itana,1, Shen-Ying Zhanga,b, Guillaume Vogta,b, Avinash Abhyankara, Melina Hermana, Patrick Nitschkec, Dror Friedd, Lluis Quintana-Murcie, Laurent Abela,b, and Jean-Laurent Casanovaa,b,f aSt. Giles Laboratory of Human Genetics of Infectious Diseases, Rockefeller Branch, The Rockefeller University, New York, NY 10065; bLaboratory of Human Genetics of Infectious Diseases, Necker Branch, Paris Descartes University, Institut National de la Santé et de la Recherche Médicale U980, Necker Medical School, 75015 Paris, France; cPlateforme Bioinformatique, Université Paris Descartes, 75116 Paris, France; dDepartment of Computer Science, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel; eUnit of Human Evolutionary Genetics, Centre National de la Recherche Scientifique, Unité de Recherche Associée 3012, Institut Pasteur, F-75015 Paris, France; and fPediatric Immunology-Hematology Unit, Necker Hospital for Sick Children, 75015 Paris, France Edited* by Bruce Beutler, University of Texas Southwestern Medical Center, Dallas, TX, and approved February 15, 2013 (received for review October 19, 2012) High-throughput genomic data reveal thousands of gene variants to detect a single mutated gene, with the other polymorphic genes per patient, and it is often difficult to determine which of these being of less interest. This goes some way to explaining why, variants underlies disease in a given individual. However, at the despite the abundance of NGS data, the discovery of disease- population level, there may be some degree of phenotypic homo- causing alleles from such data remains somewhat limited. geneity, with alterations of specific physiological pathways under- We developed the human gene connectome (HGC) to over- come this problem. -

(WASF1) (NM 003931) Human Untagged Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for SC117699 WAVE 1 (WASF1) (NM_003931) Human Untagged Clone Product data: Product Type: Expression Plasmids Product Name: WAVE 1 (WASF1) (NM_003931) Human Untagged Clone Tag: Tag Free Symbol: WASF1 Synonyms: NEDALVS; SCAR1; WAVE; WAVE1 Vector: pCMV6-XL5 E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: None This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 3 WAVE 1 (WASF1) (NM_003931) Human Untagged Clone – SC117699 Fully Sequenced ORF: >OriGene ORF within SC117699 sequence for NM_003931 edited (data generated by NextGen Sequencing) ATGCCGCTAGTGAAAAGAAACATCGATCCTAGGCACTTGTGCCACACAGCACTGCCTAGA GGCATTAAGAATGAACTGGAATGTGTAACCAATATTTCCTTGGCAAATATAATTAGACAA CTAAGTAGCCTAAGTAAATATGCTGAAGATATATTTGGAGAATTATTCAATGAAGCACAT AGTTTTTCCTTCAGAGTCAACTCATTGCAAGAACGTGTGGACCGTTTATCTGTTAGTGTT ACACAGCTTGATCCAAAGGAAGAAGAATTGTCTTTGCAAGATATAACAATGAGGAAAGCT TTCCGAAGTTCTACAATTCAAGACCAGCAGCTTTTCGATCGCAAGACTTTGCCTATTCCA TTACAGGAGACGTACGATGTTTGTGAACAGCCTCCACCTCTCAATATACTCACTCCTTAT AGAGATGATGGTAAAGAAGGTCTGAAGTTTTATACCAATCCTTCGTATTTCTTTGATCTA TGGAAAGAAAAAATGTTGCAAGATACAGAGGATAAGAGGAAGGAAAAGAGGAAGCAGAAG CAGAAAAATCTAGATCGTCCTCATGAACCAGAAAAAGTGCCAAGAGCACCTCATGACAGG CGGCGAGAATGGCAGAAGCTGGCCCAAGGTCCAGAGCTGGCTGAAGATGATGCTAATCTC -

Dissecting the Genetics of Human Communication
DISSECTING THE GENETICS OF HUMAN COMMUNICATION: INSIGHTS INTO SPEECH, LANGUAGE, AND READING by HEATHER ASHLEY VOSS-HOYNES Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Epidemiology and Biostatistics CASE WESTERN RESERVE UNIVERSITY January 2017 CASE WESTERN RESERVE UNIVERSITY SCHOOL OF GRADUATE STUDIES We herby approve the dissertation of Heather Ashely Voss-Hoynes Candidate for the degree of Doctor of Philosophy*. Committee Chair Sudha K. Iyengar Committee Member William Bush Committee Member Barbara Lewis Committee Member Catherine Stein Date of Defense July 13, 2016 *We also certify that written approval has been obtained for any proprietary material contained therein Table of Contents List of Tables 3 List of Figures 5 Acknowledgements 7 List of Abbreviations 9 Abstract 10 CHAPTER 1: Introduction and Specific Aims 12 CHAPTER 2: Review of speech sound disorders: epidemiology, quantitative components, and genetics 15 1. Basic Epidemiology 15 2. Endophenotypes of Speech Sound Disorders 17 3. Evidence for Genetic Basis Of Speech Sound Disorders 22 4. Genetic Studies of Speech Sound Disorders 23 5. Limitations of Previous Studies 32 CHAPTER 3: Methods 33 1. Phenotype Data 33 2. Tests For Quantitative Traits 36 4. Analytical Methods 42 CHAPTER 4: Aim I- Genome Wide Association Study 49 1. Introduction 49 2. Methods 49 3. Sample 50 5. Statistical Procedures 53 6. Results 53 8. Discussion 71 CHAPTER 5: Accounting for comorbid conditions 84 1. Introduction 84 2. Methods 86 3. Results 87 4. Discussion 105 CHAPTER 6: Hypothesis driven pathway analysis 111 1. Introduction 111 2. Methods 112 3. Results 116 4. -

Orbitofrontal Neuroadaptations and Cross-Species Synaptic Biomarkers in Heavy-Drinking Macaques
3646 • The Journal of Neuroscience, March 29, 2017 • 37(13):3646–3660 Neurobiology of Disease Orbitofrontal Neuroadaptations and Cross-Species Synaptic Biomarkers in Heavy-Drinking Macaques X Sudarat Nimitvilai,1* Joachim D. Uys,2* John J. Woodward,1,3 Patrick K. Randall,3 Lauren E. Ball,2 X Robert W. Williams,4 XByron C. Jones,4 X Lu Lu,4 X Kathleen A. Grant,5 and Patrick J. Mulholland1,3 Departments of 1Neuroscience, 2Cell and Molecular Pharmacology, and 3Psychiatry and Behavioral Sciences, Medical University of South Carolina, Charleston, South Carolina 29425, 4Department of Genetics, Genomics and Informatics, University of Tennessee Health Science Center, Memphis, Tennessee 38120, and 5Department of Behavioral Neuroscience, Oregon Health and Science University, Portland, Oregon 97239 Cognitive impairments, uncontrolled drinking, and neuropathological cortical changes characterize alcohol use disorder. Dysfunction of the orbitofrontal cortex (OFC), a critical cortical subregion that controls learning, decision-making, and prediction of reward outcomes, contributes to executive cognitive function deficits in alcoholic individuals. Electrophysiological and quantitative synaptomics tech- niques were used to test the hypothesis that heavy drinking produces neuroadaptations in the macaque OFC. Integrative bioinformatics and reverse genetic approaches were used to identify and validate synaptic proteins with novel links to heavy drinking in BXD mice. In drinking monkeys, evoked firing of OFC pyramidal neurons was reduced, whereas the amplitude and frequency of postsynaptic currents were enhanced compared with controls. Bath application of alcohol reduced evoked firing in neurons from control monkeys, but not drinking monkeys. Profiling of the OFC synaptome identified alcohol-sensitive proteins that control glutamate release (e.g., SV2A, synaptogyrin-1) and postsynaptic signaling (e.g., GluA1, PRRT2) with no changes in synaptic GABAergic proteins. -

WASF1 Rabbit Pab
Leader in Biomolecular Solutions for Life Science WASF1 Rabbit pAb Catalog No.: A14624 Basic Information Background Catalog No. The protein encoded by this gene, a member of the Wiskott-Aldrich syndrome protein A14624 (WASP)-family, plays a critical role downstream of Rac, a Rho-family small GTPase, in regulating the actin cytoskeleton required for membrane ruffling. It has been shown to Observed MW associate with an actin nucleation core Arp2/3 complex while enhancing actin 62kDa polymerization in vitro. Wiskott-Aldrich syndrome is a disease of the immune system, likely due to defects in regulation of actin cytoskeleton. Multiple alternatively spliced Calculated MW transcript variants encoding the same protein have been found for this gene. 61kDa Category Primary antibody Applications WB Cross-Reactivity Human Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 8936 Q92558 Immunogen A synthetic peptide corresponding to a sequence within amino acids 400 to the C- terminus of human WASF1 (NP_001020107.1). Synonyms WASF1;SCAR1;WAVE;WAVE1 Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using WASF1 antibody (A14624) at 1:3000 dilution. Secondary antibody: HRP Goat Anti-Rabbit IgG (H+L) (AS014) at 1:10000 dilution. Lysates/proteins: 25ug per lane. Blocking buffer: 3% nonfat dry milk in TBST. Detection: ECL Basic Kit (RM00020). Exposure time: 60s. Antibody | Protein | ELISA Kits | Enzyme | NGS | Service For research use only. -

Cellular Functions of WASP Family Proteins at a Glance Olga Alekhina1, Ezra Burstein2,3 and Daniel D
© 2017. Published by The Company of Biologists Ltd | Journal of Cell Science (2017) 130, 2235-2241 doi:10.1242/jcs.199570 CELL SCIENCE AT A GLANCE Cellular functions of WASP family proteins at a glance Olga Alekhina1, Ezra Burstein2,3 and Daniel D. Billadeau1,4,5,* ABSTRACT WASP family members in promoting actin dynamics at the Proteins of the Wiskott–Aldrich syndrome protein (WASP) family centrosome, influencing nuclear shape and membrane remodeling function as nucleation-promoting factors for the ubiquitously events leading to the generation of autophagosomes. Interestingly, expressed Arp2/3 complex, which drives the generation of several WASP family members have also been observed in the branched actin filaments. Arp2/3-generated actin regulates diverse nucleus where they directly influence gene expression by serving cellular processes, including the formation of lamellipodia and as molecular platforms for the assembly of epigenetic and filopodia, endocytosis and/or phagocytosis at the plasma transcriptional machinery. In this Cell Science at a Glance article membrane, and the generation of cargo-laden vesicles from and accompanying poster, we provide an update on the subcellular organelles including the Golgi, endoplasmic reticulum (ER) and the roles of WHAMM, JMY and WASH (also known as WASHC1), as endo-lysosomal network. Recent studies have also identified roles for well as their mechanisms of regulation and emerging functions within the cell. KEY WORDS: WASP, N-WASP, WAVE, WHAMM, WASH, JMY, 1Division of Oncology Research, College of Medicine, Mayo Clinic, Rochester, MN WHAMY, Arp2/3, Actin 55905, USA. 2Department of Internal Medicine, UT Southwestern Medical Center, Dallas, TX 75390-9151, USA. -

Effect of Myoferlin Depletion on Breast Cancer Cell Motility THESIS
Effect of Myoferlin Depletion on Breast Cancer Cell Motility THESIS Presented in Partial Fulfillment of the Requirements for the Degree Master of Science in the Graduate School of The Ohio State University By Leonithas Ioannis Volakis Graduate Program in Biomedical Engineering The Ohio State University 2011 Master's Examination Committee: Professor Douglas A. Kniss, Advisor Professor Samir N. Ghadiali Professor Heather M. Powell Copyright by Leonithas Ioannis Volakis 2011 Abstract Cancer metastasis requires tumor cell invasion, migration, and proliferation. Cell migration involves a complex series of processes that extend membrane protrusions to form substrate adhesions that cause cytoskeleton contraction within the cell. Further, cell migration is central to tissue repair, regeneration, development, cancer, and inflammation. The ferlins, an evolutionary conserved protein family, have been implicated as critical to maintaining plasma membranes. Specifically, myoferlin (MYOF), a mammalian ferlin, has been shown as responsible for membrane exocytosis/endocytosis and myoblast fusion. More recent studies have demonstrated that MYOF affects the anti-angiogenic response of endothelial cells and impacts the invasive ability of MDA-MB-231 breast cancer cells. It has also been noted that ablating MYOF using RNAi strategies results in consistent cell proliferation rates and a mesenchymal to epithelial phenotypic transformation. In this study, we present quantified 2D morphologic and migration differences in MDA-MB-231 wild-type (231WT) and RNAi-mediated MYOF-deficient (231MYOFKD) human breast adenocarcinoma (MDA-MB-231) cells. Morphometrics found that MYOF deficiency led to significant differences in lamellipodia number and surface area, filopodia length, and cell surface area. These marked changes could suggest that MYOF plays a role in regulating cytoskeletal arrangement in breast cancer cells. -

Functional Genomics of Cohesin Acetyltransferases in Human Cells
FUNCTIONAL GENOMICS OF COHESIN ACETYLTRANSFERASES IN HUMAN CELLS by Sadia Rahman A Dissertation Presented to the Faculty of the Louis V. Gerstner, Jr. Graduate School of Biomedical Sciences, Memorial Sloan Kettering Cancer Center in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy New York, NY March, 2014 __________________________ _________________ Prasad V. Jallepalli, MD, PhD Date Dissertation Mentor ABSTRACT Accurate chromosome segregation during cell division requires that sister chromatids be physically linked from the time of their replication until their separation at anaphase. The cohesin complex, consisting of SMC1, SMC3, RAD21 and SCC3 arranges to form a ring-shaped structure that holds sister chromatids together. Acetylation of the cohesin SMC3 subunit by acetyltransferases ESCO1 and ESCO2 is essential for cohesion establishment. In addition to cohesion, cohesin also has roles in gene expression through its regulation of chromatin architecture. Acetylation of cohesin by ESCO1/2 is regulated temporally and spatially. In human cells, it begins in G1 phase, rises in S-phase and persists until mitosis. The reaction occurs only on DNA-bound cohesin and SMC3 is quickly deacetylated after cohesin is removed from DNA. In this study, we map genome-wide ESCO1/2 and AcSMC3 sites by ChIP- Seq, study their regulation, and contribution to cohesion and gene expression functions. Genome-wide mapping of ESCO1/2 reveals that they differ in their distribution: ESCO1 has many discrete binding sites that largely overlap with cohesin/CTCF sites, whereas ESCO2 has few sites of enrichment. A monoclonal antibody against the acetylated form of cohesin was also generated in this study to map cohesin acetylation, and this shows that cohesin is already acetylated in G1 at the majority of its sites and that this depends on ESCO1.