UI Health Care Letterhead Template
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Exceptional Conservation of Horse–Human Gene Order on X Chromosome Revealed by High-Resolution Radiation Hybrid Mapping
Exceptional conservation of horse–human gene order on X chromosome revealed by high-resolution radiation hybrid mapping Terje Raudsepp*†, Eun-Joon Lee*†, Srinivas R. Kata‡, Candice Brinkmeyer*, James R. Mickelson§, Loren C. Skow*, James E. Womack‡, and Bhanu P. Chowdhary*¶ʈ *Department of Veterinary Anatomy and Public Health, ‡Department of Veterinary Pathobiology, College of Veterinary Medicine, and ¶Department of Animal Science, College of Agriculture and Life Science, Texas A&M University, College Station, TX 77843; and §Department of Veterinary Pathobiology, University of Minnesota, 295f AS͞VM, St. Paul, MN 55108 Contributed by James E. Womack, December 30, 2003 Development of a dense map of the horse genome is key to efforts ciated with the traits, once they are mapped by genetic linkage aimed at identifying genes controlling health, reproduction, and analyses with highly polymorphic markers. performance. We herein report a high-resolution gene map of the The X chromosome is the most conserved mammalian chro- horse (Equus caballus) X chromosome (ECAX) generated by devel- mosome (18, 19). Extensive comparisons of structure, organi- oping and typing 116 gene-specific and 12 short tandem repeat zation, and gene content of this chromosome in evolutionarily -markers on the 5,000-rad horse ؋ hamster whole-genome radia- diverse mammals have revealed a remarkable degree of conser tion hybrid panel and mapping 29 gene loci by fluorescence in situ vation (20–22). Until now, the chromosome has been best hybridization. The human X chromosome sequence was used as a studied in humans and mice, where the focus of research has template to select genes at 1-Mb intervals to develop equine been the intriguing patterns of X inactivation and the involve- orthologs. -

OXSR1 (A-4): Sc-271707
SANTA CRUZ BIOTECHNOLOGY, INC. OXSR1 (A-4): sc-271707 BACKGROUND APPLICATIONS Oxidative stress-responsive 1 protein (OXSR1), a protein of 527 amino acids, OXSR1 (A-4) is recommended for detection of OXSR1 of mouse, rat and belongs to the STE20 subfamily. OXSR1 is one of two human homologs of human origin by Western Blotting (starting dilution 1:100, dilution range Fray, a serine/threonine kinase expressed in Drosophila. OXSR1 binds to and 1:100-1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein phosphorylates p21-activated protein kinase (PAK1) and regulates downstream (1 ml of cell lysate)], immunofluorescence (starting dilution 1:50, dilution kinases in response to environmental stress. Endogenous OXSR1 is activated range 1:50-1:500) and solid phase ELISA (starting dilution 1:30, dilution only by osmotic stresses, notably sorbitol and to a lesser extent NaCl. OXSR1 range 1:30-1:3000). may also play a role in regulating the Actin cytoskeleton. The chloride channel Suitable for use as control antibody for OXSR1 siRNA (h): sc-61273, OXSR1 proteins SLC12A1, SLC12A2 and SLC12A6 isoform 2 interact with OXSR1, siRNA (m): sc-61274, OXSR1 shRNA Plasmid (h): sc-61273-SH, OXSR1 but SLC12A4 and SLC12A7 do not. The WNK1 and WNK4 protein kinases shRNA Plasmid (m): sc-61274-SH, OXSR1 shRNA (h) Lentiviral Particles: activate OXSR1 by phosphorlating its T-loop. The OXSR1 protein is widely sc-61273-V and OXSR1 shRNA (m) Lentiviral Particles: sc-61274-V. expressed in mammalian tissues. Molecular Weight of OXSR1: 58 kDa. REFERENCES Positive Controls: OXSR1 (h3): 293 Lysate: sc-158793, HeLa whole cell lysate: 1. -

Snapshot: Formins Christian Baarlink, Dominique Brandt, and Robert Grosse University of Marburg, Marburg 35032, Germany
SnapShot: Formins Christian Baarlink, Dominique Brandt, and Robert Grosse University of Marburg, Marburg 35032, Germany Formin Regulators Localization Cellular Function Disease Association DIAPH1/DIA1 RhoA, RhoC Cell cortex, Polarized cell migration, microtubule stabilization, Autosomal-dominant nonsyndromic deafness (DFNA1), myeloproliferative (mDia1) phagocytic cup, phagocytosis, axon elongation defects, defects in T lymphocyte traffi cking and proliferation, tumor cell mitotic spindle invasion, defects in natural killer lymphocyte function DIAPH2 Cdc42 Kinetochore Stable microtubule attachment to kinetochore for Premature ovarian failure (mDia3) chromosome alignment DIAPH3 Rif, Cdc42, Filopodia, Filopodia formation, removing the nucleus from Increased chromosomal deletion of gene locus in metastatic tumors (mDia2) Rac, RhoB, endosomes erythroblast, endosome motility, microtubule DIP* stabilization FMNL1 (FRLα) Cdc42 Cell cortex, Phagocytosis, T cell polarity Overexpression is linked to leukemia and non-Hodgkin lymphoma microtubule- organizing center FMNL2/FRL3/ RhoC ND Cell motility Upregulated in metastatic colorectal cancer, chromosomal deletion is FHOD2 associated with mental retardation FMNL3/FRL2 Constituently Stress fi bers ND ND active DAAM1 Dishevelled Cell cortex Planar cell polarity ND DAAM2 ND ND ND Overexpressed in schizophrenia patients Human (Mouse) FHOD1 ROCK Stress fi bers Cell motility FHOD3 ND Nestin, sarcomere Organizing sarcomeres in striated muscle cells Single-nucleotide polymorphisms associated with type 1 diabetes -

Cooperative Gene Regulation by Microrna Pairs and Their
Published online 29 May 2014 Nucleic Acids Research, 2014, Vol. 42, No. 12 7539–7552 doi: 10.1093/nar/gku465 Cooperative gene regulation by microRNA pairs and their identification using a computational workflow Ulf Schmitz1,*, Xin Lai2, Felix Winter1, Olaf Wolkenhauer1,3, Julio Vera2 and Shailendra K. Gupta1,4 Downloaded from 1Department of Systems Biology and Bioinformatics, University of Rostock, Rostock, Germany, 2Laboratory of Systems Tumor Immunology, Department of Dermatology, University Hospital Erlangen, Friedrich-Alexander-University Erlangen-Nuremberg, Germany, 3Stellenbosch Institute for Advanced Study (STIAS), Wallenberg Research Centre at Stellenbosch University, Stellenbosch, South Africa and 4Department of Bioinformatics, CSIR-Indian Institute of Toxicology Research, 226001 Lucknow, Uttar Pradesh, India http://nar.oxfordjournals.org/ Received December 22, 2013; Revised April 18, 2014; Accepted May 10, 2014 ABSTRACT fine-tuned through a cellular context-dependent regulation by multiple miRNAs, where miRNAs can either induce MicroRNAs (miRNAs) are an integral part of gene reg- translational repression or target mRNA degradation (4). ulation at the post-transcriptional level. Recently, it Thereby, the miRNA-target regulation machinery can real- at Universitaet Erlangen-Nuernberg, Wirtschafts- und Sozialwissenschaftliche Z on August 3, 2016 has been shown that pairs of miRNAs can repress the ize elaborate gene control functions, including noise buffer- translation of a target mRNA in a cooperative man- ing or homeostasis, and can ultimately mediate distinct tar- ner, which leads to an enhanced effectiveness and get expression patterns appropriate to the demand of differ- specificity in target repression. However, it remains ent biological processes (3,5,6). However, deregulated miR- unclear which miRNA pairs can synergize and which NAs have also been associated with the pathogenesis and genes are target of cooperative miRNA regulation. -

Leveraging Mouse Chromatin Data for Heritability Enrichment Informs Common Disease Architecture and Reveals Cortical Layer Contributions to Schizophrenia
Downloaded from genome.cshlp.org on October 5, 2021 - Published by Cold Spring Harbor Laboratory Press Research Leveraging mouse chromatin data for heritability enrichment informs common disease architecture and reveals cortical layer contributions to schizophrenia Paul W. Hook1 and Andrew S. McCallion1,2,3 1McKusick-Nathans Department of Genetic Medicine, 2Department of Comparative and Molecular Pathobiology, 3Department of Medicine, Johns Hopkins University School of Medicine, Baltimore, Maryland 21205, USA Genome-wide association studies have implicated thousands of noncoding variants across common human phenotypes. However, they cannot directly inform the cellular context in which disease-associated variants act. Here, we use open chro- matin profiles from discrete mouse cell populations to address this challenge. We applied stratified linkage disequilibrium score regression and evaluated heritability enrichment in 64 genome-wide association studies, emphasizing schizophrenia. We provide evidence that mouse-derived human open chromatin profiles can serve as powerful proxies for difficult to ob- tain human cell populations, facilitating the illumination of common disease heritability enrichment across an array of hu- man phenotypes. We demonstrate that signatures from discrete subpopulations of cortical excitatory and inhibitory neurons are significantly enriched for schizophrenia heritability with maximal enrichment in cortical layer V excitatory neu- rons. We also show that differences between schizophrenia and bipolar disorder are concentrated in excitatory neurons in cortical layers II-III, IV, and V, as well as the dentate gyrus. Finally, we leverage these data to fine-map variants in 177 schiz- ophrenia loci nominating variants in 104/177. We integrate these data with transcription factor binding site, chromatin in- teraction, and validated enhancer data, placing variants in the cellular context where they may modulate risk. -

047605V1.Full.Pdf
bioRxiv preprint doi: https://doi.org/10.1101/047605; this version posted April 8, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Assembly and Activation of Dynein-Dynactin by the Cargo Adaptor Protein Hook3 Courtney M. Schroeder1,2 and Ronald D. Vale1,2 1The Howard Hughes Medical Institute, University of California, San Francisco, San Francisco, California, USA 2Department of Cellular and Molecular Pharmacology, University of California, San Francisco, San Francisco, California, USA. Corresponding Author: Ronald D. Vale Dept. of Cellular and Molecular Pharmacology University of California, San Francisco Genentech Hall, MC 2200, Room N312A 600-16th Street San Francisco, CA 94158-2517 E-mail: [email protected] Phone: 415-476-6380 Fax: 415-514-4412 bioRxiv preprint doi: https://doi.org/10.1101/047605; this version posted April 8, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 2 Abstract Metazoan cytoplasmic dynein moves processively along microtubules with the aid of dynactin and an adaptor protein that joins dynein and dynactin into a stable ternary complex. Here, we have examined how Hook3, a cargo adaptor involved in Golgi and endosome transport, forms a motile dynein-dynactin complex. We show that the conserved Hook domain interacts directly with the dynein light intermediate chain 1 (LIC1). -

(MMP-9) in Oral Cancer Squamous Cells—Are There Therapeutical Hopes?
materials Article Lutein Treatment Effects on the Redox Status and Metalloproteinase-9 (MMP-9) in Oral Cancer Squamous Cells—Are There Therapeutical Hopes? 1 2,3 4 1 Dan Alexandru Enăs, escu , Mihaela Georgeta Moisescu , Marina Imre , Maria Greabu , Alexandra Ripszky Totan 1,* , Iulia Stanescu-Spinu 1, Marian Burcea 5, Crenguta Albu 6,* and Daniela Miricescu 1 1 Department of Biochemistry, Faculty of Dental Medicine, University of Medicine and Pharmacy Carol Davila, 8 Eroii Sanitari Blvd., Sector 5, 050474 Bucharest, Romania; [email protected] (D.A.E.); [email protected] (M.G.); [email protected] (I.S.-S.); [email protected] (D.M.) 2 Department Biophysics and Cellular Biotechnology, University of Medicine and Pharmacy Carol Davila, 8 Eroii Sanitari Blvd., Sector 5, 050474 Bucharest, Romania; [email protected] 3 Excellence Centre for Research in Biophysics and Cellular Biotechnology, University of Medicine and Pharmacy Carol Davila, 8 Eroii Sanitari Blvd., Sector 5, 050474 Bucharest, Romania 4 Department of Complete Denture, Faculty of Dental Medicine, Carol Davila University of Medicine and Pharmacy, 8 Eroii Sanitari Blvd., Sector 5, 050474 Bucharest, Romania; [email protected] 5 Department of Ophthalmology, Faculty of General Medicine, University of Medicine and Pharmacy Carol Davila, 8 Eroilor Sanitari Blvd., 050474 Bucharest, Romania; [email protected] 6 Department of Genetics, Faculty of General Medicine, University of Medicine and Pharmacy Carol Davila, 8 Eroilor Sanitari Blvd., 050474 Bucharest, Romania * Correspondence: [email protected] (A.R.T.); [email protected] (C.A.) Citation: En˘as, escu, D.A.; Moisescu, M.G.; Imre, M.; Greabu, M.; Ripszky Totan, A.; Stanescu-Spinu, I.; Burcea, Abstract: Carotenoids loaded in nanoparticles should be regarded as a promising way to increase M.; Albu, C.; Miricescu, D. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

MBNL1 Regulates Essential Alternative RNA Splicing Patterns in MLL-Rearranged Leukemia
ARTICLE https://doi.org/10.1038/s41467-020-15733-8 OPEN MBNL1 regulates essential alternative RNA splicing patterns in MLL-rearranged leukemia Svetlana S. Itskovich1,9, Arun Gurunathan 2,9, Jason Clark 1, Matthew Burwinkel1, Mark Wunderlich3, Mikaela R. Berger4, Aishwarya Kulkarni5,6, Kashish Chetal6, Meenakshi Venkatasubramanian5,6, ✉ Nathan Salomonis 6,7, Ashish R. Kumar 1,7 & Lynn H. Lee 7,8 Despite growing awareness of the biologic features underlying MLL-rearranged leukemia, 1234567890():,; targeted therapies for this leukemia have remained elusive and clinical outcomes remain dismal. MBNL1, a protein involved in alternative splicing, is consistently overexpressed in MLL-rearranged leukemias. We found that MBNL1 loss significantly impairs propagation of murine and human MLL-rearranged leukemia in vitro and in vivo. Through transcriptomic profiling of our experimental systems, we show that in leukemic cells, MBNL1 regulates alternative splicing (predominantly intron exclusion) of several genes including those essential for MLL-rearranged leukemogenesis, such as DOT1L and SETD1A.Wefinally show that selective leukemic cell death is achievable with a small molecule inhibitor of MBNL1. These findings provide the basis for a new therapeutic target in MLL-rearranged leukemia and act as further validation of a burgeoning paradigm in targeted therapy, namely the disruption of cancer-specific splicing programs through the targeting of selectively essential RNA binding proteins. 1 Division of Bone Marrow Transplantation and Immune Deficiency, Cincinnati Children’s Hospital Medical Center, Cincinnati, OH 45229, USA. 2 Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital Medical Center, Cincinnati, OH 45229, USA. 3 Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital Medical Center, Cincinnati, OH 45229, USA. -
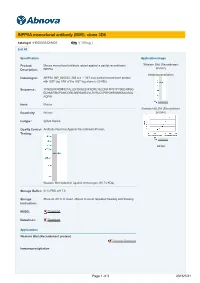
INPP5A Monoclonal Antibody (M05), Clone 3D8
INPP5A monoclonal antibody (M05), clone 3D8 Catalog # : H00003632-M05 規格 : [ 100 ug ] List All Specification Application Image Product Mouse monoclonal antibody raised against a partial recombinant Western Blot (Recombinant protein) Description: INPP5A. Immunoprecipitation Immunogen: INPP5A (NP_005530, 288 a.a. ~ 387 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. Sequence: YFNQEVFRDNNGTALLEFDKELSVFKDRLYELDISFPPSYPYSEDARQG EQYMNTRCPAWCDRILMSPSAKELVLRVSVCCPSPGHRGMWSAGSGL AQPW enlarge Host: Mouse Sandwich ELISA (Recombinant Reactivity: Human protein) Isotype: IgG2a Kappa Quality Control Antibody Reactive Against Recombinant Protein. Testing: enlarge ELISA Western Blot detection against Immunogen (36.74 KDa) . Storage Buffer: In 1x PBS, pH 7.4 Storage Store at -20°C or lower. Aliquot to avoid repeated freezing and thawing. Instruction: MSDS: Download Datasheet: Download Applications Western Blot (Recombinant protein) Protocol Download Immunoprecipitation Page 1 of 3 2016/5/21 Immunoprecipitation of INPP5A transfected lysate using anti-INPP5A monoclonal antibody and Protein A Magnetic Bead (U0007), and immunoblotted with INPP5A MaxPab rabbit polyclonal antibody. Protocol Download Sandwich ELISA (Recombinant protein) Detection limit for recombinant GST tagged INPP5A is 0.1 ng/ml as a capture antibody. Protocol Download ELISA Gene Information Entrez GeneID: 3632 GeneBank NM_005539 Accession#: Protein NP_005530 Accession#: Gene Name: INPP5A Gene Alias: 5PTASE,DKFZp434A1721,MGC116947,MGC116949 Gene inositol polyphosphate-5-phosphatase, 40kDa Description: Omim ID: 600106 Gene Ontology: Hyperlink Gene Summary: The protein encoded by this gene is a membrane-associated type I inositol 1,4,5-trisphosphate (InsP3) 5-phosphatase. InsP3 5- phosphatases hydrolyze Ins(1,4,5)P3, which mobilizes intracellular calcium and acts as a second messenger mediating cell responses to various stimulation. -

Multivariate Meta-Analysis of Differential Principal Components Underlying Human Primed and Naive-Like Pluripotent States
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.20.347666; this version posted October 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. This article is a US Government work. It is not subject to copyright under 17 USC 105 and is also made available for use under a CC0 license. October 20, 2020 To: bioRxiv Multivariate Meta-Analysis of Differential Principal Components underlying Human Primed and Naive-like Pluripotent States Kory R. Johnson1*, Barbara S. Mallon2, Yang C. Fann1, and Kevin G. Chen2*, 1Intramural IT and Bioinformatics Program, 2NIH Stem Cell Unit, National Institute of Neurological Disorders and Stroke, National Institutes of Health, Bethesda, Maryland 20892, USA Keywords: human pluripotent stem cells; naive pluripotency, meta-analysis, principal component analysis, t-SNE, consensus clustering *Correspondence to: Dr. Kory R. Johnson ([email protected]) Dr. Kevin G. Chen ([email protected]) 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.20.347666; this version posted October 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. This article is a US Government work. It is not subject to copyright under 17 USC 105 and is also made available for use under a CC0 license. ABSTRACT The ground or naive pluripotent state of human pluripotent stem cells (hPSCs), which was initially established in mouse embryonic stem cells (mESCs), is an emerging and tentative concept. To verify this important concept in hPSCs, we performed a multivariate meta-analysis of major hPSC datasets via the combined analytic powers of percentile normalization, principal component analysis (PCA), t-distributed stochastic neighbor embedding (t-SNE), and SC3 consensus clustering. -

The Role of Cdep in the Embryonic Morphogenesis of Drosophila Melanogaster
The Role of Cdep in the Embryonic Morphogenesis of Drosophila melanogaster Dissertation zur Erlangung des akademischen Grades Doctor rerum naturalium (Dr. rer.nat.) vorgelegt der Fakultät Mathematik und Naturwissenschaften der Technischen Universität Dresden von Anne Morbach Geboren am 05. August 1984 in Nordhausen, Deutschland Eingereicht am 30. Oktober 2015 Die Dissertation wurde in der Zeit von April 2012 bis Oktober 2015 im Max-Planck-Institut für molekulare Zellbiologie und Genetik angefertigt Gutachter: Prof. Dr. Elisabeth Knust Prof. Dr. Christian Dahmann Now, here, you see, it takes all the running you can do, to keep in the same place. Lewis Carroll, Through the Looking-Glass I Contents List of Figures V List of Tables VII Abstract IX List of Abbreviations XI 1 Introduction 1 1.1 Epithelial cell polarity . 1 1.1.1 Cellularization and formation of the primary epithelium . 1 1.1.1.1 Establishment of epithelial polarity and adhesion . 2 1.1.2 The epithelial polarity network . 3 1.1.3 Cell-cell adhesion . 5 1.1.3.1 Adherens junctions . 5 1.1.3.2 Septate junctions . 6 1.2 Epithelial movements in Drosophila embryonic morphogenesis . 6 1.2.1 Epithelial tube formation during Drosophila embryogenesis . 7 1.2.2 Coordinated migration of epithelial sheets during Dros. embryogenesis 7 1.2.2.1 FERM domain proteins in epithelial migration . 9 1.2.2.2 Cdep . 10 1.3 Mutagenesis with the CRISPR/Cas9 system . 12 2 Aim of My PhD Thesis Work 15 3 Preliminary Work 17 4 Results 19 4.1 A screen for novel regulators in Drosophila embryonic morphogenesis .