Supplementary Information
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Targeted Genes and Methodology Details for Neuromuscular Genetic Panels
Targeted Genes and Methodology Details for Neuromuscular Genetic Panels Reference transcripts based on build GRCh37 (hg19) interrogated by Neuromuscular Genetic Panels Next-generation sequencing (NGS) and/or Sanger sequencing is performed Motor Neuron Disease Panel to test for the presence of a mutation in these genes. Gene GenBank Accession Number Regions of homology, high GC-rich content, and repetitive sequences may ALS2 NM_020919 not provide accurate sequence. Therefore, all reported alterations detected ANG NM_001145 by NGS are confirmed by an independent reference method based on laboratory developed criteria. However, this does not rule out the possibility CHMP2B NM_014043 of a false-negative result in these regions. ERBB4 NM_005235 Sanger sequencing is used to confirm alterations detected by NGS when FIG4 NM_014845 appropriate.(Unpublished Mayo method) FUS NM_004960 HNRNPA1 NM_031157 OPTN NM_021980 PFN1 NM_005022 SETX NM_015046 SIGMAR1 NM_005866 SOD1 NM_000454 SQSTM1 NM_003900 TARDBP NM_007375 UBQLN2 NM_013444 VAPB NM_004738 VCP NM_007126 ©2018 Mayo Foundation for Medical Education and Research Page 1 of 14 MC4091-83rev1018 Muscular Dystrophy Panel Muscular Dystrophy Panel Gene GenBank Accession Number Gene GenBank Accession Number ACTA1 NM_001100 LMNA NM_170707 ANO5 NM_213599 LPIN1 NM_145693 B3GALNT2 NM_152490 MATR3 NM_199189 B4GAT1 NM_006876 MYH2 NM_017534 BAG3 NM_004281 MYH7 NM_000257 BIN1 NM_139343 MYOT NM_006790 BVES NM_007073 NEB NM_004543 CAPN3 NM_000070 PLEC NM_000445 CAV3 NM_033337 POMGNT1 NM_017739 CAVIN1 NM_012232 POMGNT2 -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Conserved and Novel Properties of Clathrin-Mediated Endocytosis in Dictyostelium Discoideum" (2012)
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2012 Conserved and Novel Properties of Clathrin- Mediated Endocytosis in Dictyostelium Discoideum Laura Macro Follow this and additional works at: http://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons Recommended Citation Macro, Laura, "Conserved and Novel Properties of Clathrin-Mediated Endocytosis in Dictyostelium Discoideum" (2012). Student Theses and Dissertations. Paper 163. This Thesis is brought to you for free and open access by Digital Commons @ RU. It has been accepted for inclusion in Student Theses and Dissertations by an authorized administrator of Digital Commons @ RU. For more information, please contact [email protected]. CONSERVED AND NOVEL PROPERTIES OF CLATHRIN- MEDIATED ENDOCYTOSIS IN DICTYOSTELIUM DISCOIDEUM A Thesis Presented to the Faculty of The Rockefeller University in Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Laura Macro June 2012 © Copyright by Laura Macro 2012 CONSERVED AND NOVEL PROPERTIES OF CLATHRIN- MEDIATED ENDOCYTOSIS IN DICTYOSTELIUM DISCOIDEUM Laura Macro, Ph.D. The Rockefeller University 2012 The protein clathrin mediates one of the major pathways of endocytosis from the extracellular milieu and plasma membrane. Clathrin functions with a network of interacting accessory proteins, one of which is the adaptor complex AP-2, to co-ordinate vesicle formation. Disruption of genes involved in clathrin-mediated endocytosis causes embryonic lethality in multicellular animals suggesting that clathrin-mediated endocytosis is a fundamental cellular process. However, loss of clathrin-mediated endocytosis genes in single cell eukaryotes, such as S.cerevisiae (yeast), does not cause lethality, suggesting that clathrin may convey specific advantages for multicellularity. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

AP4S1 Splice-Site Mutation in a Case of Spastic Paraplegia Type 52 with Polymicrogyria
CLINICAL/SCIENTIFIC NOTES OPEN ACCESS AP4S1 splice-site mutation in a case of spastic paraplegia type 52 with polymicrogyria Susana Carmona, PhD, Clara Marecos, MD, Marta Amorim, MD, Ana C. Ferreira, MD, Carla Conceição, MD, Correspondence Jos´eBr´as, PhD, Sofia T. Duarte, MD, PhD,* and Rita Guerreiro, PhD* Dr. Guerreiro [email protected] Neurol Genet 2018;4:e273. doi:10.1212/NXG.0000000000000273 Hereditary spastic paraplegias (HSPs) are a group of rare inherited neurodegenerative dis- orders that result from primary retrograde dysfunction of the long descending fibers of the corticospinal tract, causing lower limb spasticity and muscular weakness. This group of diseases has a heterogeneous clinical presentation. An extensive list of associated genes, different in- heritance patterns, and ages at onset have been reported in HSPs.1 Spastic paraplegia type 52 (SPG52) is an autosomal recessive disease caused by AP4S1 mutations. The disease is char- acterized by neonatal hypotonia that progresses to hypertonia and spasticity in early childhood, developmental delay, mental retardation, and poor or absent speech. Febrile or afebrile seizures – may also occur.2 4 Clinical case presentation We report the case of a Portuguese 2-year-old boy born to healthy nonconsanguineous parents after a full-term gestation with intrauterine growth restriction after week 37. During the first months of life, the patient presented poor weight gain, hyperammonemia with elevation of glutamine and ornithine, low citrulline, and negative orotic acid. Weight recovery and nor- malization of amino acid profile were observed after protein restriction and remained normal after reintroduction of normal diet. Genetic study of urea cycle disorders (NAGS, CPS, and OTC) was negative. -

Cldn19 Clic2 Clmp Cln3
NewbornDx™ Advanced Sequencing Evaluation When time to diagnosis matters, the NewbornDx™ Advanced Sequencing Evaluation from Athena Diagnostics delivers rapid, 5- to 7-day results on a targeted 1,722-genes. A2ML1 ALAD ATM CAV1 CLDN19 CTNS DOCK7 ETFB FOXC2 GLUL HOXC13 JAK3 AAAS ALAS2 ATP1A2 CBL CLIC2 CTRC DOCK8 ETFDH FOXE1 GLYCTK HOXD13 JUP AARS2 ALDH18A1 ATP1A3 CBS CLMP CTSA DOK7 ETHE1 FOXE3 GM2A HPD KANK1 AASS ALDH1A2 ATP2B3 CC2D2A CLN3 CTSD DOLK EVC FOXF1 GMPPA HPGD K ANSL1 ABAT ALDH3A2 ATP5A1 CCDC103 CLN5 CTSK DPAGT1 EVC2 FOXG1 GMPPB HPRT1 KAT6B ABCA12 ALDH4A1 ATP5E CCDC114 CLN6 CUBN DPM1 EXOC4 FOXH1 GNA11 HPSE2 KCNA2 ABCA3 ALDH5A1 ATP6AP2 CCDC151 CLN8 CUL4B DPM2 EXOSC3 FOXI1 GNAI3 HRAS KCNB1 ABCA4 ALDH7A1 ATP6V0A2 CCDC22 CLP1 CUL7 DPM3 EXPH5 FOXL2 GNAO1 HSD17B10 KCND2 ABCB11 ALDOA ATP6V1B1 CCDC39 CLPB CXCR4 DPP6 EYA1 FOXP1 GNAS HSD17B4 KCNE1 ABCB4 ALDOB ATP7A CCDC40 CLPP CYB5R3 DPYD EZH2 FOXP2 GNE HSD3B2 KCNE2 ABCB6 ALG1 ATP8A2 CCDC65 CNNM2 CYC1 DPYS F10 FOXP3 GNMT HSD3B7 KCNH2 ABCB7 ALG11 ATP8B1 CCDC78 CNTN1 CYP11B1 DRC1 F11 FOXRED1 GNPAT HSPD1 KCNH5 ABCC2 ALG12 ATPAF2 CCDC8 CNTNAP1 CYP11B2 DSC2 F13A1 FRAS1 GNPTAB HSPG2 KCNJ10 ABCC8 ALG13 ATR CCDC88C CNTNAP2 CYP17A1 DSG1 F13B FREM1 GNPTG HUWE1 KCNJ11 ABCC9 ALG14 ATRX CCND2 COA5 CYP1B1 DSP F2 FREM2 GNS HYDIN KCNJ13 ABCD3 ALG2 AUH CCNO COG1 CYP24A1 DST F5 FRMD7 GORAB HYLS1 KCNJ2 ABCD4 ALG3 B3GALNT2 CCS COG4 CYP26C1 DSTYK F7 FTCD GP1BA IBA57 KCNJ5 ABHD5 ALG6 B3GAT3 CCT5 COG5 CYP27A1 DTNA F8 FTO GP1BB ICK KCNJ8 ACAD8 ALG8 B3GLCT CD151 COG6 CYP27B1 DUOX2 F9 FUCA1 GP6 ICOS KCNK3 ACAD9 ALG9 -

Hereditary Spastic Paraplegia: from Genes, Cells and Networks to Novel Pathways for Drug Discovery
brain sciences Review Hereditary Spastic Paraplegia: From Genes, Cells and Networks to Novel Pathways for Drug Discovery Alan Mackay-Sim Griffith Institute for Drug Discovery, Griffith University, Brisbane, QLD 4111, Australia; a.mackay-sim@griffith.edu.au Abstract: Hereditary spastic paraplegia (HSP) is a diverse group of Mendelian genetic disorders affect- ing the upper motor neurons, specifically degeneration of their distal axons in the corticospinal tract. Currently, there are 80 genes or genomic loci (genomic regions for which the causative gene has not been identified) associated with HSP diagnosis. HSP is therefore genetically very heterogeneous. Finding treatments for the HSPs is a daunting task: a rare disease made rarer by so many causative genes and many potential mutations in those genes in individual patients. Personalized medicine through genetic correction may be possible, but impractical as a generalized treatment strategy. The ideal treatments would be small molecules that are effective for people with different causative mutations. This requires identification of disease-associated cell dysfunctions shared across geno- types despite the large number of HSP genes that suggest a wide diversity of molecular and cellular mechanisms. This review highlights the shared dysfunctional phenotypes in patient-derived cells from patients with different causative mutations and uses bioinformatic analyses of the HSP genes to identify novel cell functions as potential targets for future drug treatments for multiple genotypes. Keywords: neurodegeneration; motor neuron disease; spastic paraplegia; endoplasmic reticulum; Citation: Mackay-Sim, A. Hereditary protein-protein interaction network Spastic Paraplegia: From Genes, Cells and Networks to Novel Pathways for Drug Discovery. Brain Sci. 2021, 11, 403. -

CENTOGENE's Severe and Early Onset Disorder Gene List
CENTOGENE’s severe and early onset disorder gene list USED IN PRENATAL WES ANALYSIS AND IDENTIFICATION OF “PATHOGENIC” AND “LIKELY PATHOGENIC” CENTOMD® VARIANTS IN NGS PRODUCTS The following gene list shows all genes assessed in prenatal WES tests or analysed for P/LP CentoMD® variants in NGS products after April 1st, 2020. For searching a single gene coverage, just use the search on www.centoportal.com AAAS, AARS1, AARS2, ABAT, ABCA12, ABCA3, ABCB11, ABCB4, ABCB7, ABCC6, ABCC8, ABCC9, ABCD1, ABCD4, ABHD12, ABHD5, ACACA, ACAD9, ACADM, ACADS, ACADVL, ACAN, ACAT1, ACE, ACO2, ACOX1, ACP5, ACSL4, ACTA1, ACTA2, ACTB, ACTG1, ACTL6B, ACTN2, ACVR2B, ACVRL1, ACY1, ADA, ADAM17, ADAMTS2, ADAMTSL2, ADAR, ADARB1, ADAT3, ADCY5, ADGRG1, ADGRG6, ADGRV1, ADK, ADNP, ADPRHL2, ADSL, AFF2, AFG3L2, AGA, AGK, AGL, AGPAT2, AGPS, AGRN, AGT, AGTPBP1, AGTR1, AGXT, AHCY, AHDC1, AHI1, AIFM1, AIMP1, AIPL1, AIRE, AK2, AKR1D1, AKT1, AKT2, AKT3, ALAD, ALDH18A1, ALDH1A3, ALDH3A2, ALDH4A1, ALDH5A1, ALDH6A1, ALDH7A1, ALDOA, ALDOB, ALG1, ALG11, ALG12, ALG13, ALG14, ALG2, ALG3, ALG6, ALG8, ALG9, ALMS1, ALOX12B, ALPL, ALS2, ALX3, ALX4, AMACR, AMER1, AMN, AMPD1, AMPD2, AMT, ANK2, ANK3, ANKH, ANKRD11, ANKS6, ANO10, ANO5, ANOS1, ANTXR1, ANTXR2, AP1B1, AP1S1, AP1S2, AP3B1, AP3B2, AP4B1, AP4E1, AP4M1, AP4S1, APC2, APTX, AR, ARCN1, ARFGEF2, ARG1, ARHGAP31, ARHGDIA, ARHGEF9, ARID1A, ARID1B, ARID2, ARL13B, ARL3, ARL6, ARL6IP1, ARMC4, ARMC9, ARSA, ARSB, ARSL, ARV1, ARX, ASAH1, ASCC1, ASH1L, ASL, ASNS, ASPA, ASPH, ASPM, ASS1, ASXL1, ASXL2, ASXL3, ATAD3A, ATCAY, ATIC, ATL1, ATM, ATOH7, -

AP-4 Vesicles Unmasked by Organellar Proteomics to Reveal Their Cargo
bioRxiv preprint doi: https://doi.org/10.1101/235226; this version posted December 18, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. AP-4 vesicles unmasked by organellar proteomics to reveal their cargo and machinery Alexandra K. Davies1, Daniel N. Itzhak2, James R. Edgar1, Tara L. Archuleta3,4, Jennifer Hirst1, Lauren P. Jackson3,4, Margaret S. Robinson1*, Georg H. H. Borner2* 1Cambridge Institute for Medical Research, University of Cambridge, Cambridge CB2 0XY, UK 2Department of Proteomics and Signal Transduction, Max Planck Institute of Biochemistry, Martinsried 82152, Germany 3Department of Biological Sciences, Vanderbilt University, Nashville, TN 37235, USA 4Center for Structural Biology, Vanderbilt University, Nashville, TN 37235, USA *Correspondence: G.H.H.B., [email protected]; M.S.R., [email protected] 1 Davies et al., 2017 bioRxiv preprint doi: https://doi.org/10.1101/235226; this version posted December 18, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Adaptor protein 4 (AP-4) is an ancient membrane trafficking complex, whose function has largely remained elusive. In humans, AP-4 deficiency causes a severe neurological disorder of unknown aetiology. We apply multiple unbiased proteomic methods, including ‘Dynamic Organellar Maps’, to find proteins whose subcellular localisation depends on AP-4. We identify three highly conserved transmembrane cargo proteins, ATG9A, SERINC1 and SERINC3, and two AP-4 accessory proteins, RUSC1 and RUSC2. We demonstrate that AP-4 deficiency causes missorting of ATG9A in diverse cell types, including neuroblastoma and AP-4 patient-derived cells, as well as dysregulation of autophagy. -
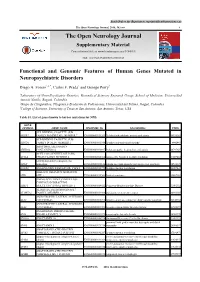
The Open Neurology Journal, 2016, 10, I-Xv I the Open Neurology Journal Supplementary Material Content List Available At
Send Orders for Reprints to [email protected] The Open Neurology Journal, 2016, 10, i-xv i The Open Neurology Journal Supplementary Material Content list available at: www.benthamopen.com/TONEUJ/ DOI: 10.2174/1874205X01610010143 Functional and Genomic Features of Human Genes Mutated in Neuropsychiatric Disorders Diego A. Forero1,4, *, Carlos F. Prada2 and George Perry3 1Laboratory of NeuroPsychiatric Genetics, Biomedical Sciences Research Group, School of Medicine, Universidad Antonio Nariño. Bogotá, Colombia 2Grupo de Citogenética, Filogenia y Evolución de Poblaciones, Universidad del Tolima. Ibagué, Colombia 3College of Sciences, University of Texas at San Antonio, San Antonio, Texas, USA Table S1. List of genes known to harbor mutations for NPD. GENE SYMBOL GENE NAME ENSEMBL ID DISORDERS PMID ATP-BINDING CASSETTE, SUB- ABCB7 FAMILY B (MDR/TAP), MEMBER 7 ENSG00000131269 X-linked sideroblastic anemia and ataxia 10196363 ATP-BINDING CASSETTE, SUB- ABCD1 FAMILY D (ALD), MEMBER 1 ENSG00000101986 X-linked adrenoleukodystrophy 8441467 ABHYDROLASE DOMAIN ABHD12 CONTAINING 12 ENSG00000100997 Polyneuropathy, hearing loss and ataxia 20797687 ACYL-COA SYNTHETASE LONG- ACSL4 CHAIN FAMILY MEMBER 4 ENSG00000068366 nonspecific X-linked mental retardation 11889465 activity-dependent neuroprotector ADNP homeobox ENSG00000101126 autism spectrum disorder and intellectual disability 24531329 AGTR2 ANGIOTENSIN II RECEPTOR, TYPE 2 ENSG00000180772 X-linked mental retardation 12089445 ABELSON HELPER INTEGRATION AHI1 SITE 1 ENSG00000135541 Joubert -

Membrane Trafficking in Health and Disease Rebecca Yarwood*, John Hellicar*, Philip G
© 2020. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2020) 13, dmm043448. doi:10.1242/dmm.043448 AT A GLANCE Membrane trafficking in health and disease Rebecca Yarwood*, John Hellicar*, Philip G. Woodman‡ and Martin Lowe‡ ABSTRACT KEY WORDS: Disease, Endocytic pathway, Genetic disorder, Membrane traffic, Secretory pathway, Vesicle Membrane trafficking pathways are essential for the viability and growth of cells, and play a major role in the interaction of cells with Introduction their environment. In this At a Glance article and accompanying Membrane trafficking pathways are essential for cells to maintain poster, we outline the major cellular trafficking pathways and discuss critical functions, to grow, and to accommodate to their chemical how defects in the function of the molecular machinery that mediates and physical environment. Membrane flux through these pathways this transport lead to various diseases in humans. We also briefly is high, and in specialised cells in some tissues can be enormous. discuss possible therapeutic approaches that may be used in the For example, pancreatic acinar cells synthesise and secrete amylase, future treatment of trafficking-based disorders. one of the many enzymes they produce, at a rate of approximately 0.5% of cellular protein mass per hour (Allfrey et al., 1953), while in Schwann cells, the rate of membrane protein export must correlate School of Biological Sciences, Faculty of Biology, Medicine and Health, with the several thousand-fold expansion of the cell surface that University of Manchester, Manchester, M13 9PT, UK. occurs during myelination (Pereira et al., 2012). The population of *These authors contributed equally to this work cell surface proteins is constantly monitored and modified via the ‡Authors for correspondence ([email protected]; endocytic pathway. -

New Clinical Insights and Physical Mapping of FOXG1-Regulatory Elements
European Journal of Human Genetics (2012) 20, 1216–1223 & 2012 Macmillan Publishers Limited All rights reserved 1018-4813/12 www.nature.com/ejhg ARTICLE 14q12 and severe Rett-like phenotypes: new clinical insights and physical mapping of FOXG1-regulatory elements Lila Allou1,15, Laetitia Lambert1,2,3,15, Daniel Amsallem4, Eric Bieth5, Patrick Edery6,7, Anne Destre´e8, Franc¸ois Rivier9,DavidAmor10, Elizabeth Thompson11, Julian Nicholl12, Michael Harbord13,ChristopheNemos1, Aline Saunier14, Aissa Moustaı¨ne14, Adeline Vigouroux5, Philippe Jonveaux1,14 and Christophe Philippe*,1,14 The Forkhead box G1 (FOXG1) gene has been implicated in severe Rett-like phenotypes. It encodes the Forkhead box protein G1, a winged-helix transcriptional repressor critical for forebrain development. Recently, the core FOXG1 syndrome was defined as postnatal microcephaly, severe mental retardation, absent language, dyskinesia, and dysgenesis of the corpus callosum. We present seven additional patients with a severe Rett-like neurodevelopment disorder associated with de novo FOXG1 point mutations (two cases) or 14q12 deletions (five cases). We expand the mutational spectrum in patients with FOXG1-related encephalopathies and precise the core FOXG1 syndrome phenotype. Dysgenesis of the corpus callosum and dyskinesia are not always present in FOXG1-mutated patients. We believe that the FOXG1 gene should be considered in severely mentally retarded patients (no speech-language) with severe acquired microcephaly ( À4toÀ6 SD) and few clinical features suggestive of Rett syndrome. Interestingly enough, three 14q12 deletions that do not include the FOXG1 gene are associated with phenotypes very reminiscent to that of FOXG1-mutation-positive patients. We physically mapped a putative long-range FOXG1-regulatory element in a 0.43 Mb DNA segment encompassing the PRKD1 locus.