Supplementary Figure S1: Protein Abundance Ratios for Selected Proteins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Airway Inflammation Mitochondrial Dysfunction Increases Allergic
Mitochondrial Dysfunction Increases Allergic Airway Inflammation Leopoldo Aguilera-Aguirre, Attila Bacsi, Alfredo Saavedra-Molina, Alexander Kurosky, Sanjiv Sur and Istvan This information is current as Boldogh of October 1, 2021. J Immunol 2009; 183:5379-5387; Prepublished online 28 September 2009; doi: 10.4049/jimmunol.0900228 http://www.jimmunol.org/content/183/8/5379 Downloaded from References This article cites 61 articles, 11 of which you can access for free at: http://www.jimmunol.org/content/183/8/5379.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on October 1, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2009 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Mitochondrial Dysfunction Increases Allergic Airway Inflammation1 Leopoldo Aguilera-Aguirre,*§ Attila Bacsi,*¶ Alfredo Saavedra-Molina,§ Alexander Kurosky,† Sanjiv Sur,‡ and Istvan Boldogh*2 The prevalence of allergies and asthma among the world’s population has been steadily increasing due to environmental factors. -

Snps) Distant from Xenobiotic Response Elements Can Modulate Aryl Hydrocarbon Receptor Function: SNP-Dependent CYP1A1 Induction S
Supplemental material to this article can be found at: http://dmd.aspetjournals.org/content/suppl/2018/07/06/dmd.118.082164.DC1 1521-009X/46/9/1372–1381$35.00 https://doi.org/10.1124/dmd.118.082164 DRUG METABOLISM AND DISPOSITION Drug Metab Dispos 46:1372–1381, September 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Single Nucleotide Polymorphisms (SNPs) Distant from Xenobiotic Response Elements Can Modulate Aryl Hydrocarbon Receptor Function: SNP-Dependent CYP1A1 Induction s Duan Liu, Sisi Qin, Balmiki Ray,1 Krishna R. Kalari, Liewei Wang, and Richard M. Weinshilboum Division of Clinical Pharmacology, Department of Molecular Pharmacology and Experimental Therapeutics (D.L., S.Q., B.R., L.W., R.M.W.) and Division of Biomedical Statistics and Informatics, Department of Health Sciences Research (K.R.K.), Mayo Clinic, Rochester, Minnesota Received April 22, 2018; accepted June 28, 2018 ABSTRACT Downloaded from CYP1A1 expression can be upregulated by the ligand-activated aryl fashion. LCLs with the AA genotype displayed significantly higher hydrocarbon receptor (AHR). Based on prior observations with AHR-XRE binding and CYP1A1 mRNA expression after 3MC estrogen receptors and estrogen response elements, we tested treatment than did those with the GG genotype. Electrophoretic the hypothesis that single-nucleotide polymorphisms (SNPs) map- mobility shift assay (EMSA) showed that oligonucleotides with the ping hundreds of base pairs (bp) from xenobiotic response elements AA genotype displayed higher LCL nuclear extract binding after (XREs) might influence AHR binding and subsequent gene expres- 3MC treatment than did those with the GG genotype, and mass dmd.aspetjournals.org sion. -

Low Abundance of the Matrix Arm of Complex I in Mitochondria Predicts Longevity in Mice
ARTICLE Received 24 Jan 2014 | Accepted 9 Apr 2014 | Published 12 May 2014 DOI: 10.1038/ncomms4837 OPEN Low abundance of the matrix arm of complex I in mitochondria predicts longevity in mice Satomi Miwa1, Howsun Jow2, Karen Baty3, Amy Johnson1, Rafal Czapiewski1, Gabriele Saretzki1, Achim Treumann3 & Thomas von Zglinicki1 Mitochondrial function is an important determinant of the ageing process; however, the mitochondrial properties that enable longevity are not well understood. Here we show that optimal assembly of mitochondrial complex I predicts longevity in mice. Using an unbiased high-coverage high-confidence approach, we demonstrate that electron transport chain proteins, especially the matrix arm subunits of complex I, are decreased in young long-living mice, which is associated with improved complex I assembly, higher complex I-linked state 3 oxygen consumption rates and decreased superoxide production, whereas the opposite is seen in old mice. Disruption of complex I assembly reduces oxidative metabolism with concomitant increase in mitochondrial superoxide production. This is rescued by knockdown of the mitochondrial chaperone, prohibitin. Disrupted complex I assembly causes premature senescence in primary cells. We propose that lower abundance of free catalytic complex I components supports complex I assembly, efficacy of substrate utilization and minimal ROS production, enabling enhanced longevity. 1 Institute for Ageing and Health, Newcastle University, Newcastle upon Tyne NE4 5PL, UK. 2 Centre for Integrated Systems Biology of Ageing and Nutrition, Newcastle University, Newcastle upon Tyne NE4 5PL, UK. 3 Newcastle University Protein and Proteome Analysis, Devonshire Building, Devonshire Terrace, Newcastle upon Tyne NE1 7RU, UK. Correspondence and requests for materials should be addressed to T.v.Z. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

G Protein-Coupled Receptors
G PROTEIN-COUPLED RECEPTORS Overview:- The completion of the Human Genome Project allowed the identification of a large family of proteins with a common motif of seven groups of 20-24 hydrophobic amino acids arranged as α-helices. Approximately 800 of these seven transmembrane (7TM) receptors have been identified of which over 300 are non-olfactory receptors (see Frederikson et al., 2003; Lagerstrom and Schioth, 2008). Subdivision on the basis of sequence homology allows the definition of rhodopsin, secretin, adhesion, glutamate and Frizzled receptor families. NC-IUPHAR recognizes Classes A, B, and C, which equate to the rhodopsin, secretin, and glutamate receptor families. The nomenclature of 7TM receptors is commonly used interchangeably with G protein-coupled receptors (GPCR), although the former nomenclature recognises signalling of 7TM receptors through pathways not involving G proteins. For example, adiponectin and membrane progestin receptors have some sequence homology to 7TM receptors but signal independently of G-proteins and appear to reside in membranes in an inverted fashion compared to conventional GPCR. Additionally, the NPR-C natriuretic peptide receptor has a single transmembrane domain structure, but appears to couple to G proteins to generate cellular responses. The 300+ non-olfactory GPCR are the targets for the majority of drugs in clinical usage (Overington et al., 2006), although only a minority of these receptors are exploited therapeutically. Signalling through GPCR is enacted by the activation of heterotrimeric GTP-binding proteins (G proteins), made up of α, β and γ subunits, where the α and βγ subunits are responsible for signalling. The α subunit (tabulated below) allows definition of one series of signalling cascades and allows grouping of GPCRs to suggest common cellular, tissue and behavioural responses. -

Supplement for TWO-SIGMA-G S1 Additional Type-I Error and Power Results
Supplement for TWO-SIGMA-G Eric Van Buren, Ming Hu, Liang Chen, John Wrobel, Kirk Wilhelmsen, Lishan Su, Yun Li, Di Wu S1 Additional Type-I Error and Power Results Independent Genes (No IGC) Genes Simulated with IGC (A) No Gene-Level Random Effects (B) No Gene-Level Random Effects Test Size = 30, Ref. Size = 30 Test Size = 30, Ref. Size = 30 0.03 0.03 0.02 0.02 0.01 0.01 Type-I Error Type-I Error Observed Set-Level Observed Set-Level 0.00 0.00 0.0000 0.0025 0.0050 0.0075 0.0100 0.0000 0.0025 0.0050 0.0075 0.0100 Significance Threshold Significance Threshold CAMERA MAST TWO-SIGMA-G CAMERA MAST TWO-SIGMA-G Genes Simulated with IGC Genes Simulated with IGC (C) Gene-Level Random Effects Present (D) Gene-Level Random Effects Test Size = 30, Ref. Size = 30 Incorrectly Absent 0.03 Test Size = 30, Ref. Size = 30 0.03 0.02 0.02 0.01 0.01 Type-I Error Type-I Error Observed Set-Level 0.00 Observed Set-Level 0.00 0.0000 0.0025 0.0050 0.0075 0.0100 0.0000 0.0025 0.0050 0.0075 0.0100 Significance Threshold Significance Threshold CAMERA MAST TWO-SIGMA-G CAMERA MAST TWO-SIGMA-G Supplementary Figure S1: Type-I error performance of CAMERA, MAST, and TWO-SIGMA-G as significance threshold varies from 0 to .01. Each panel varies the existence of IGC between genes in the test set and the presence of gene-level random effect terms in gene-level model (CAMERA never includes gene-level random effect terms). -

Anti-GNB4 (GW21993)
3050 Spruce Street, Saint Louis, MO 63103 USA Tel: (800) 521-8956 (314) 771-5765 Fax: (800) 325-5052 (314) 771-5757 email: [email protected] Product Information Anti-GNB4 antibody produced in chicken, affinity isolated antibody Catalog Number GW21993 Formerly listed as GenWay Catalog Number 15-288-21993, Guanine nucleotide-binding protein subunit beta 4 Anti- body. The product is a clear, colorless solution in phosphate – Storage Temperature Store at 20 °C buffered saline, pH 7.2, containing 0.02% sodium azide. Synonyms: Guanine nucleotide-binding protein, beta-4 Species Reactivity: Human, mouse subunit, Transducin beta chain 4 Tested Applications: WB Product Description Recommended Dilutions: Recommended starting dilution Guanine nucleotide-binding proteins (G proteins) are for Western blot analysis is 1:500, for tissue or cell staining involved as a modulator or transducer in various trans- 1:200. membrane signaling systems. The beta and gamma chains are required for the GTPase activity. for replacement of GDP Note: Optimal concentrations and conditions for each by GTP. and for G protein-effector interaction. application should be determined by the user. NCBI Accession number: NP_067642.1 Precautions and Disclaimer Swiss Prot Accession number: Q9HAV0 This product is for R&D use only, not for drug, household, or other uses. Due to the sodium azide content a material Gene Information: Human .. GNB4 (59345) safety data sheet (MSDS) for this product has been sent to Immunogen: Recombinant protein Guanine nucleotide- the attention of the safety officer of your institution. Please binding protein, beta-4 subunit consult the Material Safety Data Sheet for information regarding hazards and safe handling practices. -

Inherited Variants in Mitochondrial Biogenesis Genes May Influence Epithelial Ovarian Cancer Risk Jennifer Permuth-Wey1,2, Y. An
Author Manuscript Published OnlineFirst on March 29, 2011; DOI: 10.1158/1055-9965.EPI-10-1224 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Inherited Variants in Mitochondrial Biogenesis Genes May Influence Epithelial Ovarian Cancer Risk Jennifer Permuth-Wey1,2, Y. Ann Chen3 ,Ya-Yu Tsai1, Zhihua Chen4, Xiaotao Qu4, Johnathan M. Lancaster5, Heather Stockwell2, Getachew Dagne2, Edwin Iversen6, Harvey Risch7, Jill Barnholtz-Sloan8, Julie M. Cunningham9, Robert A. Vierkant10, Brooke L. Fridley10, Rebecca Sutphen11, John McLaughlin12, Steven A. Narod13, Ellen L. Goode10, Joellen M. Schildkraut14, David Fenstermacher4, Catherine M. Phelan1, and Thomas A. Sellers1 1Department of Cancer Epidemiology, Moffitt Cancer Center, Tampa, FL, USA. 2 Department of Epidemiology and Biostatistics, College of Public Health, University of South Florida, Tampa, FL, USA. 3 Department of Biostatistics, Moffitt Cancer Center, Tampa, FL, USA. 4 Department of Biomedical Informatics, Moffitt Cancer Center, Tampa, FL, USA. 5 Department of Women’s Oncology, Moffitt Cancer Center, Tampa, FL, USA. 6 Department of Statistical Science, Duke University Medical Center, Durham, NC, USA. 7Department of Epidemiology and Public Health, Yale University School of Medicine, New Haven, CT, USA. 8Case Comprehensive Cancer Center, Case School of Medicine, Cleveland, OH, USA. 9 Department of Laboratory Medicine and Pathology, Mayo Clinic College of Medicine, Rochester, MN, USA. 10Department of Health Sciences Research, Mayo Clinic College of Medicine, Rochester, MN, USA. 11Pediatrics Epidemiology Center, College of Medicine, University of South Florida, Tampa, FL, USA. 12Samuel Lunenfeld Research Institute, Toronto, Ontario, Canada. 13Center for Research in Women’s Health, Toronto, ON, Canada. 14Department of Community and Family Medicine, Duke University Medical Center, Durham, NC, USA. -

UQCRFS1N Assembles Mitochondrial Respiratory Complex-III Into an Asymmetric 21-Subunit Dimer
Protein Cell 2018, 9(6):586–591 https://doi.org/10.1007/s13238-018-0515-x Protein & Cell LETTER TO THE EDITOR UQCRFS1N assembles mitochondrial respiratory complex-III into an asymmetric 21-subunit dimer Dear Editor, Although several structures have been solved with very high resolution, the full length N-terminal processed peptide (1– Mitochondrial respiratory chain consists of four multimeric 78 amino acids, UQCRFS1N) of the iron-sulfur Rieske pro- protein complexes, Complex I-IV (CI, NADH dehydroge- tein (UQCRFS1) subunit has not been assigned in all of nase; CII, succinate:ubiquinone oxidoreductase; CIII, cyto- these structures (Table 1). UQCRFS1N is the N-terminal chrome bc1 complex; and CIV, cytochrome c oxidase). Cell mitochondrial targeting sequence of UQCRFS1, and after its These four complexes transfer electrons from NADH or cleavage from the precursor, this small peptide remains & FADH to oxygen and pump protons from mitochondrial 2 bound to CIII with unknown functions. In this letter, we show matrix to intermembrane space, generating electrochemical that one UQCRFS1N links the two 10-subunit CIII protomers gradient across the inner membrane which is harnessed by together to form the intact CIII, which resultantly contains complex V to synthesize ATP, providing the majority of only 21 subunits rather than previously assumed 22 subunits energy acquired by living organisms. Respiratory chain Protein (Fig. 1A and 1B). complexes were reported to interact with each other to form Firstly, we rebuilt the high-resolution crystal structures of supercomplexes, even megacomplex (Guo et al., 2017). bovine CIII (PDB: 2A06) (Huang et al., 2005) and chicken However, despite decades of intensive research, many CIII (PDB:3TGU) (Hao et al., 2012). -

4 Understanding the Role of GNA13 Deregulation in Lymphomagenesis
Integrative Genomics Reveals a Role for GNA13 in Lymphomagenesis by Adrienne Greenough University Program in Genetics and Genomics Duke University Approved: ___________________________ Sandeep Dave, Supervisor ___________________________ Fred Dietrich ___________________________ Jack Keene ___________________________ Yuan Zhuang Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the University Program in Genetics and Genomics in the Graduate School of Duke University 2014 i v ABSTRACT Integrative Genomics Reveals a Role for GNA13 in Lymphomagenesis by Adrienne Greenough University Program in Genetics and Genomics Duke University Approved: ___________________________ Sandeep Dave, Supervisor ___________________________ Fred Dietrich ___________________________ Jack Keene ___________________________ Yuan Zhuang An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the University Program in Genetics and Genomics in the Graduate School of Duke University 2014 Copyright by Adrienne Greenough 2014 Abstract Lymphomas comprise a diverse group of malignancies derived from immune cells. High throughput sequencing has recently emerged as a powerful and versatile method for analysis of the cancer genome and transcriptome. As these data continue to emerge, the crucial work lies in sorting through the wealth of information to hone in on the critical aspects that will give us a better understanding of biology and new insight for how to treat disease. Finding the important signals within these large data sets is one of the major challenges of next generation sequencing. In this dissertation, I have developed several complementary strategies to describe the genetic underpinnings of lymphomas. I begin with developing a better method for RNA sequencing that enables strand-specific total RNA sequencing and alternative splicing profiling in the same analysis. -

Multi-Functionality of Proteins Involved in GPCR and G Protein Signaling: Making Sense of Structure–Function Continuum with In
Cellular and Molecular Life Sciences (2019) 76:4461–4492 https://doi.org/10.1007/s00018-019-03276-1 Cellular andMolecular Life Sciences REVIEW Multi‑functionality of proteins involved in GPCR and G protein signaling: making sense of structure–function continuum with intrinsic disorder‑based proteoforms Alexander V. Fonin1 · April L. Darling2 · Irina M. Kuznetsova1 · Konstantin K. Turoverov1,3 · Vladimir N. Uversky2,4 Received: 5 August 2019 / Revised: 5 August 2019 / Accepted: 12 August 2019 / Published online: 19 August 2019 © Springer Nature Switzerland AG 2019 Abstract GPCR–G protein signaling system recognizes a multitude of extracellular ligands and triggers a variety of intracellular signal- ing cascades in response. In humans, this system includes more than 800 various GPCRs and a large set of heterotrimeric G proteins. Complexity of this system goes far beyond a multitude of pair-wise ligand–GPCR and GPCR–G protein interactions. In fact, one GPCR can recognize more than one extracellular signal and interact with more than one G protein. Furthermore, one ligand can activate more than one GPCR, and multiple GPCRs can couple to the same G protein. This defnes an intricate multifunctionality of this important signaling system. Here, we show that the multifunctionality of GPCR–G protein system represents an illustrative example of the protein structure–function continuum, where structures of the involved proteins represent a complex mosaic of diferently folded regions (foldons, non-foldons, unfoldons, semi-foldons, and inducible foldons). The functionality of resulting highly dynamic conformational ensembles is fne-tuned by various post-translational modifcations and alternative splicing, and such ensembles can undergo dramatic changes at interaction with their specifc partners. -
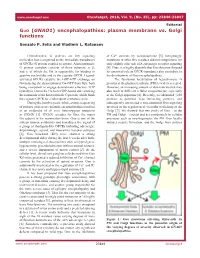
Gαo (GNAO1) Encephalopathies: Plasma Membrane Vs. Golgi Functions
www.oncotarget.com Oncotarget, 2018, Vol. 9, (No. 35), pp: 23846-23847 Editorial Gαo (GNAO1) encephalopathies: plasma membrane vs. Golgi functions Gonzalo P. Solis and Vladimir L. Katanaev Heterotrimeric G proteins are key signaling of Ca2+ currents by norepinephrine [5]. Intriguingly, molecules best recognized as the immediate transducers mutations in other five residues did not compromise (or of GPCRs (G protein coupled receptors). A heterotrimeric only slightly affected) α2A adrenergic receptor signaling G protein complex consists of three subunits, α, β, [4]. Thus, it is highly plausible that Gαo functions beyond and γ, of which the Gα is responsible for binding to its canonical role as GPCR-transducer also contribute to guanine nucleotides and to the cognate GPCR. Ligand- the development of Gαo encephalopathies. activated GPCRs catalyze the GDP-GTP exchange on The functional localization of heterotrimeric G Gα inducing the dissociation of Gα-GTP from Gβγ, both proteins at the plasma membrane (PM) is widely accepted. being competent to engage downstream effectors. GTP However, an increasing amount of data indicate that they hydrolysis returns the Gα to its GDP-bound state allowing also work at different cellular compartments, especially the formation of the heterotrimeric G protein, which binds at the Golgi apparatus [6]. Recently, we identified >250 the cognate GPCR for a subsequent activation cycle. proteins as potential Gαo interacting partners, and During the last five years, whole-exome sequencing subsequently uncovered a non-canonical Gαo signaling of patients with severe infantile encephalopathies resulted involved in the regulation of vesicular trafficking at the in an avalanche of de novo heterozygous mutations Golgi [7].