Choline Kinase Alpha Is a Novel Transcriptional Target of the Brg1 in Hepatocyte: Implication in Liver Regeneration
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

KLF2 Induced
UvA-DARE (Digital Academic Repository) The transcription factor KLF2 in vascular biology Boon, R.A. Publication date 2008 Link to publication Citation for published version (APA): Boon, R. A. (2008). The transcription factor KLF2 in vascular biology. General rights It is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), other than for strictly personal, individual use, unless the work is under an open content license (like Creative Commons). Disclaimer/Complaints regulations If you believe that digital publication of certain material infringes any of your rights or (privacy) interests, please let the Library know, stating your reasons. In case of a legitimate complaint, the Library will make the material inaccessible and/or remove it from the website. Please Ask the Library: https://uba.uva.nl/en/contact, or a letter to: Library of the University of Amsterdam, Secretariat, Singel 425, 1012 WP Amsterdam, The Netherlands. You will be contacted as soon as possible. UvA-DARE is a service provided by the library of the University of Amsterdam (https://dare.uva.nl) Download date:23 Sep 2021 Supplementary data: Genes induced by KLF2 Dekker et al. LocusLink Accession Gene Sequence Description Fold p-value ID number symbol change (FDR) 6654 AK022099 SOS1 cDNA FLJ12037 fis, clone HEMBB1001921. 100.00 5.9E-09 56999 AF086069 ADAMTS9 full length insert cDNA clone YZ35C05. 100.00 1.2E-09 6672 AF085934 SP100 full length insert cDNA clone YR57D07. 100.00 6.7E-13 9031 AF132602 BAZ1B Williams Syndrome critical region WS25 mRNA, partial sequence. -

Angiogenic Patterning by STEEL, an Endothelial-Enriched Long
Angiogenic patterning by STEEL, an endothelial- enriched long noncoding RNA H. S. Jeffrey Mana,b, Aravin N. Sukumara,b, Gabrielle C. Lamc,d, Paul J. Turgeonb,e, Matthew S. Yanb,f, Kyung Ha Kub,e, Michelle K. Dubinskya,b, J. J. David Hob,f, Jenny Jing Wangb,e, Sunit Dasg,h, Nora Mitchelli, Peter Oettgeni, Michael V. Seftonc,d,j, and Philip A. Marsdena,b,e,f,1 aInstitute of Medical Science, University of Toronto, Toronto, ON M5S 1A8, Canada; bKeenan Research Centre for Biomedical Science in the Li Ka Shing Knowledge Institute, St. Michael’s Hospital, University of Toronto, Toronto, ON M5B 1T8, Canada; cDonnelly Centre for Cellular and Biomolecular Research, University of Toronto, Toronto, ON M5S 3E2, Canada; dInstitute of Biomaterials and Biomedical Engineering, University of Toronto, Toronto, ON M5S 3G9, Canada; eDepartment of Laboratory Medicine and Pathobiology, University of Toronto, Toronto, ON M5S 1A8, Canada; fDepartment of Medical Biophysics, University of Toronto, Toronto, ON M5G 1L7, Canada; gArthur and Sonia Labatt Brain Tumour Research Institute, Hospital for SickKids, University of Toronto, Toronto, ON M5G 1X8, Canada; hDivision of Neurosurgery and Keenan Research Centre for Biomedical Science, St. Michael’s Hospital, University of Toronto, Toronto, ON M5B 1W8, Canada; iDepartment of Medicine, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA 02115; and jDepartment of Chemical Engineering and Applied Chemistry, University of Toronto, Toronto, ON M5S 3E5, Canada Edited by Napoleone Ferrara, University of California, San Diego, La Jolla, CA, and approved January 24, 2018 (received for review August 28, 2017) Endothelial cell (EC)-enriched protein coding genes, such as endothelial formation in vitro and blood vessel formation in vivo. -

Detection of Pro Angiogenic and Inflammatory Biomarkers in Patients With
www.nature.com/scientificreports OPEN Detection of pro angiogenic and infammatory biomarkers in patients with CKD Diana Jalal1,2,3*, Bridget Sanford4, Brandon Renner5, Patrick Ten Eyck6, Jennifer Laskowski5, James Cooper5, Mingyao Sun1, Yousef Zakharia7, Douglas Spitz7,9, Ayotunde Dokun8, Massimo Attanasio1, Kenneth Jones10 & Joshua M. Thurman5 Cardiovascular disease (CVD) is the most common cause of death in patients with native and post-transplant chronic kidney disease (CKD). To identify new biomarkers of vascular injury and infammation, we analyzed the proteome of plasma and circulating extracellular vesicles (EVs) in native and post-transplant CKD patients utilizing an aptamer-based assay. Proteins of angiogenesis were signifcantly higher in native and post-transplant CKD patients versus healthy controls. Ingenuity pathway analysis (IPA) indicated Ephrin receptor signaling, serine biosynthesis, and transforming growth factor-β as the top pathways activated in both CKD groups. Pro-infammatory proteins were signifcantly higher only in the EVs of native CKD patients. IPA indicated acute phase response signaling, insulin-like growth factor-1, tumor necrosis factor-α, and interleukin-6 pathway activation. These data indicate that pathways of angiogenesis and infammation are activated in CKD patients’ plasma and EVs, respectively. The pathways common in both native and post-transplant CKD may signal similar mechanisms of CVD. Approximately one in 10 individuals has chronic kidney disease (CKD) rendering CKD one of the most common diseases worldwide1. CKD is associated with a high burden of morbidity in the form of end stage kidney disease (ESKD) requiring dialysis or transplantation 2. Furthermore, patients with CKD are at signifcantly increased risk of death from cardiovascular disease (CVD)3,4. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

Impact of Consuming Extra-Virgin Olive Oil Or Nuts Within a Mediterranean Diet on DNA Methylation in Peripheral White Blood Cell
nutrients Article Impact of Consuming Extra-Virgin Olive Oil or Nuts within a Mediterranean Diet on DNA Methylation in Peripheral White Blood Cells within the PREDIMED-Navarra Randomized Controlled Trial: A Role for Dietary Lipids Ana Arpón 1,2 ID , Fermín I. Milagro 1,2,3 ID , Cristina Razquin 3,4,5, Dolores Corella 3,6 ID , Ramón Estruch 3,7, Montserrat Fitó 3,8, Amelia Marti 1,3,5 ID , Miguel A. Martínez-González 3,4,5, Emilio Ros 3,9, Jordi Salas-Salvadó 3,10 ID , José-Ignacio Riezu-Boj 1,2,5,† ID and J. Alfredo Martínez 1,2,3,5,11,*,† ID 1 Department of Nutrition, Food Sciences and Physiology, University of Navarra, 31008 Pamplona, Spain; [email protected] (A.A.); [email protected] (F.I.M.); [email protected] (A.M.); [email protected] (J.-I.R.-B.) 2 Centre for Nutrition Research, University of Navarra, 31008 Pamplona, Spain 3 Spanish Biomedical Research Centre in Physiopathology of Obesity and Nutrition (CIBERobn), Institute of Health Carlos III, 28029 Madrid, Spain; [email protected] (C.R.); [email protected] (D.C.); [email protected] (R.E.); mfi[email protected] (M.F.); [email protected] (M.A.M.-G.); [email protected] (E.R.); [email protected] (J.S.-S.) 4 Department of Preventive Medicine and Public Health, University of Navarra, 31008 Pamplona, Spain 5 Navarra Institute for Health Research (IdiSNa), 31008 Pamplona, Spain 6 Department of Preventive Medicine and Public Health, University of Valencia, 46010 Valencia, Spain 7 Department of Internal Medicine, Institut d’Investigacions Biomediques August Pi i Sunyer (IDIBAPS), Hospital Clinic, University of Barcelona, 08036 Barcelona, Spain 8 Institut de Recerca Hospital del Mar de Investigacions Mèdiques (IMIM), 08003 Barcelona, Spain 9 Lipid Clinic, Department of Endocrinology and Nutrition, Institut d'Investigacions Biomediques August Pi i Sunyer (IDIBAPS), Hospital Clinic, University of Barcelona, 08036 Barcelona, Spain. -

Inhibition of Adipocyte Differentiation by Rorα
FEBS Letters 583 (2009) 2031–2036 journal homepage: www.FEBSLetters.org Inhibition of adipocyte differentiation by RORa Hélène Duez a,b,c,1, Christian Duhem a,b,c,1, Saara Laitinen a,b,c, Prashant S. Patole a,b,c, Mouaadh Abdelkarim a,b,c, Brigitte Bois-Joyeux d, Jean-Louis Danan d, Bart Staels a,b,c,* a Institut Pasteur de Lille, Département d’Athérosclérose, Lille F-59019, France b INSERM UMR 545, Lille F-59019, France c Université Lille Nord de France, Faculté des Sciences Pharmaceutiques et Biologiques et Faculté de Médecine, Lille F-59006, France d Centre National de la Recherche Scientifique FRE3210, Faculté de Médecine René Descartes Paris 5, 75015 Paris, France article info abstract Article history: Here we show that gene expression of the nuclear receptor RORa is induced during adipogenesis, Received 8 April 2009 with RORa4 being the most abundantly expressed isoform in human and murine adipose tissue. Revised 27 April 2009 Over-expression of RORa4 in 3T3-L1 cells impairs adipogenesis as shown by the decreased expres- Accepted 8 May 2009 sion of adipogenic markers and lipid accumulation, accompanied by decreased free fatty acid and Available online 18 May 2009 glucose uptake. By contrast, mouse embryonic fibroblasts from staggerer mice, which carry a muta- Edited by Robert Barouki tion in the RORa gene, differentiate more efficiently into mature adipocytes compared to wild-type cells, a phenotype which is reversed by ectopic RORa4 restoration. Ó 2009 Federation of European Biochemical Societies. Published by Elsevier B.V. All rights reserved. Keywords: Adipogenesis RORa Nuclear receptors RAR-related orphan receptors (RORs) 1. -

Regulation of T Follicular Helper Cells by ICOS
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 6, No. 26 Editorial Regulation of T follicular helper cells by ICOS Andreas Hutloff T follicular helper (TFH) cells are gatekeepers of the (OPN-i), which facilitates nuclear translocation [3]. In the humoral immune response. Without help from this CD4+ nucleus, OPN-i dimerizes with Bcl-6 and protects it from T cell subset, B cells cannot differentiate into high-affinity proteasomal degradation. However, this OPN-i and the memory B cells and antibody-producing long-lived plasma above Klf2 pathway do act at different times. Upon ICOS cells which are the basis of protective immune responses. signaling blockade, Klf2 is upregulated within a few hours At the same time, dysregulated TFH cell responses are and TFH cells lose their typical homing receptor pattern in causative for many autoimmune disorders (reviewed in less than 24 hours [6]. In contrast, complete degradation [1]). The inducible T cell costimulator ICOS, which is of Bcl-6 takes several days [3, 6]. Therefore, the ICOS - structurally and functionally related to CD28, has been Klf2 axis first leads to emigration of TFH cells out of the known for many years as an important regulator of TFH B cell follicle (and thereby to a loss of function), whereas cells. ICOS knock-out mice as well as ICOS-deficient the osteopontin pathway acts as a second strike pathway patients have only few TFH cells and very small germinal eliminating the lineage-defining transcription factor Bcl-6. centers upon immunization, which results in severely Another important finding especially for potential compromised antigen-specific immunoglobulin levels therapeutic applications is that ICOS and the structurally and the phenotype of common variable immunodeficiency related CD28 molecule act in different phases of TFH cell in humans (reviewed in [2]). -

Choline Kinase Inhibition As a Treatment Strategy for Cancers With
Choline Kinase Inhibition as a Treatment Strategy of Cancers with Deregulated Lipid Metabolism Sebastian Trousil Imperial College London Department of Surgery and Cancer A dissertation submitted for the degree of Doctor of Philosophy 2 Declaration I declare that this dissertation is my own and original work, except where explicitly acknowledged. The copyright of this thesis rests with the author and is made available under a Creative Commons Attribution Non-Commercial No Derivatives licence. Researchers are free to copy, distribute or transmit the thesis on the condition that they attribute it, that they do not use it for commercial purposes and that they do not alter, transform or build upon it. For any reuse or redistribution, researchers must make clear to others the licence terms of this work. Abstract Aberrant choline metabolism is a characteristic shared by many human cancers. It is predominantly caused by elevated expression of choline kinase alpha, which catalyses the phosphorylation of choline to phosphocholine, an essential precursor of membrane lipids. In this thesis, a novel choline kinase inhibitor has been developed and its therapeutic potential evaluated. Furthermore the probe was used to elaborate choline kinase biology. A lead compound, ICL-CCIC-0019 (IC50 of 0.27 0.06 µM), was identified through a focused library screen. ICL-CCIC-0019 was competitive± with choline and non-competitive with ATP. In a selectivity screen of 131 human kinases, ICL-CCIC-0019 inhibited only 5 kinases more than 20% at a concentration of 10 µM(< 35% in all 131 kinases). ICL- CCIC-0019 potently inhibited cell growth in a panel of 60 cancer cell lines (NCI-60 screen) with a median GI50 of 1.12 µM (range: 0.00389–16.2 µM). -

Beta Cell Adaptation to Pregnancy Requires Prolactin Action on Both
www.nature.com/scientificreports OPEN Beta cell adaptation to pregnancy requires prolactin action on both beta and non‑beta cells Vipul Shrivastava1, Megan Lee1, Daniel Lee1, Marle Pretorius1, Bethany Radford1, Guneet Makkar1 & Carol Huang1,2,3* Pancreatic islets adapt to insulin resistance of pregnancy by up regulating β‑cell mass and increasing insulin secretion. Previously, using a transgenic mouse with global, heterozygous deletion of prolactin receptor (Prlr+/−), we found Prlr signaling is important for this adaptation. However, since Prlr is expressed in tissues outside of islets as well as within islets and prolactin signaling afects β‑cell development, to understand β‑cell‑specifc efect of prolactin signaling in pregnancy, we generated a transgenic mouse with an inducible conditional deletion of Prlr from β‑cells. Here, we found that β‑cell‑specifc Prlr reduction in adult mice led to elevated blood glucose, lowed β‑cell mass and blunted in vivo glucose‑stimulated insulin secretion during pregnancy. When we compared gene expression profle of islets from transgenic mice with global (Prlr+/−) versus β‑cell‑specifc Prlr reduction (βPrlR+/−), we found 95 diferentially expressed gene, most of them down regulated in the Prlr+/− mice in comparison to the βPrlR+/− mice, and many of these genes regulate apoptosis, synaptic vesicle function and neuronal development. Importantly, we found that islets from pregnant Prlr+/− mice are more vulnerable to glucolipotoxicity‑induced apoptosis than islets from pregnant βPrlR+/− mice. These observations suggest that down regulation of prolactin action during pregnancy in non‑β‑cells secondarily and negatively afect β‑cell gene expression, and increased β‑cell susceptibility to external insults. -

Progression of a Muscular Dystrophy Due to a Genetic Defect in Membrane Synthesis Is Driven by Large Changes in Neutral Lipid Metabolism
Progression of a Muscular Dystrophy Due to a Genetic Defect in Membrane Synthesis is Driven by Large Changes in Neutral Lipid Metabolism Mahtab Tavasoli Dalhousie University https://orcid.org/0000-0001-7962-3566 Sarah Lahire University of Reims Champagne-Ardenne Stanislav Sokolenko Dalhousie University J. Pedro Fernández-Murray1 Dalhousie University Kitipong Uaesoontrachoon Dalhousie University/Agada Biosciences Abir Lefsay Dalhousie University Joyce Rowsell Agada Biosciences Sadish Srinivassane Agada Biosciences Molly Praest Agada Biosciences Alexandra MacKinnon Agada Biosciences Melissa Mammoliti Agada Biosciences Ashley Maloney Agada Biosciences Marina Moraca Agada Biosciences Kanneboyina Nagaraju Department of Pharmaceutical Sciences, School of Pharmacy and Pharmaceutical Sciences, Binghamton University Eric Hoffman School of Pharmacy and Pharmaceutical Sciences Christopher McMaster ( [email protected] ) Dalhousie University Article Keywords: metabolism, muscular dystrophy, skeletal muscle, lipid, acylcarnitine, triacylglycerol, phosphatidylcholine, peroxisome proliferator-activated receptors (PPAR) Posted Date: May 19th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-64129/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Progression of a Muscular Dystrophy Due to a Genetic Defect in Membrane Synthesis is Driven by Large Changes in Neutral Lipid Metabolism Mahtab Tavasoli1, Sarah Lahire2, Stanislav Sokolenko3, J. Pedro Fernandez-Murray1, Kitipong Uaesoontrachoon4, -

CHKL (CHKB) (NM 005198) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP310253 CHKL (CHKB) (NM_005198) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Recombinant protein of human choline kinase beta (CHKB), transcript variant 1 Species: Human Expression Host: HEK293T Tag: C-Myc/DDK Predicted MW: 45.1 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 25 mM Tris.HCl, pH 7.3, 100 mM glycine, 10% glycerol Preparation: Recombinant protein was captured through anti-DDK affinity column followed by conventional chromatography steps. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_005189 Locus ID: 1120 UniProt ID: Q9Y259, A0A024R4X4 RefSeq Size: 1595 Cytogenetics: 22q13.33 RefSeq ORF: 1185 Synonyms: CHETK; CHKL; CK; CKB; CKEKB; EK; EKB; MDCMC This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 CHKL (CHKB) (NM_005198) Human Recombinant Protein – TP310253 Summary: Choline kinase (CK) and ethanolamine kinase (EK) catalyze the phosphorylation of choline/ethanolamine to phosphocholine/phosphoethanolamine. This is the first enzyme in the biosynthesis of phosphatidylcholine/phosphatidylethanolamine in all animal cells. The highly purified CKs from mammalian sources and their recombinant gene products have been shown to have EK activity also, indicating that both activities reside on the same protein. -
Targeted Genes Common Elements In
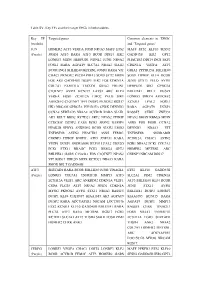
Table SV. Key TFs and their target DEGs in hub modules. Key TF Targeted genes Common elements in ‘DEGs’ (module) and ‘Targeted genes’ JUN HOMER2 ATF3 VEGFA FOSB NR4A3 MAFF ETS2 MAFF ETS2 KLF10 SESN2 (Purple) JOSD1 ATF3 RARA ATF3 BCOR DDIT4 IER2 GADD45B IER2 GPT2 LONRF3 MIDN HERPUD1 NDNL2 JUNB NR4A2 PHACTR3 DDIT4 ING1 SKP2 FOSL1 RARA AGPAT9 SLC7A1 NR4A2 SIAH2 CDKN1A VEGFA ATF3 BCOR ING1 BHLHE40 METRNL JOSD1 RARA VIT GRIA2 PPP1R15A BHLHE40 CHAC1 PKNOX2 PLCB4 PHF13 SOX9 SYT2 MIDN SOX9 ITPRIP KLF4 BCOR FOS AK5 GADD45B DUSP1 IER2 FOS CDKN1A JUNB STX11 PELO AVPI1 COL7A1 FAM131A TBCCD1 GRIA2 ERLIN1 HERPUD1 SIK1 GPRC5A C1QTNF7 AVPI1 KCNJ15 LATS2 ARC KLF4 BHLHE41 RELT DUSP2 VEGFA FOSB ZC3H12A LMO2 PELO SIK1 LONRF3 SPRY4 ARHGEF2 ARHGEF2 C1QTNF7 TFPI DUSP2 PKNOX2 RGS17 KCNJ15 LPAL2 FOSL1 HK1 NRCAM GPRC5A PPP1R15A CERK DENND3 RARA AGPAT9 DUSP1 CCNA2 SERTAD3 NR4A2 ACVR1B RARA SIAH1 RASSF5 CERK ZNF331 AK5 RELT BRD2 KCTD21 SKP2 NPAS2 ITPRIP NPAS2 SMOX RBM24 MIDN CCDC85C DUSP2 CARS EGR1 SESN2 RASSF5 ASNS FOS FOSB CCNA2 PIP4K2B SPRY4 ANKRD52 BCOR SIAH2 LMO2 DENND3 NR4A3 VIT TNFRSF1B ASTN2 PHACTR3 ASNS FEM1C TNFRSF1B SNORA80B CSRNP3 ITPRIP BNIP3L ATF3 ZNF331 RARA ZC3H12A CHAC1 ASTN2 VEZF1 DUSP1 SNORA80B KLF10 LPAL2 UBE2O EGR1 NR4A2 PCK1 COL7A1 PCK1 STX11 RRAGC PCK1 RBM24 GPT2 HOMER2 METRNL ARC BHLHE41 GARS C15orf41 FOS C1QTNF7 NPAS2 CSRNP3 NRCAM RGS17 VIT RGS17 UBE2O SOX9 KCTD21 NR4A3 RARA SMOX RELT GADD45B ATF3 SERTAD1 RARA BCOR BHLHE40 JUNB TINAGL1 ETS2 KLF10 GADD45B (Purple) LONRF3 COL7A1 TNFRSF1B MMP13 ATF3 SLC2A1 PIM2 CDKN1A ZC3H12A VEZF1 ARC ANKRD52