Targeting the Ubiquitination/Deubiquitination Process to Regulate Immune Checkpoint Pathways
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ENSG Gene Encodes Effector TCR Pathway Costimulation Inhibitory/Exhaustion Synapse/Adhesion Chemokines/Receptors
ENSG Gene Encodes Effector TCR pathway Costimulation Inhibitory/exhaustion Synapse/adhesion Chemokines/receptors ENSG00000111537 IFNG IFNg x ENSG00000109471 IL2 IL-2 x ENSG00000232810 TNF TNFa x ENSG00000271503 CCL5 CCL5 x x ENSG00000139187 KLRG1 Klrg1 x ENSG00000117560 FASLG Fas ligand x ENSG00000121858 TNFSF10 TRAIL x ENSG00000134545 KLRC1 Klrc1 / NKG2A x ENSG00000213809 KLRK1 Klrk1 / NKG2D x ENSG00000188389 PDCD1 PD-1 x x ENSG00000117281 CD160 CD160 x x ENSG00000134460 IL2RA IL-2 receptor x subunit alpha ENSG00000110324 IL10RA IL-10 receptor x subunit alpha ENSG00000115604 IL18R1 IL-18 receptor 1 x ENSG00000115607 IL18RAP IL-18 receptor x accessory protein ENSG00000081985 IL12RB2 IL-12 receptor x beta 2 ENSG00000186810 CXCR3 CXCR3 x x ENSG00000005844 ITGAL CD11a x ENSG00000160255 ITGB2 CD18; Integrin x x beta-2 ENSG00000156886 ITGAD CD11d x ENSG00000140678 ITGAX; CD11c x x Integrin alpha-X ENSG00000115232 ITGA4 CD49d; Integrin x x alpha-4 ENSG00000169896 ITGAM CD11b; Integrin x x alpha-M ENSG00000138378 STAT4 Stat4 x ENSG00000115415 STAT1 Stat1 x ENSG00000170581 STAT2 Stat2 x ENSG00000126561 STAT5a Stat5a x ENSG00000162434 JAK1 Jak1 x ENSG00000100453 GZMB Granzyme B x ENSG00000145649 GZMA Granzyme A x ENSG00000180644 PRF1 Perforin 1 x ENSG00000115523 GNLY Granulysin x ENSG00000100450 GZMH Granzyme H x ENSG00000113088 GZMK Granzyme K x ENSG00000057657 PRDM1 Blimp-1 x ENSG00000073861 TBX21 T-bet x ENSG00000115738 ID2 ID2 x ENSG00000176083 ZNF683 Hobit x ENSG00000137265 IRF4 Interferon x regulatory factor 4 ENSG00000140968 IRF8 Interferon -

The Effect of Temperature Adaptation on the Ubiquitin–Proteasome Pathway in Notothenioid Fishes Anne E
© 2017. Published by The Company of Biologists Ltd | Journal of Experimental Biology (2017) 220, 369-378 doi:10.1242/jeb.145946 RESEARCH ARTICLE The effect of temperature adaptation on the ubiquitin–proteasome pathway in notothenioid fishes Anne E. Todgham1,*, Timothy A. Crombie2 and Gretchen E. Hofmann3 ABSTRACT proliferation to compensate for the effects of low temperature on ’ There is an accumulating body of evidence suggesting that the sub- aerobic metabolism (Johnston, 1989; O Brien et al., 2003; zero Antarctic marine environment places physiological constraints Guderley, 2004). Recently, there has been an accumulating body – on protein homeostasis. Levels of ubiquitin (Ub)-conjugated proteins, of literature to suggest that protein homeostasis the maintenance of – 20S proteasome activity and mRNA expression of many proteins a functional protein pool has been highly impacted by evolution involved in both the Ub tagging of damaged proteins as well as the under these cold and stable conditions. different complexes of the 26S proteasome were measured to Maintaining protein homeostasis is a fundamental physiological examine whether there is thermal compensation of the Ub– process, reflecting a dynamic balance in synthetic and degradation proteasome pathway in Antarctic fishes to better understand the processes. There are numerous lines of evidence to suggest efficiency of the protein degradation machinery in polar species. Both temperature compensation of protein synthesis in Antarctic Antarctic (Trematomus bernacchii, Pagothenia borchgrevinki)and invertebrates (Whiteley et al., 1996; Marsh et al., 2001; Robertson non-Antarctic (Notothenia angustata, Bovichtus variegatus) et al., 2001; Fraser et al., 2002) and fish (Storch et al., 2005). In notothenioids were included in this study to investigate the zoarcid fishes, it has been demonstrated that Antarctic eelpouts mechanisms of cold adaptation of this pathway in polar species. -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -
![UBE2E1 (Ubch6) [Untagged] E2 – Ubiquitin Conjugating Enzyme](https://docslib.b-cdn.net/cover/0534/ube2e1-ubch6-untagged-e2-ubiquitin-conjugating-enzyme-320534.webp)
UBE2E1 (Ubch6) [Untagged] E2 – Ubiquitin Conjugating Enzyme
UBE2E1 (UbcH6) [untagged] E2 – Ubiquitin Conjugating Enzyme Alternate Names: UbcH6, UbcH6, Ubiquitin conjugating enzyme UbcH6 Cat. No. 62-0019-100 Quantity: 100 µg Lot. No. 1462 Storage: -70˚C FOR RESEARCH USE ONLY NOT FOR USE IN HUMANS CERTIFICATE OF ANALYSIS Page 1 of 2 Background Physical Characteristics The enzymes of the ubiquitylation Species: human Protein Sequence: pathway play a pivotal role in a num- GPLGSPGIPGSTRAAAM SDDDSRAST ber of cellular processes including Source: E. coli expression SSSSSSSSNQQTEKETNTPKKKESKVSMSKN regulated and targeted proteasomal SKLLSTSAKRIQKELADITLDPPPNCSAGP degradation of substrate proteins. Quantity: 100 μg KGDNIYEWRSTILGPPGSVYEGGVFFLDIT FTPEYPFKPPKVTFRTRIYHCNINSQGVI Three classes of enzymes are in- Concentration: 1 mg/ml CLDILKDNWSPALTISKVLLSICSLLTDCNPAD volved in the process of ubiquitylation; PLVGSIATQYMTNRAEHDRMARQWTKRYAT activating enzymes (E1s), conjugating Formulation: 50 mM HEPES pH 7.5, enzymes (E2s) and protein ligases 150 mM sodium chloride, 2 mM The residues underlined remain after cleavage and removal (E3s). UBE2E1 is a member of the E2 dithiothreitol, 10% glycerol of the purification tag. ubiquitin-conjugating enzyme family UBE2E1 (regular text): Start bold italics (amino acid and cloning of the human gene was Molecular Weight: ~23 kDa residues 1-193) Accession number: AAH09139 first described by Nuber et al. (1996). UBE2E1 shares 74% sequence ho- Purity: >98% by InstantBlue™ SDS-PAGE mology with UBE2D1 and contains an Stability/Storage: 12 months at -70˚C; N-terminal extension of approximately aliquot as required 40 amino acids. A tumour suppressor candidate, tumour-suppressing sub- chromosomal transferable fragment Quality Assurance cDNA (TSSC5) is located in the re- gion of human chromosome 11p15.5 Purity: Protein Identification: linked with Beckwith-Wiedemann syn- 4-12% gradient SDS-PAGE Confirmed by mass spectrometry. -

SLPI and Soluble BTLA As Immunological Markers in Severe Bacterial Infections
SLPI and soluble BTLA as immunological markers in severe bacterial infections To my family Örebro Studies in Medicine 211 ANNA LANGE SLPI and soluble BTLA as immunological markers in severe bacterial infections © Anna Lange, 2020 Title: SLPI and soluble BTLA as immunological markers in severe bacterial infections Publisher: Örebro University 2020 www.oru.se/publikationer Print: Örebro University, Repro 04/2020 ISSN 1652-4063 ISBN 978-91-7529-335-6 Abstract Anna Lange (2020): SLPI and soluble BTLA as immunological markers in severe bacterial infections. Örebro Studies in Medicine 211. Clinical presentation, and outcome of infections are affected by host-, and etiology- (focus of infection and pathogen) related factors. The im- mune response is controlled by a network of regulating pathways. This thesis focuses on Secretory Leukocyte Protease Inhibitor (SLPI), a protease inhibitor with anti-inflammatory properties, and the previously non-studied soluble isoform of B and T lymphocyte attenuator (sBTLA), a membrane-associated regulatory protein. Plasma concentrations of SLPI and sBTLA were assessed in relation to etiology, severity, mortality, and markers of inflammation and immunosuppression, in i) community- acquired pneumonia (CAP) (SLPI), ii) intensive care unit (ICU) treated severe sepsis and septic shock (sBTLA), and iii) dynamically in BSI (SLPI and sBTLA). Main findings were: higher expression of SLPI in pneumonia, com- pared to other sources, higher initial concentrations in Streptococcus pneumoniae, and Staphylococcus aureus BSI, compared to Escherichia coli BSI, and higher SLPI concentrations in sepsis compared to non-septic BSI. Interestingly, men with pneumonia had higher plasma levels of SLPI, both in CAP and BSI. Likewise, sBTLA was associated with severity, but preferentially at higher organ failure scores. -

Gain of UBE2D1 Facilitates Hepatocellular Carcinoma
Zhou et al. Journal of Experimental & Clinical Cancer Research (2018) 37:290 https://doi.org/10.1186/s13046-018-0951-8 RESEARCH Open Access Gain of UBE2D1 facilitates hepatocellular carcinoma progression and is associated with DNA damage caused by continuous IL-6 Chuanchuan Zhou1,2, Fengrui Bi1, Jihang Yuan1, Fu Yang1 and Shuhan Sun1* Abstract Background: Hepatocellular carcinoma (HCC) is the most common type of liver cancer with increasing incidence and poor prognosis. Ubiquitination regulators are reported to play crucial roles in HCC carcinogenesis. UBE2D1, one of family member of E2 ubiquitin conjugating enzyme, mediates the ubiquitination and degradation of tumor suppressor protein p53. However, the expression and functional roles of UBE2D1 in HCC was unknown. Methods: Immunohistochemistry (IHC), western blotting, and real-time PCR were used to detect the protein, transcription and genomic levels of UBE2D1 in HCC tissues with paired nontumor tissues, precancerous lesions and hepatitis liver tissues. Four HCC cell lines and two immortalized hepatic cell lines were used to evaluate the functional roles and underlying mechanisms of UBE2D1 in HCC initiation and progression in vitro and in vivo. The contributors to UBE2D1 genomic amplification were first evaluated by performing a correlation analysis between UBE2D1 genomic levels with clinical data of HCC patients, and then evaluated in HCC and hepatic cell lines. Results: Expression of UBE2D1 was significantly increased in HCC tissues and precancerous lesions and was associated with reduced survival of HCC patients. Upregulation of UBE2D1 promoted HCC growth in vitro and in vivo by decreasing the p53 in ubiquitination-dependent pathway. High expression of UBE2D1 was attributed to the recurrent genomic copy number gain, which was associated with high serum IL-6 level of HCC patients. -

UBE2B Sirna Set I Sirna Duplexes Targeted Against Three Exon Regions
Catalog # Aliquot Size U211-911-05 3 x 5 nmol U211-911-20 3 x 20 nmol U211-911-50 3 x 50 nmol UBE2B siRNA Set I siRNA duplexes targeted against three exon regions Catalog # U211-911 Lot # Z2109-16 Specificity Formulation UBE2B siRNAs are designed to specifically knock-down The siRNAs are supplied as a lyophilized powder and human UBE2B expression. shipped at room temperature. Product Description Reconstitution Protocol UBE2B siRNA is a pool of three individual synthetic siRNA Briefly centrifuge the tubes (maximum RCF 4,000g) to duplexes designed to knock-down human UBE2B mRNA collect lyophilized siRNA at the bottom of the tube. expression. Each siRNA is 19-25 bases in length. The gene Resuspend the siRNA in 50 µl of DEPC-treated water accession number is NM_003337. (supplied by researcher), which results in a 1x stock solution (10 µM). Gently pipet the solution 3-5 times to mix Gene Aliases and avoid the introduction of bubbles. Optional: aliquot E2-17kDa; HHR6B; HR6B; RAD6B; UBC2 1x stock solutions for storage. Storage and Stability Related Products The lyophilized powder is stable for at least 4 weeks at room temperature. It is recommended that the Product Name Catalog Number lyophilized and resuspended siRNAs are stored at or UBE2A Protein U210-30H below -20oC. After resuspension, siRNA stock solutions ≥2 UBE2B Protein U211-30H µM can undergo up to 50 freeze-thaw cycles without UBE2C Protein U212-30H significant degradation. For long-term storage, it is UBE2D1 (UBCH5A) U213-30H recommended that the siRNA is stored at -70oC. For most Protein favorable performance, avoid repeated handling and UBE2D2 (UBC4) Protein U214-30H multiple freeze/thaw cycles. -

The X-Linked Intellectual Disability Gene Product and E3 Ubiquitin Ligase KLHL15 Degrades Doublecortin Proteins to Constrain Neuronal Dendritogenesis
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.02.324285; this version posted October 2, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. KLHL15 degrades doublecortin proteins The X-linked intellectual disability gene product and E3 ubiquitin ligase KLHL15 degrades doublecortin proteins to constrain neuronal dendritogenesis Jianing Song1,2, Ronald A. Merrill1,2, Andrew Y. Usachev1, and Stefan Strack1* From the 1Department of Neuroscience and Pharmacology and the Iowa Neuroscience Institute, University of Iowa, Iowa City, Iowa 52242 2 These authors contributed equally to the work. *To whom correspondence should be addressed: Stefan Strack: Dept. of Neuroscience and Pharmacology, The University of Iowa, Iowa City, IA 52242; [email protected]; Tel. (319) 384-4439; Fax. (319) 335-8930 Running Title: KLHL15 degrades doublecortin proteins Keywords: Kelch-like 15, KLHL15, E3 ubiquitin ligase, protein phosphatase 2A, signal transduction, doublecortin, doublecortin-like kinases, microtubule-associated protein, ubiquitination, proteasomal degradation, protein turnover, neurite outgrowth, dendritic complexity, pulse-chase, HaloTag ______________________________________________________________________________ ABSTRACT doublecortin (DCX), also an X-linked disease gene, and doublecortin-like kinases 1 and 2 Proper brain development and function (DCLK1/2) as bona fide KLHL15 interactors requires finely controlled mechanisms -
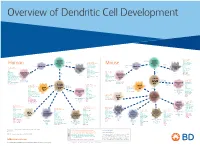
View Dendritic Cell Development Poster
Overview of Dendritic Cell Development Lineage–, CD45+, Common CD117 (c-kit) Common MHCII+, CD11c+ – + CD207 (Langerin) Myeloid CD117 (c-kit) Lineage , CD45 , Myeloid Progenitor MHCII (HLA-DR)+, CD11c+ Progenitor CD324 (E-Cadherin) Human Mouse CD326 (EpCAM) CD207 (Langerin) TGFb1 Cells CD11b, CD115 Cells CD14 Monocyte CD324 (E-Cadherin) Monocyte M-CSF CD11b – + Ly6C Langerhans CD24 Lineage , CD45 , M-CSF CD326 (EpCAM) MHCII (HLA-DR)+, CD11c+ Langerhans CD11blo Zbtb46– Cells CD172a (Sirp-α) CD16 CD1ahi, CD1c CD205 (DEC-205) Cells CSF F4/80 CD64 CD172a (Sirp-α) Lineage–, CD45+, FLT3L TLR3, TLR11 CD1a, CD1c Inflammatory CD369 (Dectin-1/CLEC7A) MHCII+, CD11c+ +/– CSF IL-15 CD8–, CD14– CD11b, CD14 CD371 (CLEC12A) CD64 Monocyte- FLT3L Inflammatory CD370 (Clec9a)– CD172a (Sirp-α) IL-15 CLEC6A CD11b derived lo Monocyte- CD206, CD209 (DC-SIGN) TLR1, TLR2, TLR3 , TLR6 CD209a (DC-SIGN) CD367 (DCIR/CLEC4A) DCs CD14– CD272 (BTLA)lo derived CD369 (Dectin-1/CLEC7A) DCs Common Ly-6C – + CD371 (CLEC12A) CD117 (c-kit) Lineage , CD45 , IL-1β, IL-6, IL-10, TLR1-6, TLR7-8, TLR10 Dendritic + lo CLEC6A – – CD135/FLT3 MHCII , CD11c IL-12, IL-23, TNF CD8a , CD14 IL-1β, IL-6 IL-10, Precursor TLR3lo, TLR4, TLR7, TLR8 CD45R (B220) IL-12, IL-23, TNF Plasmacytoid CD207 (Langerin)– Cells CD317 (BST-2) Common Lineage–, CD45+, FLT3L DCs Lineage–, CD45+, + Ly6C + lo/– CD207 IFN Type I + + Dendritic CD135/FLT3 MHCII (HLA-DR) , CD11c Lineage–, CD45+, IRF7, IRF8, BATF3hi Siglec-H MHCII (HLA-DR) , CD11c hi – + CD123 + + Dermal SpiB, Zbtb46 CD1a, CD64 CD1a Precursor CD117 (c-kit) -

UBE2C Is Upregulated by Estrogen and Promotes Epithelial–Mesenchymal Transition Via P53 in Endometrial Cancer
Published OnlineFirst October 29, 2019; DOI: 10.1158/1541-7786.MCR-19-0561 MOLECULAR CANCER RESEARCH | CANCER GENES AND NETWORKS UBE2C Is Upregulated by Estrogen and Promotes Epithelial–Mesenchymal Transition via p53 in Endometrial Cancer Yan Liu1, Rong Zhao1, Shuqi Chi1, Wei Zhang1, Chengyu Xiao1, Xing Zhou1, Yingchao Zhao2, and Hongbo Wang1 ABSTRACT ◥ Ubiquitin-conjugating enzyme E2C (UBE2C) plays important inhibited endometrial cancer cell proliferation, migration, invasion, roles in tumor progression; nevertheless, its function in endometrial and epithelial–mesenchymal transition (EMT), whereas UBE2C cancer remains unclear. This study elucidated the impact of UBE2C overexpression exerted the opposite effects. UBE2C downregulation on endometrial cancer and its underlying mechanism. Human increased p53 and its downstream p21 expression, with p53 over- endometrial cancer and normal endometrial tissues were acquired expression reversing the EMT-promoting effects of UBE2C. UBE2C from patients at Wuhan Union Hospital and UBE2C expression was enhanced p53 ubiquitination to facilitate its degradation in endo- detected by Western blotting and qRT-PCR. Endometrial cancer metrial cancer cells. Estradiol (E2) induced UBE2C expression via cells were transfected with a UBE2C overexpression plasmid or estrogen receptor a, which binds directly to the UBE2C promoter UBE2C-specific short hairpin RNA (shRNA) to up- or downregu- element. Silencing of UBE2C inhibited E2-promoted migration, late UBE2C expression, respectively. CCK8 and transwell assays -

BTLA−HVEM Checkpoint Axis Regulates Hepatic Homeostasis and Inflammation in a Cona-Induced Hepatitis Model in Zebrafish
BTLA−HVEM Checkpoint Axis Regulates Hepatic Homeostasis and Inflammation in a ConA-Induced Hepatitis Model in Zebrafish This information is current as Wei Shi, Tong Shao, Jiang-yuan Li, Dong-dong Fan, Ai-fu of September 27, 2021. Lin, Li-xin Xiang and Jian-zhong Shao J Immunol published online 27 September 2019 http://www.jimmunol.org/content/early/2019/09/26/jimmun ol.1900458 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2019/09/26/jimmunol.190045 Material 8.DCSupplemental http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 27, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2019 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Published September 27, 2019, doi:10.4049/jimmunol.1900458 The Journal of Immunology BTLA–HVEM Checkpoint Axis Regulates Hepatic Homeostasis and Inflammation in a ConA-Induced Hepatitis Model in Zebrafish Wei Shi,* Tong Shao,* Jiang-yuan Li,* Dong-dong Fan,* Ai-fu Lin,* Li-xin Xiang,* and Jian-zhong Shao*,† The BTLA2HVEM checkpoint axis plays extensive roles in immunomodulation and diseases, including cancer and autoimmune disorders. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement.