Proteome-Wide Survey of the Autoimmune Target Repertoire In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cytokine Nomenclature
RayBiotech, Inc. The protein array pioneer company Cytokine Nomenclature Cytokine Name Official Full Name Genbank Related Names Symbol 4-1BB TNFRSF Tumor necrosis factor NP_001552 CD137, ILA, 4-1BB ligand receptor 9 receptor superfamily .2. member 9 6Ckine CCL21 6-Cysteine Chemokine NM_002989 Small-inducible cytokine A21, Beta chemokine exodus-2, Secondary lymphoid-tissue chemokine, SLC, SCYA21 ACE ACE Angiotensin-converting NP_000780 CD143, DCP, DCP1 enzyme .1. NP_690043 .1. ACE-2 ACE2 Angiotensin-converting NP_068576 ACE-related carboxypeptidase, enzyme 2 .1 Angiotensin-converting enzyme homolog ACTH ACTH Adrenocorticotropic NP_000930 POMC, Pro-opiomelanocortin, hormone .1. Corticotropin-lipotropin, NPP, NP_001030 Melanotropin gamma, Gamma- 333.1 MSH, Potential peptide, Corticotropin, Melanotropin alpha, Alpha-MSH, Corticotropin-like intermediary peptide, CLIP, Lipotropin beta, Beta-LPH, Lipotropin gamma, Gamma-LPH, Melanotropin beta, Beta-MSH, Beta-endorphin, Met-enkephalin ACTHR ACTHR Adrenocorticotropic NP_000520 Melanocortin receptor 2, MC2-R hormone receptor .1 Activin A INHBA Activin A NM_002192 Activin beta-A chain, Erythroid differentiation protein, EDF, INHBA Activin B INHBB Activin B NM_002193 Inhibin beta B chain, Activin beta-B chain Activin C INHBC Activin C NM005538 Inhibin, beta C Activin RIA ACVR1 Activin receptor type-1 NM_001105 Activin receptor type I, ACTR-I, Serine/threonine-protein kinase receptor R1, SKR1, Activin receptor-like kinase 2, ALK-2, TGF-B superfamily receptor type I, TSR-I, ACVRLK2 Activin RIB ACVR1B -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Secretome Screening Reveals Immunomodulating Functions Of
www.nature.com/scientificreports OPEN Secretome screening reveals immunomodulating functions of IFNα‑7, PAP and GDF‑7 on regulatory T‑cells Mei Ding1*, Rajneesh Malhotra2, Tomas Ottosson2, Magnus Lundqvist3, Aman Mebrahtu3, Johan Brengdahl1, Ulf Gehrmann2, Elisabeth Bäck4, Douglas Ross‑Thriepland5, Ida Isaksson6, Björn Magnusson1, Kris F. Sachsenmeier7, Hanna Tegel3, Sophia Hober3, Mathias Uhlén3, Lorenz M. Mayr5, Rick Davies5, Johan Rockberg3 & Lovisa Holmberg Schiavone1* Regulatory T cells (Tregs) are the key cells regulating peripheral autoreactive T lymphocytes. Tregs exert their function by suppressing efector T cells. Tregs have been shown to play essential roles in the control of a variety of physiological and pathological immune responses. However, Tregs are unstable and can lose the expression of FOXP3 and suppressive functions as a consequence of outer stimuli. Available literature suggests that secreted proteins regulate Treg functional states, such as diferentiation, proliferation and suppressive function. Identifcation of secreted proteins that afect Treg cell function are highly interesting for both therapeutic and diagnostic purposes in either hyperactive or immunosuppressed populations. Here, we report a phenotypic screening of a human secretome library in human Treg cells utilising a high throughput fow cytometry technology. Screening a library of 575 secreted proteins allowed us to identify proteins stabilising or destabilising the Treg phenotype as suggested by changes in expression of Treg marker proteins FOXP3 and/or CTLA4. Four proteins including GDF‑7, IL‑10, PAP and IFNα‑7 were identifed as positive regulators that increased FOXP3 and/or CTLA4 expression. PAP is a phosphatase. A catalytic‑dead version of the protein did not induce an increase in FOXP3 expression. -

Silkworm Z Chromosome Is Enriched in Testis-Specific Genes
Supporting Information http://www.genetics.org/cgi/content/full/genetics.108.099994/DC1 Silkworm Z Chromosome is Enriched in Testis-Specific Genes K. P. Arunkumar, Kazuei Mita and J. Nagaraju Copyright © 2009 by the Genetics Society of America DOI: 10.1534/genetics.108.099994 2 SI K. Arunkumar et al. File S1 Gene Ontology annotation GO annotation generates a dynamic controlled vocabulary that can be applied to all organisms, even while knowledge of gene and protein roles in cells is still accumulating and changing. To this end, the Seqdblite FASTA sequence flat file was downloaded from the GO database. By running BLAST against Seqdblite, closest homologue was identified. From BLAST output, molecular functions, biological processes and cellular localisation were parsed by building an in-house GO database in MySQL from the GO-term-database flat file, downloaded from Gene Ontology Database Downloads (http://www.godatabase.org/dev/). The Perl-DBI was used to interface with MySQL, to extract the parent terms of each individual GO term that are obtained by parsing BLAST output. The output was then represented graphically. All ESTs were assigned a biological process, molecular function and cellular component using Gene Ontology (GO) database. The closest annotated homologue in the GO database was used for assigning these categories. The results of the GO annotation are graphically represented in Figures S1-3. Many of the gene products were found to be localized in cell (42%). In cell, gene products were predominant in intracellular region (78%) which comprised of localizations in intracellular organelle (38%) and cytoplasm (29%). The other localizations were organelle (29%) followed by protein complex (18%) (Figure S1). -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Affiliations
Supplementary material Proteome-wide survey of the autoimmune target repertoire in autoimmune polyendocrine syndrome type 1 *Nils Landegren1,2, Donald Sharon3,4, Eva Freyhult2,5,6,, Åsa Hallgren1,2, Daniel Eriksson1,2, Per-Henrik Edqvist7, Sophie Bensing8, Jeanette Wahlberg9, Lawrence M. Nelson10, Jan Gustafsson11, Eystein S Husebye12, Mark S Anderson13, Michael Snyder3, Olle Kämpe1,2 Nils Landegren and Donald Sharon contributed equally to the work Affiliations 1Department of Medicine (Solna), Karolinska University Hospital, Karolinska Institutet, Sweden 2Science for Life Laboratory, Department of Medical Sciences, Uppsala University, Sweden 3Department of Genetics, Stanford University, California, USA 4Department of Molecular, Cellular, and Developmental Biology, Yale University, Connecticut, USA 1 5Department of Medical Sciences, Cancer Pharmacology and Computational Medicine, Uppsala University 6Bioinformatics Infrastructure for Life Sciences 7Department of Immunology, Genetics and Pathology, Uppsala University, Sweden and Science for Life Laboratory 8 Department of Molecular Medicine and Surgery, Karolinska Institutet, Stockholm, Sweden 9Department of Endocrinology and Department of Medical and Health Sciences and Department of Clinical and Experimental Medicine, Linköping University, Linköping, Sweden 10Integrative Reproductive Medicine Group, Intramural Research Program on Reproductive and Adult Endocrinology, National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD 20892, USA. 11Department -

Heatmaps - the Gene Expression Edition
Heatmaps - the gene expression edition Jeff Oliver 20 July, 2020 An application of heatmap visualization to investigate differential gene expression. Learning objectives 1. Manipulate data into a ‘tidy’ format 2. Visualize data in a heatmap 3. Become familiar with ggplot syntax for customizing plots Heatmaps for differential gene expression Heatmaps are a great way of displaying three-dimensional data in only two dimensions. But how can we easily translate tabular data into a format for heatmap plotting? By taking advantage of “data munging” and graphics packages, heatmaps are relatively easy to produce in R. Getting started We are going to start by isolating different types of information by imposing structure in our file managment. That is, we are going to put our input data in one folder and any output such as plots or analytical results in a different folder. We can use the dir.create to create these two folders: dir.create("data") dir.create("output") For this lesson, we will use a subset of data on a study of gene expression in cells infected with the influenza virus (doi: 10.4049/jimmunol.1501557). The study infected human plasmacytoid dendritic cells with the influenza virus, and compared gene expression in those cells to gene expression in uninfected cells. Thegoal was to see how the flu virus affected the function of these immune system cells. The data for this lesson is available at: http://tinyurl.com/flu-expression-data or https://jcoliver.github.io/ learn-r/data/GSE68849-expression.csv. Download this comma separated file and put it in the data folder. -

Microarray Analysis of Colorectal Cancer Stromal Tissue Reveals Upregulation of Two Oncogenic Mirna Clusters
Published OnlineFirst March 27, 2012; DOI: 10.1158/1078-0432.CCR-11-1078 Clinical Cancer Human Cancer Biology Research Microarray Analysis of Colorectal Cancer Stromal Tissue Reveals Upregulation of Two Oncogenic miRNA Clusters Naohiro Nishida1,4, Makoto Nagahara2, Tetsuya Sato3, Koshi Mimori1, Tomoya Sudo1, Fumiaki Tanaka1, Kohei Shibata1, Hideshi Ishii1,4, Kenichi Sugihara2, Yuichiro Doki4, and Masaki Mori1,4 Abstract Purpose: Cancer stroma plays an important role in the progression of cancer. Although alterations in miRNA expression have been explored in various kinds of cancers, the expression of miRNAs in cancer stroma has not been explored in detail. Experimental Design: Using a laser microdissection technique, we collected RNA samples specific for epithelium or stroma from 13 colorectal cancer tissues and four normal tissues, and miRNA microarray and gene expression microarray were carried out. The expression status of miRNAs was confirmed by reverse transcriptase PCR. Furthermore, we investigated whether miRNA expression status in stromal tissue could influence the clinicopathologic factors. Results: Oncogenic miRNAs, including two miRNA clusters, miR-17-92a and miR-106b-25 cluster, were upregulated in cancer stromal tissues compared with normal stroma. Gene expression profiles from cDNA microarray analyses of the same stromal tissue samples revealed that putative targets of these miRNA clusters, predicted by Target Scan, such as TGFBR2, SMAD2, and BMP family genes, were significantly downregulated in cancer stromal tissue. Downregulated putative targets were also found to be involved in cytokine interaction and cellular adhesion. Importantly, expression of miR-25 and miR-92a in stromal tissues was associated with a variety of clinicopathologic factors. Conclusions: Oncogenic miRNAs were highly expressed in cancer stroma. -

75 2. INTRODUCTION Triple-Negative Breast Cancer (TNBC)
[Frontiers in Bioscience, Scholar, 11, 75-88, March 1, 2019] The persisting puzzle of racial disparity in triple negative breast cancer: looking through a new lens Chakravarthy Garlapati1, Shriya Joshi1, Bikram Sahoo1, Shobhna Kapoor2, Ritu Aneja1 1Department of Biology, Georgia State University, Atlanta, GA, USA, 2Department of Chemistry, Indian Institute of Technology Bombay, Powai, India TABLE OF CONTENTS 1. Abstract 2. Introduction 3. Dissecting the TNBC racially disparate burden 3.1. Does race influence TNBC onset and progression? 3.2. Tumor microenvironment in TNBC and racial disparity 3.3. Differential gene signatures and pathways in racially distinct TNBC 3.4. Our Perspective: Looking racial disparity through a new lens 4. Conclusion 5. Acknowledgement 6. References 1. ABSTRACT 2. INTRODUCTION Triple-negative breast cancer (TNBC) Triple-negative breast cancer (TNBC), is characterized by the absence of estrogen a subtype of breast cancer (BC), accounts for and progesterone receptors and absence 15-20% of all BC diagnoses in the US. It has of amplification of human epidermal growth been recognized that women of African descent factor receptor (HER2). This disease has no are twice as likely to develop TNBC than approved treatment with a poor prognosis women of European descent (1). As the name particularly in African-American (AA) as foretells, TNBCs lack estrogen, progesterone, compared to European-American (EA) and human epidermal growth factor receptors. patients. Gene ontology analysis showed Unfortunately, TNBCs are defined by what they specific gene pathways that are differentially “lack” rather than what they “have” and thus this regulated and gene signatures that are negative nomenclature provides no actionable differentially expressed in AA as compared to information on “druggable” targets. -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -
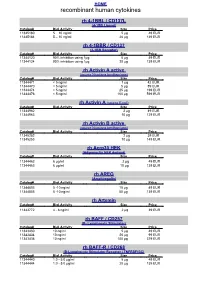
Recombinant Human Cytokines
HOME recombinant human cytokines rh 4-1BBL / CD137L (4-1BB Ligand) Catalog# Biol.Activity Size Price 11345180 5 – 10 ng/ml 5 µg 49 EUR 11345184 5 – 10 ng/ml 20 µg 139 EUR rh 4-1BBR / CD137 (4-1BB Receptor) Catalog# Biol.Activity Size Price 11344120 90% inhibition using 1µg 5 µg 49 EUR 11344124 90% inhibition using 1µg 20 µg 139 EUR rh Activin A active (source Nicotiana benthamiana) Catalog# Biol.Activity Size Price 11344471 < 5 ng/ml 1 µg 42 EUR 11344470 < 5 ng/ml 5 µg 89 EUR 11344474 < 5 ng/ml 25 µg 199 EUR 11344476 < 5 ng/ml 100 µg 599 EUR rh Activin A (source E.coli) Catalog# Biol.Activity Size Price 11344962 2 µg 49 EUR 11344963 10 µg 129 EUR rh Activin B active (source Nicotiana benthamiana) Catalog# Biol.Activity Size Price 11345252 2 µg 39 EUR 11345253 10 µg 149 EUR rh Acrp30 HEK (Adiponectin HEK derived) Catalog# Biol.Activity Size Price 11344462 6 µg/ml 2 µg 49 EUR 11344463 6 µg/ml 10 µg 139 EUR rh AREG (Amphiregulin) Catalog# Biol.Activity Size Price 11344803 5 -10 ng/ml 10 µg 49 EUR 11344805 5 -10 ng/ml 50 µg 139 EUR rh Artemin Catalog# Biol.Activity Size Price 11343772 4 - 8 ng/ml 2 µg 39 EUR rh BAFF / CD257 (B- Lymphocyte Stimulator) Catalog# Biol.Activity Size Price 11343430 10 ng/ml 5 µg 49 EUR 11343434 10 ng/ml 20 µg 99 EUR 11343436 10 ng/ml 100 µg 379 EUR rh BAFF-R / CD268 (B-Lymphocyte Stimulator Receptor / TNFRSF13C) Catalog# Biol.Activity Size Price 11344440 1.0 - 5.0 µg/ml 5 µg 49 EUR 11344444 1.0 - 5.0 µg/ml 20 µg 139 EUR HOME recombinant human cytokines rh BCA-1 (CXCL13) Catalog# Biol.Activity Size Price 11344180 -

Prevalent Homozygous Deletions of Type I Interferon and Defensin Genes in Human Cancers Associate with Immunotherapy Resistance
Author Manuscript Published OnlineFirst on April 4, 2018; DOI: 10.1158/1078-0432.CCR-17-3008 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Prevalent Homozygous Deletions of Type I Interferon and Defensin Genes in Human Cancers Associate with Immunotherapy Resistance Zhenqing Ye1; Haidong Dong2,3; Ying Li1; Tao Ma1,4; Haojie Huang4; Hon-Sing Leong4; Jeanette Eckel-Passow1; Jean-Pierre A. Kocher1; Han Liang5; LiguoWang1,4,* 1Division of Biomedical Statistics and Informatics, Department of Health Sciences, Mayo Clinic, Rochester, Minnesota 55905, United States of America 2Department of Immunology, College of Medicine, Mayo Clinic, Rochester, Minnesota 55905, United States of America 3Department of Urology, Mayo Clinic, Rochester, Minnesota 55905, United States of America 4Department of Biochemistry and Molecular Biology, Mayo Clinic, Rochester, Minnesota 55905, United States of America 5Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center, Houston, Texas 77030, United States of America Running title: Pervasive Deletion of Interferon/Defensin in Human Cancers Keywords: Homozygous deletion, Type-I interferon, Defensin, immunotherapy resistance, cancer * Correspondence to: Liguo Wang, Ph.D. Associate Professor Division of Biomedical Statistics and Informatics, Mayo Clinic 200 1st St SW Rochester, MN 55905, USA Phone: +1-507-284-8728 Fax: +1-507-284-0745 Email: [email protected] The authors declare no potential conflicts of interest. 1 Downloaded from clincancerres.aacrjournals.org on September 29, 2021. © 2018 American Association for Cancer Research. Author Manuscript Published OnlineFirst on April 4, 2018; DOI: 10.1158/1078-0432.CCR-17-3008 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited.