Redalyc.Assessment of Non-Cultured Aquatic Fungal Diversity from Differenthabitats in Mexico
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Checklist of Calicioid Lichens and Fungi for Genera with Members in Temperate Western North America Draft: 2012-03-13
Draft: 2012-03-13 Checklist of Calicioids – E. B. Peterson Checklist of Calicioid Lichens and Fungi For Genera with Members in Temperate Western North America Draft: 2012-03-13 by E. B. Peterson Calicium abietinum, EBP#4640 1 Draft: 2012-03-13 Checklist of Calicioids – E. B. Peterson Genera Acroscyphus Lév. Brucea Rikkinen Calicium Pers. Chaenotheca Th. Fr. Chaenothecopsis Vainio Coniocybe Ach. = Chaenotheca "Cryptocalicium" – potentially undescribed genus; taxonomic placement is not known but there are resemblances both to Mycocaliciales and Onygenales Cybebe Tibell = Chaenotheca Cyphelium Ach. Microcalicium Vainio Mycocalicium Vainio Phaeocalicium A.F.W. Schmidt Sclerophora Chevall. Sphinctrina Fr. Stenocybe (Nyl.) Körber Texosporium Nádv. ex Tibell & Hofsten Thelomma A. Massal. Tholurna Norman Additional genera are primarily tropical, such as Pyrgillus, Tylophoron About the Species lists Names in bold are believed to be currently valid names. Old synonyms are indented and listed with the current name following (additional synonyms can be found in Esslinger (2011). Names in quotes are nicknames for undescribed species. Names given within tildes (~) are published, but may not be validly published. Underlined species are included in the checklist for North America north of Mexico (Esslinger 2011). Names are given with authorities and original citation date where possible, followed by a colon. Additional citations are given after the colon, followed by a series of abbreviations for states and regions where known. States and provinces use the standard two-letter abbreviation. Regions include: NAm = North America; WNA = western North America (west of the continental divide); Klam = Klamath Region (my home territory). For those not known from North America, continental distribution may be given: SAm = South America; EUR = Europe; ASIA = Asia; Afr = Africa; Aus = Australia. -

Characterization of Two Undescribed Mucoralean Species with Specific
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 26 March 2018 doi:10.20944/preprints201803.0204.v1 1 Article 2 Characterization of Two Undescribed Mucoralean 3 Species with Specific Habitats in Korea 4 Seo Hee Lee, Thuong T. T. Nguyen and Hyang Burm Lee* 5 Division of Food Technology, Biotechnology and Agrochemistry, College of Agriculture and Life Sciences, 6 Chonnam National University, Gwangju 61186, Korea; [email protected] (S.H.L.); 7 [email protected] (T.T.T.N.) 8 * Correspondence: [email protected]; Tel.: +82-(0)62-530-2136 9 10 Abstract: The order Mucorales, the largest in number of species within the Mucoromycotina, 11 comprises typically fast-growing saprotrophic fungi. During a study of the fungal diversity of 12 undiscovered taxa in Korea, two mucoralean strains, CNUFC-GWD3-9 and CNUFC-EGF1-4, were 13 isolated from specific habitats including freshwater and fecal samples, respectively, in Korea. The 14 strains were analyzed both for morphology and phylogeny based on the internal transcribed 15 spacer (ITS) and large subunit (LSU) of 28S ribosomal DNA regions. On the basis of their 16 morphological characteristics and sequence analyses, isolates CNUFC-GWD3-9 and CNUFC- 17 EGF1-4 were confirmed to be Gilbertella persicaria and Pilobolus crystallinus, respectively.To the 18 best of our knowledge, there are no published literature records of these two genera in Korea. 19 Keywords: Gilbertella persicaria; Pilobolus crystallinus; mucoralean fungi; phylogeny; morphology; 20 undiscovered taxa 21 22 1. Introduction 23 Previously, taxa of the former phylum Zygomycota were distributed among the phylum 24 Glomeromycota and four subphyla incertae sedis, including Mucoromycotina, Kickxellomycotina, 25 Zoopagomycotina, and Entomophthoromycotina [1]. -

Supplementary Table S2: New Taxonomic Assignment of Sequences of Basal Fungal Lineages
Supplementary Table S2: New taxonomic assignment of sequences of basal fungal lineages. Fungal sequences were subjected to BLAST-N analysis and checked for their taxonomic placement in the eukaryotic guide-tree of the SILVA release 111. Sequences were classified depending on combined results from the methods mentioned above as well as literature searches. Accession Name New classification Clustering of the sequence in the Best BLAST-N hit number based on combined results eukaryotic guide tree of SILVA Name Accession number E.value Identity AB191431 Uncultured fungus Chytridiomycota Chytridiomycota Basidiobolus haptosporus AF113413.1 0.0 91 AB191432 Unculltured eukaryote Blastocladiomycota Blastocladiomycota Rhizophlyctis rosea NG_017175.1 0.0 91 AB252775 Uncultured eukaryote Chytridiomycota Chytridiomycota Blastocladiales sp. EF565163.1 0.0 91 AB252776 Uncultured eukaryote Fungi Nucletmycea_Fonticula Rhizophydium sp. AF164270.2 0.0 87 AB252777 Uncultured eukaryote Chytridiomycota Chytridiomycota Basidiobolus haptosporus AF113413.1 0.0 91 AB275063 Uncultured fungus Chytridiomycota Chytridiomycota Catenomyces sp. AY635830.1 0.0 90 AB275064 Uncultured fungus Chytridiomycota Chytridiomycota Endogone lactiflua DQ536471.1 0.0 91 AB433328 Nuclearia thermophila Nuclearia Nucletmycea_Nuclearia Nuclearia thermophila AB433328.1 0.0 100 AB468592 Uncultured fungus Basal clone group I Chytridiomycota Physoderma dulichii DQ536472.1 0.0 90 AB468593 Uncultured fungus Basal clone group I Chytridiomycota Physoderma dulichii DQ536472.1 0.0 91 AB468594 Uncultured -

Characterization of Fungi from Hypersaline Environments of Solar Salterns Using Morphological and Molecular Techniques
mycological research 110 (2006) 962–970 available at www.sciencedirect.com journal homepage: www.elsevier.com/locate/mycres Characterization of fungi from hypersaline environments of solar salterns using morphological and molecular techniques Sharon A. CANTRELLa,*, Lilliam CASILLAS-MARTI´NEZb, Marirosa MOLINAc aDepartment of Biology, School of Science and Technology, Universidad del Turabo, P. O. Box 3030, Gurabo, PR 00778, USA bBiology Department, University of Puerto Rico, CUH Station, Humacao, PR 00791, USA cEnvironmental Protection Agency, ORD, Athens, GA 30602 USA article info abstract Article history: The Cabo Rojo Solar Salterns located on the southwest coast of Puerto Rico are composed of Received 22 December 2005 two main ecosystems (i.e., salt ponds and microbial mats). Even though these locations are Received in revised form characterized by high solar radiation (mean light intensity of 39 mol photons mÀ2 dÀ1)they 24 April 2006 harbour a diverse microscopic life. We used morphological and molecular techniques to Accepted 1 June 2006 identify a series of halotolerant fungi. A total of 183 isolates and 36 species were cultured Published online 10 August 2006 in this study. From the water from the salt ponds, 86 isolates of 26 species were cultured. Corresponding Editor: John Dighton The halotolerant fungi isolated from water were: Cladosporium cladosporioides, nine Aspergillus sp., five Penicillium sp. and the black yeast Hortaea werneckii. A distinctive isolate with a blue Keywords: mycelium was cultured from the salt ponds, representing a new species of Periconia based Evaporation ponds on morphology and rDNA analysis. Forty-four isolates from eight species were cultured Halophilic fungi from the sediments around the salt ponds. -

Fungal Evolution: Major Ecological Adaptations and Evolutionary Transitions
Biol. Rev. (2019), pp. 000–000. 1 doi: 10.1111/brv.12510 Fungal evolution: major ecological adaptations and evolutionary transitions Miguel A. Naranjo-Ortiz1 and Toni Gabaldon´ 1,2,3∗ 1Department of Genomics and Bioinformatics, Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Dr. Aiguader 88, Barcelona 08003, Spain 2 Department of Experimental and Health Sciences, Universitat Pompeu Fabra (UPF), 08003 Barcelona, Spain 3ICREA, Pg. Lluís Companys 23, 08010 Barcelona, Spain ABSTRACT Fungi are a highly diverse group of heterotrophic eukaryotes characterized by the absence of phagotrophy and the presence of a chitinous cell wall. While unicellular fungi are far from rare, part of the evolutionary success of the group resides in their ability to grow indefinitely as a cylindrical multinucleated cell (hypha). Armed with these morphological traits and with an extremely high metabolical diversity, fungi have conquered numerous ecological niches and have shaped a whole world of interactions with other living organisms. Herein we survey the main evolutionary and ecological processes that have guided fungal diversity. We will first review the ecology and evolution of the zoosporic lineages and the process of terrestrialization, as one of the major evolutionary transitions in this kingdom. Several plausible scenarios have been proposed for fungal terrestralization and we here propose a new scenario, which considers icy environments as a transitory niche between water and emerged land. We then focus on exploring the main ecological relationships of Fungi with other organisms (other fungi, protozoans, animals and plants), as well as the origin of adaptations to certain specialized ecological niches within the group (lichens, black fungi and yeasts). -
Cuivre Bryophytes
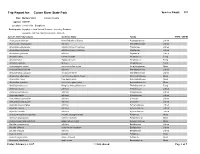
Trip Report for: Cuivre River State Park Species Count: 335 Date: Multiple Visits Lincoln County Agency: MODNR Location: Lincoln Hills - Bryophytes Participants: Bryophytes from Natural Resource Inventory Database Bryophyte List from NRIDS and Bruce Schuette Species Name (Synonym) Common Name Family COFC COFW Acarospora unknown Identified only to Genus Acarosporaceae Lichen Acrocordia megalospora a lichen Monoblastiaceae Lichen Amandinea dakotensis a button lichen (crustose) Physiaceae Lichen Amandinea polyspora a button lichen (crustose) Physiaceae Lichen Amandinea punctata a lichen Physiaceae Lichen Amanita citrina Citron Amanita Amanitaceae Fungi Amanita fulva Tawny Gresette Amanitaceae Fungi Amanita vaginata Grisette Amanitaceae Fungi Amblystegium varium common willow moss Amblystegiaceae Moss Anisomeridium biforme a lichen Monoblastiaceae Lichen Anisomeridium polypori a crustose lichen Monoblastiaceae Lichen Anomodon attenuatus common tree apron moss Anomodontaceae Moss Anomodon minor tree apron moss Anomodontaceae Moss Anomodon rostratus velvet tree apron moss Anomodontaceae Moss Armillaria tabescens Ringless Honey Mushroom Tricholomataceae Fungi Arthonia caesia a lichen Arthoniaceae Lichen Arthonia punctiformis a lichen Arthoniaceae Lichen Arthonia rubella a lichen Arthoniaceae Lichen Arthothelium spectabile a lichen Uncertain Lichen Arthothelium taediosum a lichen Uncertain Lichen Aspicilia caesiocinerea a lichen Hymeneliaceae Lichen Aspicilia cinerea a lichen Hymeneliaceae Lichen Aspicilia contorta a lichen Hymeneliaceae Lichen -

Depsidone Derivatives and a Cyclopeptide Produced by Marine Fungus Aspergillus Unguis Under Chemical Induction and by Its Plasma Induced Mutant
molecules Article Depsidone Derivatives and a Cyclopeptide Produced by Marine Fungus Aspergillus unguis under Chemical Induction and by Its Plasma Induced Mutant Wen-Cong Yang 1,† ID , Hai-Yan Bao 1,2,†, Ya-Yue Liu 1, Ying-Ying Nie 1,3, Jing-Ming Yang 1, Peng-Zhi Hong 1,3 and Yi Zhang 1,3,* ID 1 Research Institute for Marine Drugs and Nutrition, College of Food Science and Technology, Guangdong Provincial Key Laboratory of Aquatic Product Processing and Safety, Guangdong Province Engineering Laboratory for Marine Biological Products, Guangdong Ocean University, Zhanjiang 524088, China; [email protected] (W.-C.Y.); [email protected] (H.-Y.B.); [email protected] (Y.-Y.L.); [email protected] (Y.-Y.N.); [email protected] (J.-M.Y.); [email protected] (P.-Z.H.) 2 School of Environmental and Chemical Engineering, Dalian Jiaotong University, Dalian 116028, China 3 Shenzhen Institute of Guangdong Ocean University, Shenzhen 518120, China * Correspondence: [email protected]; Tel.: +86-759-2396046 † These authors contributed equally to this work. Academic Editor: Francesco Epifano Received: 17 August 2018; Accepted: 30 August 2018; Published: 3 September 2018 Abstract: A new depsidone derivative (1), aspergillusidone G, was isolated from a marine fungus Aspergillus unguis, together with eight known depsidones (2–9) and a cyclic peptide (10): agonodepside A (2), nornidulin (3), nidulin (4), aspergillusidone F (5), unguinol (6), aspergillusidone C(7), 2-chlorounguinol (8), aspergillusidone A (9), and unguisin A (10). Compounds 1–4 and 7–9 were obtained from the plasma induced mutant of this fungus, while 5, 6, and 10 were isolated from the original strain under chemical induction. -

Rhizaria, Cercozoa)
Protist, Vol. 166, 363–373, July 2015 http://www.elsevier.de/protis Published online date 28 May 2015 ORIGINAL PAPER Molecular Phylogeny of the Widely Distributed Marine Protists, Phaeodaria (Rhizaria, Cercozoa) a,1 a a b Yasuhide Nakamura , Ichiro Imai , Atsushi Yamaguchi , Akihiro Tuji , c d Fabrice Not , and Noritoshi Suzuki a Plankton Laboratory, Graduate School of Fisheries Sciences, Hokkaido University, Hakodate, Hokkaido 041–8611, Japan b Department of Botany, National Museum of Nature and Science, Tsukuba 305–0005, Japan c CNRS, UMR 7144 & Université Pierre et Marie Curie, Station Biologique de Roscoff, Equipe EPPO - Evolution du Plancton et PaléoOcéans, Place Georges Teissier, 29682 Roscoff, France d Institute of Geology and Paleontology, Graduate School of Science, Tohoku University, Sendai 980–8578, Japan Submitted January 1, 2015; Accepted May 19, 2015 Monitoring Editor: David Moreira Phaeodarians are a group of widely distributed marine cercozoans. These plankton organisms can exhibit a large biomass in the environment and are supposed to play an important role in marine ecosystems and in material cycles in the ocean. Accurate knowledge of phaeodarian classification is thus necessary to better understand marine biology, however, phylogenetic information on Phaeodaria is limited. The present study analyzed 18S rDNA sequences encompassing all existing phaeodarian orders, to clarify their phylogenetic relationships and improve their taxonomic classification. The mono- phyly of Phaeodaria was confirmed and strongly supported by phylogenetic analysis with a larger data set than in previous studies. The phaeodarian clade contained 11 subclades which generally did not correspond to the families and orders of the current classification system. Two families (Challengeri- idae and Aulosphaeridae) and two orders (Phaeogromida and Phaeocalpida) are possibly polyphyletic or paraphyletic, and consequently the classification needs to be revised at both the family and order levels by integrative taxonomy approaches. -

For Review Only 377 Algomyces Stechlinensis Clustered Together with Environmental Clones from a Eutrophic 378 Lake in France (Jobard Et Al
Journal of Eukaryotic Microbiology Page 18 of 43 1 Running head: Parasitic chytrids of volvocacean algae. 2 3 Title: Diversity and Hidden Host Specificity of Chytrids infecting Colonial 4 Volvocacean Algae. 5 Authors: Silke Van den Wyngaerta, Keilor Rojas-Jimeneza,b, Kensuke Setoc, Maiko Kagamic, 6 Hans-Peter Grossarta,d 7 a Department of ExperimentalFor Limnology, Review Leibniz-Institute Only of Freshwater Ecology and Inland 8 Fisheries, Alte Fischerhuette 2, D-16775 Stechlin, Germany 9 b Universidad Latina de Costa Rica, Campus San Pedro, Apdo. 10138-1000, San Jose, Costa Rica 10 c Department of Environmental Sciences, Faculty of Science, Toho University, Funabashi, Chiba, 11 Japan 12 d Institute of Biochemistry and Biology, Potsdam University, Maulbeerallee 2, 14476 Potsdam, 13 Germany 14 15 Corresponding Author: 16 Silke Van den Wyngaert, Department of Experimental Limnology, Leibniz-Institute of 17 Freshwater Ecology and Inland Fisheries, Alte Fischerhuette 2, D-16775 Stechlin, Germany 18 Telephone number: +49 33082 69972; Fax number: +49 33082 69917; e-mail: [email protected], 19 [email protected] 20 21 22 23 1 Page 19 of 43 Journal of Eukaryotic Microbiology 24 ABSTRACT 25 Chytrids are zoosporic fungi that play an important, but yet understudied, ecological role in 26 aquatic ecosystems. Many chytrid species have been morphologically described as parasites on 27 phytoplankton. However, the majority of them have rarely been isolated and lack DNA sequence 28 data. In this study we isolated and cultivated three parasitic chytrids, infecting a common 29 volvocacean host species, Yamagishiella unicocca. In order to identify the chytrids, we 30 characterized morphology and life cycle, and analyzed phylogenetic relationships based on 18S 31 and 28S rDNA genes. -

Phaeocalicium (Mycocaliciaceae, Ascomycetes) in Northern Europe
Ann. Bot. Fennici 33: 205–221 ISSN 0003-3847 Helsinki 30 October 1996 © Finnish Zoological and Botanical Publishing Board 1996 Phaeocalicium (Mycocaliciaceae, Ascomycetes) in Northern Europe Leif Tibell Tibell, L., Department of Systematic Botany, Villavägen 6, S-752 36 Uppsala, Sweden Received 7 February 1996, accepted 28 May 1996 The taxonomy, distribution and ecology of eight species of Phaeocalicium A. F. W. Schmidt (Mycocaliciaceae, Ascomycetes) occurring in the Nordic countries and Greenland are de- scribed. They are parasitic or saprophytic mainly on thin twigs of trees and shrubs such as Alnus, Betula, Populus and Salix and are often quite host-specific. A key to the species is supplied. Two new species, Ph. boreale Tibell and Ph. flabelliforme Tibell are described. A lectotype is selected for Calicium praecedens Nyl., Mycocalicium pusiolum (Ach.) Räsänen var. macrospora Räsänen and Stenocybe tremulicola Norrl. ex Nyl., and a neotype is se- lected for Phaeocalicium populneum (Brond. ex Duby) A. F. W. Schmidt. The new combi- nation Ph. tremulicola (Norrl. ex Nyl.) Tibell is proposed. Key words: Ascomycetes, Caliciales, Mycocaliciaceae, Phaeocalicium, taxonomy INTRODUCTION cybe, as conceived by Schmidt (1970), are unsatis- factory and in need of revision. Thus as an example The ascomycete genus Phaeocalicium belongs to spore septation does not seem to be consistent evi- Mycocaliciaceae in Caliciales s. lat. (Tibell 1984). dence for generic delimitation. In the present paper The North European species occur as saprobes and/ a group of similar and presumably closely related or weak parasites mostly on thin, decaying branches species has provisionally been accomodated in of deciduous trees or shrubs. -

Preliminary Classification of Leotiomycetes
Mycosphere 10(1): 310–489 (2019) www.mycosphere.org ISSN 2077 7019 Article Doi 10.5943/mycosphere/10/1/7 Preliminary classification of Leotiomycetes Ekanayaka AH1,2, Hyde KD1,2, Gentekaki E2,3, McKenzie EHC4, Zhao Q1,*, Bulgakov TS5, Camporesi E6,7 1Key Laboratory for Plant Diversity and Biogeography of East Asia, Kunming Institute of Botany, Chinese Academy of Sciences, Kunming 650201, Yunnan, China 2Center of Excellence in Fungal Research, Mae Fah Luang University, Chiang Rai, 57100, Thailand 3School of Science, Mae Fah Luang University, Chiang Rai, 57100, Thailand 4Landcare Research Manaaki Whenua, Private Bag 92170, Auckland, New Zealand 5Russian Research Institute of Floriculture and Subtropical Crops, 2/28 Yana Fabritsiusa Street, Sochi 354002, Krasnodar region, Russia 6A.M.B. Gruppo Micologico Forlivese “Antonio Cicognani”, Via Roma 18, Forlì, Italy. 7A.M.B. Circolo Micologico “Giovanni Carini”, C.P. 314 Brescia, Italy. Ekanayaka AH, Hyde KD, Gentekaki E, McKenzie EHC, Zhao Q, Bulgakov TS, Camporesi E 2019 – Preliminary classification of Leotiomycetes. Mycosphere 10(1), 310–489, Doi 10.5943/mycosphere/10/1/7 Abstract Leotiomycetes is regarded as the inoperculate class of discomycetes within the phylum Ascomycota. Taxa are mainly characterized by asci with a simple pore blueing in Melzer’s reagent, although some taxa have lost this character. The monophyly of this class has been verified in several recent molecular studies. However, circumscription of the orders, families and generic level delimitation are still unsettled. This paper provides a modified backbone tree for the class Leotiomycetes based on phylogenetic analysis of combined ITS, LSU, SSU, TEF, and RPB2 loci. In the phylogenetic analysis, Leotiomycetes separates into 19 clades, which can be recognized as orders and order-level clades. -

Epiphytic Seed Microbiomes of Wheat, Canola, and Lentil
EPIPHYTIC SEED MICROBIOMES OF WHEAT, CANOLA, AND LENTIL A Thesis Submitted to the College of Graduate and Postdoctoral Studies In Partial Fulfillment of the Requirements For the Degree of Doctor of Philosophy In the Department of Food and Bioproduct Sciences University of Saskatchewan Saskatoon By Zayda Piedad Morales Moreira © Copyright Zayda Piedad Morales Moreira, June, 2021. All rights reserved. Unless otherwise noted, copyright of the material in this thesis belongs to the author PERMISSION TO USE In presenting this thesis in partial fulfilment of the requirements for a Postgraduate degree from the University of Saskatchewan, I agree that the Libraries of this University may make it freely available for inspection. I further agree that permission for copying of this thesis in any manner, in whole or in part, for scholarly purposes may be granted by the professor or professors who supervised my thesis work or, in their absence, by the Head of the Department or the Dean of the College in which my thesis work was done. It is understood that any copying, publication, or use of this thesis or parts thereof for financial gain shall not be allowed without my written permission. It is also understood that due recognition shall be given to me and to the University of Saskatchewan in any scholarly use which may be made of any material in my thesis. Requests for permission to copy or to make other use of material in this thesis in whole or part should be addressed to: Head of the Department of Food and Bioproduct Sciences University of Saskatchewan 51 Campus Drive University of Saskatchewan Saskatoon, Saskatchewan, S7N 5A8 Canada OR Dean of the College of Graduate and Postdoctoral Studies University of Saskatchewan 107 Administration Place Saskatoon, Saskatchewan S7N 5A2 Canada i ABSTRACT Microorganisms are found colonizing all plant organs including seeds.