Coordinating Cell Cycle Exit and Differentiation in the Mammalian
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Title of the Dissertation
UNIVERSITY OF CALIFORNIA SAN DIEGO Novel network-based integrated analyses of multi-omics data reveal new insights into CD8+ T cell differentiation and mouse embryogenesis A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioinformatics and Systems Biology by Kai Zhang Committee in charge: Professor Wei Wang, Chair Professor Pavel Arkadjevich Pevzner, Co-Chair Professor Vineet Bafna Professor Cornelis Murre Professor Bing Ren 2018 Copyright Kai Zhang, 2018 All rights reserved. The dissertation of Kai Zhang is approved, and it is accept- able in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California San Diego 2018 iii EPIGRAPH The only true wisdom is in knowing you know nothing. —Socrates iv TABLE OF CONTENTS Signature Page ....................................... iii Epigraph ........................................... iv Table of Contents ...................................... v List of Figures ........................................ viii List of Tables ........................................ ix Acknowledgements ..................................... x Vita ............................................. xi Abstract of the Dissertation ................................. xii Chapter 1 General introduction ............................ 1 1.1 The applications of graph theory in bioinformatics ......... 1 1.2 Leveraging graphs to conduct integrated analyses .......... 4 1.3 References .............................. 6 Chapter 2 Systematic -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid
The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid Leukemia Cells Supported Only by Exogenous Cytokines Is Associated with a Patient Subset with Adverse Outcome Annette K. Brenner, Elise Aasebø, Maria Hernandez-Valladares, Frode Selheim, Frode Berven, Ida-Sofie Grønningsæter, Sushma Bartaula-Brevik and Øystein Bruserud Supplementary Material S2 of S31 Table S1. Detailed information about the 68 AML patients included in the study. # of blasts Viability Proliferation Cytokine Viable cells Change in ID Gender Age Etiology FAB Cytogenetics Mutations CD34 Colonies (109/L) (%) 48 h (cpm) secretion (106) 5 weeks phenotype 1 M 42 de novo 241 M2 normal Flt3 pos 31.0 3848 low 0.24 7 yes 2 M 82 MF 12.4 M2 t(9;22) wt pos 81.6 74,686 low 1.43 969 yes 3 F 49 CML/relapse 149 M2 complex n.d. pos 26.2 3472 low 0.08 n.d. no 4 M 33 de novo 62.0 M2 normal wt pos 67.5 6206 low 0.08 6.5 no 5 M 71 relapse 91.0 M4 normal NPM1 pos 63.5 21,331 low 0.17 n.d. yes 6 M 83 de novo 109 M1 n.d. wt pos 19.1 8764 low 1.65 693 no 7 F 77 MDS 26.4 M1 normal wt pos 89.4 53,799 high 3.43 2746 no 8 M 46 de novo 26.9 M1 normal NPM1 n.d. n.d. 3472 low 1.56 n.d. no 9 M 68 MF 50.8 M4 normal D835 pos 69.4 1640 low 0.08 n.d. -

Predicting Coupling Probabilities of G-Protein Coupled Receptors Gurdeep Singh1,2,†, Asuka Inoue3,*,†, J
Published online 30 May 2019 Nucleic Acids Research, 2019, Vol. 47, Web Server issue W395–W401 doi: 10.1093/nar/gkz392 PRECOG: PREdicting COupling probabilities of G-protein coupled receptors Gurdeep Singh1,2,†, Asuka Inoue3,*,†, J. Silvio Gutkind4, Robert B. Russell1,2,* and Francesco Raimondi1,2,* 1CellNetworks, Bioquant, Heidelberg University, Im Neuenheimer Feld 267, 69120 Heidelberg, Germany, 2Biochemie Zentrum Heidelberg (BZH), Heidelberg University, Im Neuenheimer Feld 328, 69120 Heidelberg, Germany, 3Graduate School of Pharmaceutical Sciences, Tohoku University, Sendai, Miyagi 980-8578, Japan and 4Department of Pharmacology and Moores Cancer Center, University of California, San Diego, La Jolla, CA 92093, USA Received February 10, 2019; Revised April 13, 2019; Editorial Decision April 24, 2019; Accepted May 01, 2019 ABSTRACT great use in tinkering with signalling pathways in living sys- tems (5). G-protein coupled receptors (GPCRs) control multi- Ligand binding to GPCRs induces conformational ple physiological states by transducing a multitude changes that lead to binding and activation of G-proteins of extracellular stimuli into the cell via coupling to situated on the inner cell membrane. Most of mammalian intra-cellular heterotrimeric G-proteins. Deciphering GPCRs couple with more than one G-protein giving each which G-proteins couple to each of the hundreds receptor a distinct coupling profile (6) and thus specific of GPCRs present in a typical eukaryotic organism downstream cellular responses. Determining these coupling is therefore critical to understand signalling. Here, profiles is critical to understand GPCR biology and phar- we present PRECOG (precog.russelllab.org): a web- macology. Despite decades of research and hundreds of ob- server for predicting GPCR coupling, which allows served interactions, coupling information is still missing for users to: (i) predict coupling probabilities for GPCRs many receptors and sequence determinants of coupling- specificity are still largely unknown. -

VSX1 Mutational Analysis in a Series of Italian Patients Affected by Keratoconus: Detection of a Novel Mutation
VSX1 Mutational Analysis in a Series of Italian Patients Affected by Keratoconus: Detection of a Novel Mutation Luigi Bisceglia,1 Marilena Ciaschetti,2 Patrizia De Bonis,1 Pablo Alberto Perafan Campo,2 Costantina Pizzicoli,3 Costanza Scala,1 Michele Grifa,1 Pio Ciavarella,1 Nicola Delle Noci,3 Filippo Vaira,4 Claudio Macaluso,5 and Leopoldo Zelante1 PURPOSE. Keratoconus is a noninflammatory corneal disorder Most cases of keratoconus appear to be sporadic, but a that is clinically and genetically heterogeneous. Mutations in positive family history has been documented in 6% to 10% of the VSX1 (visual system homeobox 1) gene have been identi- patients.5 Both recessive and dominant patterns of inheritance fied for two distinct, inherited corneal dystrophies: posterior have been described.6–8 Autosomal dominant inheritance has polymorphous corneal dystrophy and keratoconus. To evalu- more frequently been reported in families, showing incom- ate the possible role of the VSX1 gene in a series of Italian plete penetrance and variable expressivity. Subtle videokerato- patients, 80 keratoconus-affected subjects were screened for graphic anomalies have been reported among relatives of pa- mutations. tients with keratoconus, allowing the detection of low- ETHODS expressivity forms of keratoconus, usually referred to as M . The diagnosis of keratoconus was made on the basis 9–11 of clinical examination and corneal topography. The whole subclinical or forme fruste keratoconus. Multifactorial in- heritance and a major gene model have also been pro- coding region and the exon–intron junctions of the VSX1 gene 12,13 were analyzed by direct sequencing. posed. In some cases nongenetic causes have been postu- lated, such as eye rubbing or rigid contact lens wear, which RESULTS. -

A Flexible Microfluidic System for Single-Cell Transcriptome Profiling
www.nature.com/scientificreports OPEN A fexible microfuidic system for single‑cell transcriptome profling elucidates phased transcriptional regulators of cell cycle Karen Davey1,7, Daniel Wong2,7, Filip Konopacki2, Eugene Kwa1, Tony Ly3, Heike Fiegler2 & Christopher R. Sibley 1,4,5,6* Single cell transcriptome profling has emerged as a breakthrough technology for the high‑resolution understanding of complex cellular systems. Here we report a fexible, cost‑efective and user‑ friendly droplet‑based microfuidics system, called the Nadia Instrument, that can allow 3′ mRNA capture of ~ 50,000 single cells or individual nuclei in a single run. The precise pressure‑based system demonstrates highly reproducible droplet size, low doublet rates and high mRNA capture efciencies that compare favorably in the feld. Moreover, when combined with the Nadia Innovate, the system can be transformed into an adaptable setup that enables use of diferent bufers and barcoded bead confgurations to facilitate diverse applications. Finally, by 3′ mRNA profling asynchronous human and mouse cells at diferent phases of the cell cycle, we demonstrate the system’s ability to readily distinguish distinct cell populations and infer underlying transcriptional regulatory networks. Notably this provided supportive evidence for multiple transcription factors that had little or no known link to the cell cycle (e.g. DRAP1, ZKSCAN1 and CEBPZ). In summary, the Nadia platform represents a promising and fexible technology for future transcriptomic studies, and other related applications, at cell resolution. Single cell transcriptome profling has recently emerged as a breakthrough technology for understanding how cellular heterogeneity contributes to complex biological systems. Indeed, cultured cells, microorganisms, biopsies, blood and other tissues can be rapidly profled for quantifcation of gene expression at cell resolution. -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -
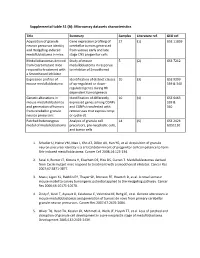
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

Supplementary Table 1. Pain and PTSS Associated Genes (N = 604
Supplementary Table 1. Pain and PTSS associated genes (n = 604) compiled from three established pain gene databases (PainNetworks,[61] Algynomics,[52] and PainGenes[42]) and one PTSS gene database (PTSDgene[88]). These genes were used in in silico analyses aimed at identifying miRNA that are predicted to preferentially target this list genes vs. a random set of genes (of the same length). ABCC4 ACE2 ACHE ACPP ACSL1 ADAM11 ADAMTS5 ADCY5 ADCYAP1 ADCYAP1R1 ADM ADORA2A ADORA2B ADRA1A ADRA1B ADRA1D ADRA2A ADRA2C ADRB1 ADRB2 ADRB3 ADRBK1 ADRBK2 AGTR2 ALOX12 ANO1 ANO3 APOE APP AQP1 AQP4 ARL5B ARRB1 ARRB2 ASIC1 ASIC2 ATF1 ATF3 ATF6B ATP1A1 ATP1B3 ATP2B1 ATP6V1A ATP6V1B2 ATP6V1G2 AVPR1A AVPR2 BACE1 BAMBI BDKRB2 BDNF BHLHE22 BTG2 CA8 CACNA1A CACNA1B CACNA1C CACNA1E CACNA1G CACNA1H CACNA2D1 CACNA2D2 CACNA2D3 CACNB3 CACNG2 CALB1 CALCRL CALM2 CAMK2A CAMK2B CAMK4 CAT CCK CCKAR CCKBR CCL2 CCL3 CCL4 CCR1 CCR7 CD274 CD38 CD4 CD40 CDH11 CDK5 CDK5R1 CDKN1A CHRM1 CHRM2 CHRM3 CHRM5 CHRNA5 CHRNA7 CHRNB2 CHRNB4 CHUK CLCN6 CLOCK CNGA3 CNR1 COL11A2 COL9A1 COMT COQ10A CPN1 CPS1 CREB1 CRH CRHBP CRHR1 CRHR2 CRIP2 CRYAA CSF2 CSF2RB CSK CSMD1 CSNK1A1 CSNK1E CTSB CTSS CX3CL1 CXCL5 CXCR3 CXCR4 CYBB CYP19A1 CYP2D6 CYP3A4 DAB1 DAO DBH DBI DICER1 DISC1 DLG2 DLG4 DPCR1 DPP4 DRD1 DRD2 DRD3 DRD4 DRGX DTNBP1 DUSP6 ECE2 EDN1 EDNRA EDNRB EFNB1 EFNB2 EGF EGFR EGR1 EGR3 ENPP2 EPB41L2 EPHB1 EPHB2 EPHB3 EPHB4 EPHB6 EPHX2 ERBB2 ERBB4 EREG ESR1 ESR2 ETV1 EZR F2R F2RL1 F2RL2 FAAH FAM19A4 FGF2 FKBP5 FLOT1 FMR1 FOS FOSB FOSL2 FOXN1 FRMPD4 FSTL1 FYN GABARAPL1 GABBR1 GABBR2 GABRA2 GABRA4 -

Mir-205: a Potential Biomedicine for Cancer Therapy
cells Review miR-205: A Potential Biomedicine for Cancer Therapy Neeraj Chauhan 1,2 , Anupam Dhasmana 1,2, Meena Jaggi 1,2, Subhash C. Chauhan 1,2 and Murali M. Yallapu 1,2,* 1 Department of Immunology and Microbiology, School of Medicine, University of Texas Rio Grande Valley, McAllen, TX 78504, USA; [email protected] (N.C.); [email protected] (A.D.); [email protected] (M.J.); [email protected] (S.C.C.) 2 South Texas Center of Excellence in Cancer Research, School of Medicine, University of Texas Rio Grande Valley, McAllen, TX 78504, USA * Correspondence: [email protected]; Tel.: +1-(956)-296-1734 Received: 3 June 2020; Accepted: 21 August 2020; Published: 25 August 2020 Abstract: microRNAs (miRNAs) are a class of small non-coding RNAs that regulate the expression of their target mRNAs post transcriptionally. miRNAs are known to regulate not just a gene but the whole gene network (signaling pathways). Accumulating evidence(s) suggests that miRNAs can work either as oncogenes or tumor suppressors, but some miRNAs have a dual nature since they can act as both. miRNA 205 (miR-205) is one such highly conserved miRNA that can act as both, oncomiRNA and tumor suppressor. However, most reports confirm its emerging role as a tumor suppressor in many cancers. This review focuses on the downregulated expression of miR-205 and discusses its dysregulation in breast, prostate, skin, liver, gliomas, pancreatic, colorectal and renal cancers. This review also confers its role in tumor initiation, progression, cell proliferation, epithelial to mesenchymal transition, and tumor metastasis. -

Molecular Markers and Potential Therapeutic Targets in Non-WNT/Non
Menyhárt et al. Journal of Hematology & Oncology (2019) 12:29 https://doi.org/10.1186/s13045-019-0712-y REVIEW Open Access Molecular markers and potential therapeutic targets in non-WNT/non-SHH (group 3 and group 4) medulloblastomas Otília Menyhárt1,2, Felice Giangaspero3,4 and Balázs Győrffy1,2* Abstract Childhood medulloblastomas (MB) are heterogeneous and are divided into four molecular subgroups. The provisional non-wingless-activated (WNT)/non-sonic hedgehog-activated (SHH) category combining group 3 and group 4 represents over two thirds of all MBs, coupled with the highest rates of metastases and least understood pathology. The molecular era expanded our knowledge about molecular aberrations involved in MB tumorigenesis, and here, we review processes leading to non-WNT/non-SHH MB formations. The heterogeneous group 3 and group 4 MBs frequently harbor rare individual genetic alterations, yet the emerging profiles suggest that infrequent events converge on common, potentially targetable signaling pathways. A mutual theme is the altered epigenetic regulation, and in vitro approaches targeting epigenetic machinery are promising. Growing evidence indicates the presence of an intermediate, mixed signature group along group 3 and group 4, and future clarifications are imperative for concordant classification, as misidentifying patient samples has serious implications for therapy and clinical trials. To subdue the high MB mortality, we need to discern mechanisms of disease spread and recurrence. Current preclinical models do not represent the full scale of group 3 and group 4 heterogeneity: all of existing group 3 cell lines are MYC-amplified and most mouse models resemble MYC-activated MBs. Clinical samples provide a wealth of information about the genetic divergence between primary tumors and metastatic clones, but recurrent MBs are rarely resected.