[C1C2], Internal
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supporting Information
Supporting Information Koyanagi et al. 10.1073/pnas.0806215105 SI Materials and Methods CNGB, NM137763; human CNGB1, NM001297; human Amino Acid Sequences. The accession numbers of amino acid CNGB3, NM 019098; box jellyfish CNG, AB435552; fruit fly sequences used for analyses are as follows: Box jellyfish opsin, CNG4, NM 167441; fruit fly CNG3, NM 137871; human AB435549; sea anemone opsin, BR000662; hydra opsin, Con- CNGA4, NM 001037329; fruit fly CNGA, NM 057768; human CNGA1, NM000087; human CNGA2, NM005140; human tig39347:487820–488855 (http://hydrazome.metazome.net); hy- drozaon jellyfish opsin, AB332435; human encephalopsin, CNGA3, NM 001298. AF140242; ragworm c-opsin, AY692353; human rhodopsin, Western Blot Analysis. The proteins extracted from half a rhopalia U49742; human blue, M13299; human red, Z68193; lamprey were separated by 12% SDS/PAGE, transferred onto a PVDF parapinopsin, AB116380; lizard parietopsin, DQ100320; am- membrane, and incubated with 1:500 diluted anti-box jellyfish phioxus peropsin, AB050610; human peropsin, AF012270; hu- opsin antiserum. Visualization was carried out by the ABC man rgr, U15790; squid retinochrome, X57143; human melan- method (Vectastain) and by staining with 3,3Ј-diaminobenzidine opsin, AF147788; amphioxus melanopsin, AB205400; squid (Sigma). rhodopsin, X70498; fruit fly Rh1, K02315; human neuropsin, AY377391; amphioxus rhodopsin, AB050606; scallop Go- In Situ Hybridization. Digoxigenin-labeled antisense and sense RNA rhodopsin, AB006455; human muscarinic acetylcholine receptor probes for the box jellyfish opsin were synthesized by using the DIG M1 (CHRM1), NM000738; human melatonin receptor 1A RNA labeling kit (Roche). Sections were pretreated with protein- (MTNR1A), NM005958; Arabidopsis AKT1, U06745; Arabi- ase K and hybridized with each RNA probe. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Expression of Melatonin Receptors 1A/1B, Calmodulin and Estrogen Receptor 2 in Deep Paravertebral Muscles Revisited
MOLECULAR MEDICINE REPORTS 14: 5719-5724, 2016 Etiopathogenesis of adolescent idiopathic scoliosis: Expression of melatonin receptors 1A/1B, calmodulin and estrogen receptor 2 in deep paravertebral muscles revisited JOSEF ZAMECNIK1*, LENKA KRSKOVA1*, JAROMIR HACEK1, IVANA STETKAROVA2 and MARTIN KRBEC3 1Department of Pathology and Molecular Medicine, 2nd Faculty of Medicine, Charles University in Prague and University Hospital Motol, 15006 Prague; 2Department of Neurology, Charles University; 3Department of Orthopedics and Traumatology, 3rd Faculty of Medicine, Charles University in Prague and University Hospital Královské Vinohrady, 10034 Prague, Czech Republic Received October 28, 2015; Accepted October 11, 2016 DOI: 10.3892/mmr.2016.5927 Abstract. The pathogenesis of adolescent idiopathic scoliosis addition, no difference in expression was detected between the (AIS), including the associated local changes in deep para- patients with AIS and the controls. With regards to MTNR1A vertebral muscles, is poorly understood. The asymmetric and MTNR1B, their expression was very weak in paravertebral expression of several molecules involved in the melatonin muscles, and in the majority of cases their expression could signaling pathway, including melatonin receptors 1A/1B not be detected by repeated RT-qPCR analysis. Therefore, (MTNR1A/MTNR1B), estrogen receptor 2 (ESR2) and these data do not support the previously suggested role of the calmodulin (CALM1), has previously been suggested to be asymmetric expression of molecules involved in the melatonin associated with AIS. However, this hypothesis is based on signaling pathway in deep paravertebral muscles in the patho- single studies in which the data were obtained by different genesis of AIS. methodological approaches. Therefore, to evaluate the symmetry of the mRNA expression levels of these molecules, Introduction 18 patients with AIS and 10 non-scoliotic controls were enrolled in the present study. -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -
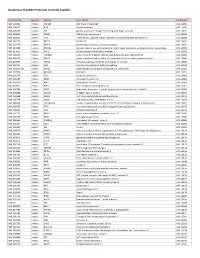
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) -

Genetic Variations of Melatonin Receptor Type 1A Are Associated with the Clinicopathologic Development of Urothelial Cell Carcin
Int. J. Med. Sci. 2017, Vol. 14 1130 Ivyspring International Publisher International Journal of Medical Sciences 2017; 14(11): 1130-1135. doi: 10.7150/ijms.20629 Research Paper Genetic Variations of Melatonin Receptor Type 1A are Associated with the Clinicopathologic Development of Urothelial Cell Carcinoma Yung-Wei Lin1, 2, Shian-Shiang Wang3, 4, 5, Yu-Ching Wen2, 6, Min-Che Tung1, 7, Liang-Ming Lee2, 6, Shun-Fa Yang5, 8, Ming-Hsien Chien1, 9 1. Graduate Institute of Clinical Medicine, Taipei Medical University, Taipei, Taiwan; 2. Department of Urology, Wan Fang Hospital, Taipei Medical University, Taipei, Taiwan; 3. Division of Urology, Department of Surgery, Taichung Veterans General Hospital, Taichung, Taiwan; 4. School of Medicine, Chung Shan Medical University, Taichung, Taiwan; 5. Institute of Medicine, Chung Shan Medical University, Taichung, Taiwan; 6. Department of Urology, School of Medicine, College of Medicine, Taipei Medical University, Taipei, Taiwan; 7. Department of Surgery, Tungs' Taichung Metro Harbor Hospital, Taichung, Taiwan; 8. Department of Medical Research, Chung Shan Medical University Hospital, Taichung, Taiwan; 9. Department of Medical Education and Research, Wan Fang Hospital, Taipei Medical University, Taipei, Taiwan. Corresponding authors: Ming-Hsien Chien, PhD, Graduate Institute of Clinical Medicine, Taipei Medical University, 250 Wu-Hsing Street, Taipei 11031, Taiwan; Phone: 886-2-27361661, ext. 3237; Fax: 886-2-27390500; E-mail: [email protected] or Shun-Fa Yang, PhD, Institute of Medicine, Chung Shan Medical University, 110 Chien-Kuo N. Road, Section 1, Taichung 402, Taiwan; Phone: 886-4-2473959, ext. 34253; Fax: 886-4-24723229; E-mail: [email protected] © Ivyspring International Publisher. -

Neurotransmission Alphabetical 28/7/05 15:52 Page 1
neuroscience - Neurotransmission alphabetical 28/7/05 15:52 Page 1 Abcam’s range of Neuroscience receptor, channel and ligand antibodies includes over 320 tried and tested products www.abcam.com Neurotransmission - receptor and channel signaling www.abcam.com GPCRs TYROSINE KINASES ION CHANNELS Neurotrophins rin u BDNF rt NGF u NT-3 Glutamate GABA ACh GDNF Ne Artemin Persepin NT-4/5 Glutamate Anandamide ATP Glycine ␥ 2-AG 1 2 3 4 / ␣ ␣ ␣ ␣ R / u A GFR GFR GFR GFR NTR mGL X1, 2, 3 CB1/2 2 EphA/B* Ephrin RET TrkA/B/C p75 AMPA/KainateNMDAR P GABA GlyR nAChR IP3 released Lyn Ephexin Homer DAG Adenylate Fyn PI3K Gr Src PSD95 Rapsin i PD2RGS3 Shc Syntrophin b K PLC Gi/q cyclase Gi 4 NcK PI3 Shc + 2+ 2+ PI3K PLC ␥ Na Ca Ca P PI3K Grb2 PLC/␥ PKC nNOS ␥ FAK PiP3/4 SOS MEK CaM Cl- Cl- ER NFkB PDEI PKA Rac RhoA RasGAP Src PTEN PKC Ras GTPase ERK CaMK 2+ PKK2 Calcineurin Ca released ROCK ERK FRS2 AKT/PKB Raf JNK nNOS SynGAP Grb2 actin MEK 1/2 P Ser133 Inhibition Cl- Src SOS Cell survival CREB of signaling RAS P CREB ERK 1/2 P P MAPK GSK3 ELK RSK NFB CREB CREB gene Long term Cytoskeleton transcription synaptic dynamics, neurite ASK1 Bad CRE plasticity (LTP) P P extension CREB CREB Short term MKK3/6 Apoptosis plasticity Others Other GPCRs - ARA9 ACh (Muscarinic): M1, M2, M3, M4 CRE p38MAPK c-fos Potassium - ASIC, ASIC3 Adrenalin: Alpha1b, 1c, 1d - KChIP2 - DDR 1, 2 ATP: P2Y MSK1 c-j c-fos gene - KCNQ 3, 5 - ENSA Dopamine: D1, D4 u transcription - Kv beta 2 - GJB1 n GABA: GABAB receptor - Kv 1.2 - SLC31A1 ␦ P P Opioid: µ, , CREB CREB AP1 - Kv -

Initial, Transient, and Specific Interaction Between G Protein
Sato T. et al. Medical Research Archives, vol. 6, issue 9, September 2018 Page 1 of 25 ARTICLE Initial, transient, and specific interaction between G protein-coupled receptor and target G protein in parallel signal processing: a case of olfactory discrimination of cancer-induced odors Takaaki Sato1, Mutsumi Matsukawa2, Yoichi Mizutani3, Toshio Iijima4, Hiroyoshi Matsumura5 Authors’ affiliations: 1 Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology, Osaka, Japan 2 Division of Anatomical Science, Department of Functional Morphology, Nihon University School of Medicine, Tokyo, Japan 3 Department of Medical Engineering, Faculty of Health Science, Aino University, Osaka, Japan 4 Graduate School of Life Sciences, Tohoku University, Sendai, Japan 5 College of Life Sciences, Ritsumeikan University, Kusatsu, Japan * Corresponding author: Takaaki Sato, Biomedical Research Institute, National Institute of Ad- vanced Industrial Science and Technology, 1-8-31 Midorioka, Ikeda, Osaka 563-8577, Japan, E-mail: [email protected] Abstract: G protein-coupled receptors (GPCRs) detect and distinguish between various subtypes of extracellular sig- nals, such as neurotransmitters, hormones, light, and odorous chemicals. As determinants for robust and appropriate cellular responses, common and unique features of interactions between GPCRs and their target G proteins provide insights into structure-based drug design for treatment of GPCR-related diseases. Re- cently, we found that the hydrophobic core buried between GPCR helix 8 and TM1–2 is essential for acti- vation of both specific and nonspecific G proteins. Furthermore, the 2nd residue of helix 8 is responsible for initial, transient, and specific interaction with a target G protein. Analysis of human and murine olfactory receptors (ORs) and other class-A GPCRs revealed that several amino acids, such as Glu, Gln, and Asp, are conserved at this position. -

Gene Expression Signature for Biliary Atresia and a Role for Interleukin-8
SUPPLEMENTARY “PATIENTS AND METHODS” “Gene expression signature for biliary atresia and a role for Interleukin-8 in pathogenesis of experimental disease” Bessho K, et al. PATIENTS AND METHODS Patients. Liver biopsies, serum samples and clinical data were obtained from infants with cholestasis enrolled into a prospective study (ClinicalTrials.gov Identifier: NCT00061828) of the NIDDK-funded Childhood Liver Disease Research and Education Network (www.childrennetwork.org) or from infants evaluated at Cincinnati Children’s Hospital Medical Center. For subjects with biliary atresia (BA), liver biopsies were obtained from 64 infants during the preoperative workup or at the time of intraoperative cholangiogram, with ages ranging from 22-169 days after birth (Supplementary Table 7). For subjects with intrahepatic cholestasis (serving as diseased controls, and referred to as non- BA), liver biopsy samples were obtained percutaneously or intraoperatively from 14 infants at the time of diagnostic evaluation, with ages ranging from 19-189 days. Their diagnosis were Alagille syndrome (N=1), multidrug resistance protein-3 deficiency (N=2), alpha-1-antitrypsin deficiency (N=2) and cholestasis with unknown etiology (N=9) (Supplementary Table 7). Representative liver biopsy photomicrographs are shown in Supplementary Figure 6A-D. A third group of normal controls (NC) consisted of liver biopsy samples obtained from 7 deceased-donor children aged 22-42 months as described previously (1). This group serves as a reference cohort, with the median levels of gene expression used to normalize gene expression across all patients in the BA and non-BA groups. This greatly facilitates the visual identification of key differences in gene expression levels between BA and non-BA groups. -

Table SII. Significantly Differentially Expressed Mrnas of GSE23558 Data Series with the Criteria of Adjusted P<0.05 And
Table SII. Significantly differentially expressed mRNAs of GSE23558 data series with the criteria of adjusted P<0.05 and logFC>1.5. Probe ID Adjusted P-value logFC Gene symbol Gene title A_23_P157793 1.52x10-5 6.91 CA9 carbonic anhydrase 9 A_23_P161698 1.14x10-4 5.86 MMP3 matrix metallopeptidase 3 A_23_P25150 1.49x10-9 5.67 HOXC9 homeobox C9 A_23_P13094 3.26x10-4 5.56 MMP10 matrix metallopeptidase 10 A_23_P48570 2.36x10-5 5.48 DHRS2 dehydrogenase A_23_P125278 3.03x10-3 5.40 CXCL11 C-X-C motif chemokine ligand 11 A_23_P321501 1.63x10-5 5.38 DHRS2 dehydrogenase A_23_P431388 2.27x10-6 5.33 SPOCD1 SPOC domain containing 1 A_24_P20607 5.13x10-4 5.32 CXCL11 C-X-C motif chemokine ligand 11 A_24_P11061 3.70x10-3 5.30 CSAG1 chondrosarcoma associated gene 1 A_23_P87700 1.03x10-4 5.25 MFAP5 microfibrillar associated protein 5 A_23_P150979 1.81x10-2 5.25 MUCL1 mucin like 1 A_23_P1691 2.71x10-8 5.12 MMP1 matrix metallopeptidase 1 A_23_P350005 2.53x10-4 5.12 TRIML2 tripartite motif family like 2 A_24_P303091 1.23x10-3 4.99 CXCL10 C-X-C motif chemokine ligand 10 A_24_P923612 1.60x10-5 4.95 PTHLH parathyroid hormone like hormone A_23_P7313 6.03x10-5 4.94 SPP1 secreted phosphoprotein 1 A_23_P122924 2.45x10-8 4.93 INHBA inhibin A subunit A_32_P155460 6.56x10-3 4.91 PICSAR P38 inhibited cutaneous squamous cell carcinoma associated lincRNA A_24_P686965 8.75x10-7 4.82 SH2D5 SH2 domain containing 5 A_23_P105475 7.74x10-3 4.70 SLCO1B3 solute carrier organic anion transporter family member 1B3 A_24_P85099 4.82x10-5 4.67 HMGA2 high mobility group AT-hook 2 A_24_P101651 -

S41467-019-12987-9.Pdf
Corrected: Author correction ARTICLE https://doi.org/10.1038/s41467-019-12987-9 OPEN Macrophage-associated wound healing contributes to African green monkey SIV pathogenesis control Fredrik Barrenas1,2,16, Kevin Raehtz3,4,16, Cuiling Xu5, Lynn Law6,7, Richard R. Green6,7, Guido Silvestri8,9, Steven E. Bosinger8,9, Andrew Nishida1, Qingsheng Li 10, Wuxun Lu10, Jianshui Zhang10, Matthew J. Thomas6,11, Jean Chang6,7, Elise Smith6,7, Jeffrey M. Weiss1, Reem A. Dawoud8, George H. Richter5, Anita Trichel12, Dongzhu Ma13, Xinxia Peng 14, Jan Komorowski 2,15, Cristian Apetrei3,4, Ivona Pandrea4,5 & Michael Gale Jr 6,7,11* 1234567890():,; Natural hosts of simian immunodeficiency virus (SIV) avoid AIDS despite lifelong infection. Here, we examined how this outcome is achieved by comparing a natural SIV host, African green monkey (AGM) to an AIDS susceptible species, rhesus macaque (RM). To asses gene expression profiles from acutely SIV infected AGMs and RMs, we developed a systems biology approach termed Conserved Gene Signature Analysis (CGSA), which compared RNA sequencing data from rectal AGM and RM tissues to various other species. We found that AGMs rapidly activate, and then maintain, evolutionarily conserved regenerative wound healing mechanisms in mucosal tissue. The wound healing protein fibronectin shows distinct tissue distribution and abundance kinetics in AGMs. Furthermore, AGM monocytes exhibit an embryonic development and repair/regeneration signature featuring TGF-β and con- comitant reduced expression of inflammatory genes compared to RMs. This regenerative wound healing process likely preserves mucosal integrity and prevents inflammatory insults that underlie immune exhaustion in RMs. 1 Department of Microbiology, University of Washington, Seattle, WA, USA. -

Human/Mouse Annexin A13 Antibody Antigen Affinity-Purified Polyclonal Goat Igg Catalog Number: AF4149
Human/Mouse Annexin A13 Antibody Antigen Affinity-purified Polyclonal Goat IgG Catalog Number: AF4149 DESCRIPTION Species Reactivity Human/Mouse Specificity Detects endogenous human and mouse Annexin A13 in Western blots. In Western blots, this antibody did not crossreact with recombinant human Annexin A1, A2, A3, A4, A5, A6, A7, A8, A9, A10, or A11. Source Polyclonal Goat IgG Purification Antigen Affinitypurified Immunogen E. coliderived recombinant human Annexin A13 Met1His316 Accession # P27216 Formulation Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. See Certificate of Analysis for details. *Small pack size (SP) is supplied either lyophilized or as a 0.2 μm filtered solution in PBS. APPLICATIONS Please Note: Optimal dilutions should be determined by each laboratory for each application. General Protocols are available in the Technical Information section on our website. Recommended Sample Concentration Western Blot 1 µg/mL See Below DATA Western Blot Detection of Human/Mouse Annexin A13 by Western Blot. Western blot shows lysates of human and mouse small intestine tissue. PVDF membrane was probed with 1 µg/mL of Human/Mouse Annexin A13 Antigen Affinitypurified Polyclonal Antibody (Catalog # AF4149) followed by HRPconjugated AntiGoat IgG Secondary Antibody (Catalog # HAF017). A specific band was detected for Annexin A13 at approximately 36 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1. PREPARATION AND STORAGE Reconstitution Reconstitute at 0.2 mg/mL in sterile PBS. Shipping The product is shipped at ambient temperature. Upon receipt, store it immediately at the temperature recommended below.