Methyl Directed DNA Mismatch Repair in Vibrio Cholerae
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Restriction Endonucleases
Molecular Biology Problem Solver: A Laboratory Guide. Edited by Alan S. Gerstein Copyright © 2001 by Wiley-Liss, Inc. ISBNs: 0-471-37972-7 (Paper); 0-471-22390-5 (Electronic) 9 Restriction Endonucleases Derek Robinson, Paul R. Walsh, and Joseph A. Bonventre Background Information . 226 Which Restriction Enzymes Are Commercially Available? . 226 Why Are Some Enzymes More Expensive Than Others? . 227 What Can You Do to Reduce the Cost of Working with Restriction Enzymes? . 228 If You Could Select among Several Restriction Enzymes for Your Application, What Criteria Should You Consider to Make the Most Appropriate Choice? . 229 What Are the General Properties of Restriction Endonucleases? . 232 What Insight Is Provided by a Restriction Enzyme’s Quality Control Data? . 233 How Stable Are Restriction Enzymes? . 236 How Stable Are Diluted Restriction Enzymes? . 236 Simple Digests . 236 How Should You Set up a Simple Restriction Digest? . 236 Is It Wise to Modify the Suggested Reaction Conditions? . 237 Complex Restriction Digestions . 239 How Can a Substrate Affect the Restriction Digest? . 239 Should You Alter the Reaction Volume and DNA Concentration? . 241 Double Digests: Simultaneous or Sequential? . 242 225 Genomic Digests . 244 When Preparing Genomic DNA for Southern Blotting, How Can You Determine If Complete Digestion Has Been Obtained? . 244 What Are Your Options If You Must Create Additional Rare or Unique Restriction Sites? . 247 Troubleshooting . 255 What Can Cause a Simple Restriction Digest to Fail? . 255 The Volume of Enzyme in the Vial Appears Very Low. Did Leakage Occur during Shipment? . 259 The Enzyme Shipment Sat on the Shipping Dock for Two Days. -

Dna Repair Mechanisms
DNA REPAIR MECHANISMS: A. P. NAGRALE DNA REPAIR •One of the main objectives of biological system is to maintain base sequences of DNA from one generation to the other. •DNA is relatively fragile, easily damaged molecule. •The DNA of a cell is subjected to damage from a variety of environmental factors like radiations (X-rays, UV-rays), chemicals mutagens, physical factors etc. •The survival of the cell depends on its ability to repair this damage. •DNA damage is of two types – Monoadduct and Diadduct. Monoadduct damage involves alterations in a single nitrogen base for example deamination reactions of chemical mutagen like HNO2. Diadduct damage are alterations involving more than one nitrogen base, for example Pyrimidine - Pyrimidine dimer produced by UV- radiations. •There are three principal types of repair mechanisms. •Light Repair (Enzymatic Photoreacivation) •Ultraviolet light causes the damage in DNA of bacteria and phages. UV-radiations (254 nm wavelength) cause the formation of covalent bond between adjacent pyrimidines to form dimer on the same strand of DNA (intrastrand bonding). •The UV- light forms abnormal structure Thymine dimmer (T=T) in DNA. As a result, DNA replication cannot proceed and gene expression is also prevented. •Damage to DNA caused by UV- light can be repaired after exposing the cells to visible light called photoreactivation or light repair. •This photoreactivation requires a specific enzyme that binds to the defective site on the DNA. •In this mechanism an enzyme DNA photolyase cleaves T=T dimmer and reverse to monomeric stage. •This enzyme is activated only when exposed to visible light. This enzyme absorbs the light energy, binds of cyclobutane ring to defective site of DNA and absorbed energy promotes the cleavage of covalent bond between two thymine molecules. -

Structure and Function of Nucleases in DNA Repair: Shape, Grip and Blade of the DNA Scissors
Oncogene (2002) 21, 9022 – 9032 ª 2002 Nature Publishing Group All rights reserved 0950 – 9232/02 $25.00 www.nature.com/onc Structure and function of nucleases in DNA repair: shape, grip and blade of the DNA scissors Tatsuya Nishino1 and Kosuke Morikawa*,1 1Department of Structural Biology, Biomolecular Engineering Research Institute (BERI), 6-2-3 Furuedai, Suita, Osaka 565-0874, Japan DNA nucleases catalyze the cleavage of phosphodiester mismatched nucleotides. They also recognize the bonds. These enzymes play crucial roles in various DNA replication or recombination intermediates to facilitate repair processes, which involve DNA replication, base the following reaction steps through the cleavage of excision repair, nucleotide excision repair, mismatch DNA strands (Table 1). repair, and double strand break repair. In recent years, Nucleases can be regarded as molecular scissors, new nucleases involved in various DNA repair processes which cleave phosphodiester bonds between the sugars have been reported, including the Mus81 : Mms4 (Eme1) and the phosphate moieties of DNA. They contain complex, which functions during the meiotic phase and conserved minimal motifs, which usually consist of the Artemis : DNA-PK complex, which processes a V(D)J acidic and basic residues forming the active site. recombination intermediate. Defects of these nucleases These active site residues coordinate catalytically cause genetic instability or severe immunodeficiency. essential divalent cations, such as magnesium, Thus, structural biology on various nuclease actions is calcium, manganese or zinc, as a cofactor. However, essential for the elucidation of the molecular mechanism the requirements for actual cleavage, such as the types of complex DNA repair machinery. Three-dimensional and the numbers of metals, are very complicated, but structural information of nucleases is also rapidly are not common among the nucleases. -

SUPPLEMENTARY INFORMATION in Silico Signature Prediction
SUPPLEMENTARY INFORMATION In Silico Signature Prediction Modeling in Cytolethal Distending Toxin-Producing Escherichia coli Strains Maryam Javadi, Mana Oloomi*, Saeid Bouzari Department of Molecular Biology, Pasteur Institute of Iran, Tehran 13164, Iran http://www.genominfo.org/src/sm/gni-15-69-s001.pdf Supplementary Table 6. Aalphabetic abbreviation and description of putative conserved domains Alphabetic Abbreviation Description 17 Large terminase protein 2_A_01_02 Multidrug resistance protein 2A0115 Benzoate transport; [Transport and binding proteins, Carbohydrates, organic alcohols] 52 DNA topisomerase II medium subunit; Provisional AAA_13 AAA domain; This family of domains contain a P-loop motif AAA_15 AAA ATPase domain; This family of domains contain a P-loop motif AAA_21 AAA domain AAA_23 AAA domain ABC_RecF ATP-binding cassette domain of RecF; RecF is a recombinational DNA repair ATPase ABC_SMC_barmotin ATP-binding cassette domain of barmotin, a member of the SMC protein family AcCoA-C-Actrans Acetyl-CoA acetyltransferases AHBA_syn 3-Amino-5-hydroxybenzoic acid synthase family (AHBA_syn) AidA Type V secretory pathway, adhesin AidA [Cell envelope biogenesis] Ail_Lom Enterobacterial Ail/Lom protein; This family consists of several bacterial and phage Ail_Lom proteins AIP3 Actin interacting protein 3; Aip3p/Bud6p is a regulator of cell and cytoskeletal polarity Aldose_epim_Ec_YphB Aldose 1-epimerase, similar to Escherichia coli YphB AlpA Predicted transcriptional regulator [Transcription] AntA AntA/AntB antirepressor AraC AraC-type -

Open Matthew R Moreau Ph.D. Dissertation Finalfinal.Pdf
The Pennsylvania State University The Graduate School Department of Veterinary and Biomedical Sciences Pathobiology Program PATHOGENOMICS AND SOURCE DYNAMICS OF SALMONELLA ENTERICA SEROVAR ENTERITIDIS A Dissertation in Pathobiology by Matthew Raymond Moreau 2015 Matthew R. Moreau Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy May 2015 The Dissertation of Matthew R. Moreau was reviewed and approved* by the following: Subhashinie Kariyawasam Associate Professor, Veterinary and Biomedical Sciences Dissertation Adviser Co-Chair of Committee Bhushan M. Jayarao Professor, Veterinary and Biomedical Sciences Dissertation Adviser Co-Chair of Committee Mary J. Kennett Professor, Veterinary and Biomedical Sciences Vijay Kumar Assistant Professor, Department of Nutritional Sciences Anthony Schmitt Associate Professor, Veterinary and Biomedical Sciences Head of the Pathobiology Graduate Program *Signatures are on file in the Graduate School iii ABSTRACT Salmonella enterica serovar Enteritidis (SE) is one of the most frequent common causes of morbidity and mortality in humans due to consumption of contaminated eggs and egg products. The association between egg contamination and foodborne outbreaks of SE suggests egg derived SE might be more adept to cause human illness than SE from other sources. Therefore, there is a need to understand the molecular mechanisms underlying the ability of egg- derived SE to colonize the chicken intestinal and reproductive tracts and cause disease in the human host. To this end, the present study was carried out in three objectives. The first objective was to sequence two egg-derived SE isolates belonging to the PFGE type JEGX01.0004 to identify the genes that might be involved in SE colonization and/or pathogenesis. -

Roles of Methylation and Sequestration in the Mechanisms of DNA Replication in Some Members of the Enterobacteriaceae Family
Chapter 12 Roles of Methylation and Sequestration in the Mechanisms of DNA Replication in some Members of the Enterobacteriaceae Family Amine Aloui, Alya El May, Saloua Kouass Sahbani and Ahmed Landoulsi Additional information is available at the end of the chapter http://dx.doi.org/10.5772/51724 1. Introduction When growing cells divide, they need to copy their genetic material and distribute it to en‐ sure that each daughter cell receives one copy. This is a challenging task especially when the enormous length of the DNA compared to the cell size is considered. During DNA replica‐ tion, organization of the chromosomes is even more demanding, since replication forks con‐ tinuously produce new DNA. This DNA contains all the information required to build the cells and tissues of a prokaryotic or an eukaryotic organism. The exact replication of this in‐ formation in any species assures its genetic continuity from generation to generation and is critical to the normal development of an individual. The information stored in DNA is ar‐ ranged in hereditary units known as genes that control the identifiable traits of an organism. Discovery of the structure of DNA and subsequent elucidation of how DNA directs synthe‐ sis of RNA, which then directs assembly of proteins -the so-called central dogma - were monumental achievements that marked the early days of molecular biology. However, the simplified representation of the central dogma as DNA → RNA → protein does not reflect the role of proteins in the synthesis of nucleic acids. Moreover, proteins are largely responsi‐ ble for regulating DNA replication and gene expression, the entire process whereby the in‐ formation encoded in DNA is decoded into the proteins that characterize various cell types. -

Review DNA Repair Nucleases
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by RERO DOC Digital Library CMLS, Cell. Mol. Life Sci. 61 (2004) 336–354 1420-682X/04/030336-19 DOI 10.1007/s00018-003-3223-4 CMLS Cellular and Molecular Life Sciences © Birkhäuser Verlag, Basel, 2004 Review DNA repair nucleases T. M. Martia and O. Fleck b, * Institute of Cell Biology, University of Bern, Baltzerstrasse 4, 3012 Bern (Switzerland), Fax: +41 31 631 4684, e-mail: [email protected] a Present address: UCSF Comprehensive Cancer Center, University of California, 2340 Sutter Street, Box 0808, San Francisco, California 94143 (USA) b Present address: Department of Genetics, Institute of Molecular Biology, University of Copenhagen, Øster Farimagsgade 2A, 1353 Copenhagen K (Denmark), Fax: +45 35 32 2113, e-mail: [email protected] Received 12 June 2003; received after revision 29 July 2003; accepted 16 September 2003 Abstract. Stability of DNA largely depends on accuracy of DNA. Flap endonucleases cleave DNA flap structures of repair mechanisms, which remove structural anomalies at or near the junction between single-stranded and dou- induced by exogenous and endogenous agents or intro- ble-stranded regions. DNA nucleases play a crucial role in duced by DNA metabolism, such as replication. Most re- mismatch repair, nucleotide excision repair, base excision pair mechanisms include nucleolytic processing of DNA, repair and double-strand break repair. In addition, nucle- where nucleases cleave a phosphodiester bond between a olytic repair functions are required during replication to deoxyribose and a phosphate residue, thereby producing remove misincorporated nucleotides, Okazaki fragments 5′-terminal phosphate and 3′-terminal hydroxyl groups. -

Identifying Functional Roles for Alkb in the Adaptive
IDENTIFYING FUNCTIONAL ROLES FOR ALKB IN THE ADAPTIVE RESPONSE OF ESCHERICHIA COLI TO ALKYLATION DAMAGE SUNEET DINGLAY A thesis submitted for Ph.D.; 2000 Imperial Cancer Research Fund, Clare Hall Laboratories, Blanche Lane, South Mimms, Herts, EN6 3LD & University College London, Gower Street, London, WC1E 6BT ProQuest Number: 10608883 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a com plete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. uest ProQuest 10608883 Published by ProQuest LLC(2017). Copyright of the Dissertation is held by the Author. All rights reserved. This work is protected against unauthorized copying under Title 17, United States C ode Microform Edition © ProQuest LLC. ProQuest LLC. 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, Ml 48106- 1346 In loving memory of my Papa ji. ABSTRACT In 1977 a novel, inducible and error free DNA repair system in Escherichia coli came to light. It protected E. coli against the mutagenic and cytotoxic effects of alkylating agents such as N-methyl-N’-nitro-N-nitrosoguanidine (MNNG) and methyl methanesulfonate (MMS), and was termed the ‘adaptive response of E. coli to alkylation damage’. This response consists of four inducible genes; ada, aidB, alkA and alkB. The ada gene product encodes an 06-methylguanine- DNA methyltransferase, and is also the positive regulator of the response. The alkA gene product encodes a 3-methyladenine- DNA glycosylase, and aidB shows homology to several mammalian acyl coenzyme A dehydrogenases. -

Deciphering Bacterial Epigenomes Using Modern Sequencing Technologies
REVIEWS Deciphering bacterial epigenomes using modern sequencing technologies John Beaulaurier, Eric E. Schadt and Gang Fang * Abstract | Prokaryotic DNA contains three types of methylation: N6-methyladenine, N4-methylcytosine and 5-methylcytosine. The lack of tools to analyse the frequency and distribution of methylated residues in bacterial genomes has prevented a full understanding of their functions. Now , advances in DNA sequencing technology , including single- molecule, real- time sequencing and nanopore- based sequencing, have provided new opportunities for systematic detection of all three forms of methylated DNA at a genome- wide scale and offer unprecedented opportunities for achieving a more complete understanding of bacterial epigenomes. Indeed, as the number of mapped bacterial methylomes approaches 2,000, increasing evidence supports roles for methylation in regulation of gene expression, virulence and pathogen–host interactions. Phase variation DNA methylation was discovered in bacteria more than MTases has been found to play important regulatory 1 2–12 A means by which reversible a half century ago . It is now known that modification roles in bacteria . RM systems protect cells from variation of protein expression of the four canonical DNA bases by methylation can act invading DNA by methylating endogenous DNA and is achieved in bacteria, often in as an epigenetic regulator — that is, it can impart dis- cleaving non- methylated foreign DNA2,4. RM sys- an ON/OFF manner. The process creates phenotypic tinct and reversible regulatory states to identical genetic tems are divided into three main categories based diversity in a clonally expanded sequences. In eukaryotes, epigenetic regulation can on the subunits involved and the precise site of DNA population and allows the occur at multiple levels: DNA methylation, nucleosome restriction13–16 (Fig. -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Homing Endonucleases: from Microbial Genetic Invaders to Reagents for Targeted DNA Modification
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Structure Review Homing Endonucleases: From Microbial Genetic Invaders to Reagents for Targeted DNA Modification Barry L. Stoddard1,* 1Division of Basic Sciences, Fred Hutchinson Cancer Research Center, 1100 Fairview Avenue N., A3-025, Seattle, WA 98109, USA *Correspondence: [email protected] DOI 10.1016/j.str.2010.12.003 Homing endonucleases are microbial DNA-cleaving enzymes that mobilize their own reading frames by generating double strand breaks at specific genomic invasion sites. These proteins display an economy of size, and yet recognize long DNA sequences (typically 20 to 30 base pairs). They exhibit a wide range of fidelity at individual nucleotide positions in a manner that is strongly influenced by host constraints on the coding sequence of the targeted gene. The activity of these proteins leads to site-specific recombination events that can result in the insertion, deletion, mutation, or correction of DNA sequences. Over the past fifteen years, the crystal structures of representatives from several homing endonuclease families have been solved, and methods have been described to create variants of these enzymes that cleave novel DNA targets. Engineered homing endonucleases proteins are now being used to generate targeted genomic modifications for a variety of biotech and medical applications. Endonuclease enzymes are ubiquitous catalysts that are A series of studies conducted in several laboratories over the involved in genomic modification, rearrangement, protection, ensuing years led to the discovery of homing endonucleases and and repair. Their specificity spans at least nine orders of magni- mobile introns and inteins within a wide variety of additional tude, ranging from nonspecific degradative enzymes up to microbial genomes (reviewed in Belfort and Perlman, 1995). -
Q 297 Suppl USE
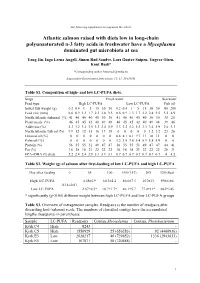
The following supplement accompanies the article Atlantic salmon raised with diets low in long-chain polyunsaturated n-3 fatty acids in freshwater have a Mycoplasma dominated gut microbiota at sea Yang Jin, Inga Leena Angell, Simen Rød Sandve, Lars Gustav Snipen, Yngvar Olsen, Knut Rudi* *Corresponding author: [email protected] Aquaculture Environment Interactions 11: 31–39 (2019) Table S1. Composition of high- and low LC-PUFA diets. Stage Fresh water Sea water Feed type High LC-PUFA Low LC-PUFA Fish oil Initial fish weight (g) 0.2 0.4 1 5 15 30 50 0.2 0.4 1 5 15 30 50 80 200 Feed size (mm) 0.6 0.9 1.3 1.7 2.2 2.8 3.5 0.6 0.9 1.3 1.7 2.2 2.8 3.5 3.5 4.9 North Atlantic fishmeal (%) 41 40 40 40 40 30 30 41 40 40 40 40 30 30 35 25 Plant meals (%) 46 45 45 42 40 49 48 46 45 45 42 40 49 48 39 46 Additives (%) 3.3 3.2 3.2 3.5 3.3 3.4 3.9 3.3 3.2 3.2 3.5 3.3 3.4 3.9 2.6 3.3 North Atlantic fish oil (%) 9.9 12 12 15 16 17 18 0 0 0 0 0 1.2 1.2 23 26 Linseed oil (%) 0 0 0 0 0 0 0 6.8 8.1 8.1 9.7 11 10 11 0 0 Palm oil (%) 0 0 0 0 0 0 0 3.2 3.8 3.8 5.4 5.9 5.8 5.9 0 0 Protein (%) 56 55 55 51 49 47 47 56 55 55 51 49 47 47 44 41 Fat (%) 16 18 18 21 22 22 22 16 18 18 21 22 22 22 28 31 EPA+DHA (% diet) 2.2 2.4 2.4 2.9 3.1 3.1 3.1 0.7 0.7 0.7 0.7 0.7 0.7 0.7 4 4.2 Table S2.