Temporally-Divergent Regulatory Mechanisms Govern Neuronal Development and Diversification in the Neocortex
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Palmitic Acid Effects on Hypothalamic Neurons
bioRxiv preprint doi: https://doi.org/10.1101/2021.08.03.454666; this version posted August 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Running title: Oleic and palmitic acid effects on hypothalamic neurons Concentration-dependent change in hypothalamic neuronal transcriptome by the dietary fatty acids: oleic and palmitic acids Fabiola Pacheco Valencia1^, Amanda F. Marino1^, Christos Noutsos1, Kinning Poon1* 1Department of Biological Sciences, SUNY Old Westbury, Old Westbury NY, United States ^Authors contributed equally to this work *Corresponding Author: Kinning Poon 223 Store Hill Rd Old Westbury, NY 11568, USA 1-516-876-2735 [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2021.08.03.454666; this version posted August 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Prenatal high-fat diet exposure increases hypothalamic neurogenesis events in embryos and programs offspring to be obesity-prone. The molecular mechanism involved in these dietary effects of neurogenesis are unknown. This study investigated the effects of oleic and palmitic acids, which are abundant in a high-fat diet, on the hypothalamic neuronal transcriptome and how these changes impact neurogenesis events. The results show differential effects of low and high concentrations of oleic or palmitic acid treatment on differential gene transcription. -

EBF1 Drives Hallmark B Cell Gene Expression by Enabling the Interaction of PAX5 with the MLL H3K4 Methyltransferase Complex Charles E
www.nature.com/scientificreports OPEN EBF1 drives hallmark B cell gene expression by enabling the interaction of PAX5 with the MLL H3K4 methyltransferase complex Charles E. Bullerwell1, Philippe Pierre Robichaud1,2, Pierre M. L. Deprez1, Andrew P. Joy1, Gabriel Wajnberg1, Darwin D’Souza1,3, Simi Chacko1, Sébastien Fournier1, Nicolas Crapoulet1, David A. Barnett1,2, Stephen M. Lewis1,2 & Rodney J. Ouellette1,2* PAX5 and EBF1 work synergistically to regulate genes that are involved in B lymphocyte diferentiation. We used the KIS-1 difuse large B cell lymphoma cell line, which is reported to have elevated levels of PAX5 expression, to investigate the mechanism of EBF1- and PAX5-regulated gene expression. We demonstrate the lack of expression of hallmark B cell genes, including CD19, CD79b, and EBF1, in the KIS-1 cell line. Upon restoration of EBF1 expression we observed activation of CD19, CD79b and other genes with critical roles in B cell diferentiation. Mass spectrometry analyses of proteins co-immunoprecipitated with PAX5 in KIS-1 identifed components of the MLL H3K4 methylation complex, which drives histone modifcations associated with transcription activation. Immunoblotting showed a stronger association of this complex with PAX5 in the presence of EBF1. Silencing of KMT2A, the catalytic component of MLL, repressed the ability of exogenous EBF1 to activate transcription of both CD19 and CD79b in KIS-1 cells. We also fnd association of PAX5 with the MLL complex and decreased CD19 expression following silencing of KMT2A in other human B cell lines. These data support an important role for the MLL complex in PAX5-mediated transcription regulation. -
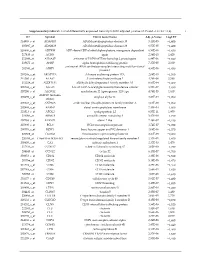
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Mutations in PAX2 Associate with Adult-Onset FSGS
BASIC RESEARCH www.jasn.org Mutations in PAX2 Associate with Adult-Onset FSGS † ‡ | Moumita Barua,* Emilia Stellacci, Lorenzo Stella, Astrid Weins,§ Giulio Genovese,* ¶** † †† ‡‡ Valentina Muto, Viviana Caputo, Hakan R. Toka,* Victoria T. Charoonratana,* † Marco Tartaglia, and Martin R. Pollak* *Division of Nephrology, Department of Medicine, Beth Israel Deaconess Medical Center, Boston, Massachusetts; †Department of Hematology, Oncology and Molecular Medicine, National Institute of Health, Rome, Italy; ‡Department of Chemical Science and Technology, University of Rome Tor Vergata, Rome, Italy; §Department of Pathology, and ‡‡Division of Nephrology, Department of Medicine, Brigham and Women’s Hospital, Boston, Massachusetts; |Stanley Center for Psychiatric Research, Cambridge, Massachusetts; ¶Program in Medical and Population Genetics, Broad Institute of MIT and Harvard, Cambridge, Massachusetts; **Department of Genetics, Harvard Medical School, Boston, Massachusetts; and ††Department of Experimental Medicine, University “La Sapienza,” Rome, Italy ABSTRACT FSGS is characterized by the presence of partial sclerosis of some but not all glomeruli. Studies of familial FSGS have been instrumental in identifying podocytes as critical elements in maintaining glomerular function, but underlying mutations have not been identified for all forms of this genetically heterogeneous condition. Here, exome sequencing in members of an index family with dominant FSGS revealed a nonconservative, disease-segregating variant in the PAX2 transcription factor -

Transcriptional Repression by FEZF2 Restricts Alternative Identities of Cortical Projection Neurons
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Cell Rep Manuscript Author . Author manuscript; Manuscript Author available in PMC 2021 August 02. Published in final edited form as: Cell Rep. 2021 June 22; 35(12): 109269. doi:10.1016/j.celrep.2021.109269. Transcriptional repression by FEZF2 restricts alternative identities of cortical projection neurons Jeremiah Tsyporin1,7, David Tastad1,7, Xiaokuang Ma2, Antoine Nehme2, Thomas Finn1, Liora Huebner1, Guoping Liu3, Daisy Gallardo1, Amr Makhamreh1, Jacqueline M. Roberts1, Solomon Katzman4, Nenad Sestan5, Susan K. McConnell6, Zhengang Yang3, Shenfeng Qiu2, Bin Chen1,8,* 1Department of Molecular, Cell, and Developmental Biology, University of California, Santa Cruz, 1156 High Street, Santa Cruz, CA 95064, USA 2Department of Basic Medical Sciences, University of Arizona College of Medicine – Phoenix, Phoenix, AZ 85004, USA 3State Key Laboratory of Medical Neurobiology and MOE Frontiers Center for Brain Science, Institute for Translational Brain Research, Institutes of Brain Science, Department of Neurology, Zhongshan Hospital, Fudan University, Shanghai 200032, China 4Genomics Institute, University of California, Santa Cruz, Santa Cruz, CA 95064, USA 5Department of Neuroscience, Yale School of Medicine, New Haven, CT 06520, USA 6Department of Biology, Stanford University, Stanford, CA 94305, USA 7These authors contributed equally 8Lead contact SUMMARY Projection neuron subtype identities in the cerebral cortex are established by expressing pan- cortical and subtype-specific effector genes that execute terminal differentiation programs bestowing neurons with a glutamatergic neuron phenotype and subtype-specific morphology, physiology, and axonal projections. Whether pan-cortical glutamatergic and subtype-specific characteristics are regulated by the same genes or controlled by distinct programs remains largely unknown. -

Characterization of Acute Myeloid Leukemia with Del(9Q) – Impact of the Genes in the Minimally Deleted Region T ⁎ Isabel S
Leukemia Research 76 (2019) 15–23 Contents lists available at ScienceDirect Leukemia Research journal homepage: www.elsevier.com/locate/leukres Research paper Characterization of acute myeloid leukemia with del(9q) – Impact of the genes in the minimally deleted region T ⁎ Isabel S. Naarmann-de Vriesa, , Yvonne Sackmanna,b, Felicitas Kleina,b, Antje Ostareck-Lederera, Dirk H. Ostarecka, Edgar Jostb, Gerhard Ehningerc, Tim H. Brümmendorfb, Gernot Marxa, ⁎⁎ Christoph Rölligc, Christian Thiedec, Martina Crysandtb, a Department of Intensive Care Medicine, University Hospital RWTH Aachen University, Aachen, Germany b Department of Hematology, Oncology, Hemostaseology and Stem Cell Transplantation, University Hospital RWTH Aachen University, Aachen, Germany c Medical Department I, University Hospital of the Technical University Dresden, Dresden, Germany ARTICLE INFO ABSTRACT Keywords: Acute myeloid leukemia is an aggressive disease that arises from clonal expansion of malignant hematopoietic AML precursor cells of the bone marrow. Deletions on the long arm of chromosome 9 (del(9q)) are observed in 2% of del(9q) acute myeloid leukemia patients. Our deletion analysis in a cohort of 31 del(9q) acute myeloid leukemia patients HNRNPK further supports the importance of a minimally deleted region composed of seven genes potentially involved in CEBPA leukemogenesis: GKAP1, KIF27, C9ORF64, HNRNPK, RMI1, SLC28A3 and NTRK2. Importantly, among them HNRNPK, encoding heterogeneous nuclear ribonucleoprotein K is proposed to function in leukemogenesis. We show that expression of HNRNPK and the other genes of the minimally deleted region is significantly reduced in patients with del(9q) compared with normal karyotype acute myeloid leukemia. Also, two mRNAs interacting with heterogeneous nuclear ribonucleoprotein K, namely CDKN1A and CEBPA are significantly downregulated. -

Dissecting the Genetics of Human Communication
DISSECTING THE GENETICS OF HUMAN COMMUNICATION: INSIGHTS INTO SPEECH, LANGUAGE, AND READING by HEATHER ASHLEY VOSS-HOYNES Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Epidemiology and Biostatistics CASE WESTERN RESERVE UNIVERSITY January 2017 CASE WESTERN RESERVE UNIVERSITY SCHOOL OF GRADUATE STUDIES We herby approve the dissertation of Heather Ashely Voss-Hoynes Candidate for the degree of Doctor of Philosophy*. Committee Chair Sudha K. Iyengar Committee Member William Bush Committee Member Barbara Lewis Committee Member Catherine Stein Date of Defense July 13, 2016 *We also certify that written approval has been obtained for any proprietary material contained therein Table of Contents List of Tables 3 List of Figures 5 Acknowledgements 7 List of Abbreviations 9 Abstract 10 CHAPTER 1: Introduction and Specific Aims 12 CHAPTER 2: Review of speech sound disorders: epidemiology, quantitative components, and genetics 15 1. Basic Epidemiology 15 2. Endophenotypes of Speech Sound Disorders 17 3. Evidence for Genetic Basis Of Speech Sound Disorders 22 4. Genetic Studies of Speech Sound Disorders 23 5. Limitations of Previous Studies 32 CHAPTER 3: Methods 33 1. Phenotype Data 33 2. Tests For Quantitative Traits 36 4. Analytical Methods 42 CHAPTER 4: Aim I- Genome Wide Association Study 49 1. Introduction 49 2. Methods 49 3. Sample 50 5. Statistical Procedures 53 6. Results 53 8. Discussion 71 CHAPTER 5: Accounting for comorbid conditions 84 1. Introduction 84 2. Methods 86 3. Results 87 4. Discussion 105 CHAPTER 6: Hypothesis driven pathway analysis 111 1. Introduction 111 2. Methods 112 3. Results 116 4. -

SNP-Array Lesions in Core Binding Factor Acute Myeloid Leukemia
www.impactjournals.com/oncotarget/ Oncotarget, 2018, Vol. 9, (No. 5), pp: 6478-6489 Research Paper SNP-array lesions in core binding factor acute myeloid leukemia Nicolas Duployez1,2, Elise Boudry-Labis2,3, Christophe Roumier1,2, Nicolas Boissel4, Arnaud Petit5,6, Sandrine Geffroy1,2, Nathalie Helevaut1,2, Karine Celli-Lebras4, Christine Terré7, Odile Fenneteau8, Wendy Cuccuini9, Isabelle Luquet10, Hélène Lapillonne6,11, Catherine Lacombe12, Pascale Cornillet13, Norbert Ifrah14, Hervé Dombret4, Guy Leverger5,6, Eric Jourdan15 and Claude Preudhomme1,2 1CHU Lille, Laboratory of Hematology, Biology and Pathology Center, Lille, France 2Univ. Lille, INSERM, UMR-S 1172, JPARC, Lille, France 3CHU Lille, Institute of Medical Genetics, Jeanne de Flandre Hospital, Lille, France 4APHP, Department of Hematology, Saint Louis Hospital, Paris, France 5APHP, Department of Pediatric Hematology and Oncology, GH HUEP, Trousseau Hospital, Paris, France 6Sorbonne Universites, UPMC Univ Paris 06, UMR-S 938, CDR Saint-Antoine, GRC n°07, GRC MyPAC, Paris, France 7CH Versailles, Department of Genetics, Le Chesnay, France 8APHP, Laboratory of Hematology, Robert Debré University Hospital, Paris, France 9APHP, Department of Cytogenetics, Saint Louis Hospital, Paris, France 10CHU Toulouse, Laboratory of Hematology, Toulouse, France 11APHP, Laboratory of Hematology, GH HUEP, Trousseau Hospital, Paris, France 12APHP, Goelamsthèque, Cochin Hospital, Paris, France 13CHU Reims, Laboratory of Hematology, Reims, France 14CHU Angers, INSERM CRCINA, Department of Hematology, Angers, France 15CHU Nîmes, Department of Hematology, Nîmes, France Correspondence to: Claude Preudhomme, email: [email protected] Nicolas Duployez, email: [email protected] Keywords: acute myeloid leukemia; core binding factor; SNP-array; sequencing Received: November 21, 2017 Accepted: January 03, 2018 Published: January 08, 2018 Copyright: Duployez et al. -

Mouse Tle4 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Tle4 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Tle4 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Tle4 gene (NCBI Reference Sequence: NM_011600 ; Ensembl: ENSMUSG00000024642 ) is located on Mouse chromosome 19. 20 exons are identified, with the ATG start codon in exon 1 and the TAA stop codon in exon 20 (Transcript: ENSMUST00000052011). Exon 7 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Tle4 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-68P6 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a knock-out allele are runted and die around 4 weeks of age with leukocytopenia, B cell lymphopenia, reduced bone mineralization and reduced hematopoietic stem cell number and function. Exon 7 starts from about 16.86% of the coding region. The knockout of Exon 7 will result in frameshift of the gene. The size of intron 6 for 5'-loxP site insertion: 26814 bp, and the size of intron 7 for 3'-loxP site insertion: 1396 bp. The size of effective cKO region: ~702 bp. The cKO region does not have any other known gene. Page 1 of 7 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 7 8 20 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Tle4 Homology arm cKO region loxP site Page 2 of 7 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Prioritization of Putatively Detrimental Variants in Euploid Miscarriages
medRxiv preprint doi: https://doi.org/10.1101/2021.01.02.20248961; this version posted March 12, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC 4.0 International license . Prioritization of putatively detrimental variants in euploid miscarriages Silvia Buonaiuto1+, Immacolata Di Biase2+, Valentina Aleotti3, Amin Ravaei3, Adriano De Marino4, Gianluca Damaggio5, Marco Chierici6, Madhuri Pulijala7, Palmira D’Ambrosio2, Gabriella Esposito2, Qasim Ayub7, Cesare Furlanello9, Pantaleo Greco10, Antonio Capalbo4, Michele Rubini3, Sebastiano Di Biase2, and Vincenza Colonna11* 1University of Campania Luigi Vanvitelli, 81100, Italy 2MeriGen Research, Naples, 80131, Italy 3Department of Neurosciences and Rehabilitation, University of Ferrara, Ferrara, 44121, Italy. 4Igenomix Italy, Marostica, 36063, Italy 5University of Naples Federico II, Italy 6Fondazione Bruno Kessler, DSH Lab, Trento, 38123, Italy 7Monash University Malaysia Genomics Facility, Tropical Medicine and Biology Multidisciplinary Platform, 47500 Bandar Sunway, Malaysia. 8School of Science, Monash University Malaysia, 47500 Bandar Sunway, Malaysia. 9HK3 Lab, Rovereto,38068, Italy 10Department of Medical Sciences, University of Ferrara, Ferrara, 44121,Italy. 11Institute of Genetics and Biophysics, National Research Council, Naples, 80111, Italy *Correspondence: [email protected] +these authors contributed equally to this work ABSTRACT Miscarriage is the spontaneous termination of a pregnancy before 24 weeks of gestation. We studied the genome of euploid miscarried embryos from mothers in the range of healthy adult individuals to understand genetic susceptibility to miscarriage not caused by chromosomal aneuploidies. We developed GP , a pipeline that we used to prioritize 439 unique variants in 399 genes, including genes known to be associated with miscarriages. -
Temporally-Divergent Regulatory Mechanisms Govern Neuronal Development and Diversification in the Neocortex
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.23.263434; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Temporally-Divergent Regulatory Mechanisms Govern Neuronal Development and Diversification in the Neocortex Wen Yuan1, Sai Ma1,2,3, Juliana R. Brown1, Kwanho Kim1,2,4, Vanessa Murek5, Lucia Trastulla5, Alexander Meissner1,6,7, Simona Lodato8, Ashwin Shetty1, Joshua Z. Levin2,4, Jason D. Buenrostro1,7, Michael J. Ziller5*, and Paola Arlotta1,4* Affiliations: 1Department of Stem Cell and Regenerative Biology, Harvard University, Cambridge, MA 02138, USA 2Klarman Cell Observatory, Broad Institute of MIT and Harvard, Cambridge, MA, 02142, USA 3Department of Biology and Koch Institute, Massachusetts Institute of Technology, Cambridge, MA, 02142 USA 4Stanley Center for Psychiatric Research, Broad Institute of Harvard and MIT, Cambridge, MA 02142, USA 5Max Planck Institute of Psychiatry, 80804 Munich, Germany 6Department of Genome Regulation, Max Planck Institute for Molecular Genetics, 14195 Berlin, Germany 7Broad Institute of MIT and Harvard, Cambridge, MA 02142, USA 8Department of Biomedical Sciences and Humanitas Clinical and Research Center – IRCCS, Humanitas University, Milan, Italy *Correspondence to: [email protected] and [email protected] Abstract: Mammalian neocortical neurons span one of the most diverse cell type spectra of any tissue. The regulatory strategies that neurons use during progressive development and maturation remain unclear. We present an integrated single-cell epigenomic and transcriptional analysis of individual classes of neurons from both mouse and marmoset neocortex, sampled during both early postmitotic stages of identity acquisition and later stages of neuronal plasticity and circuit integration. -
Large Scale Molecular Studies of Pituitary Neuroendocrine Tumors: Novel Markers, Mechanisms and Translational Perspectives
cancers Review Large Scale Molecular Studies of Pituitary Neuroendocrine Tumors: Novel Markers, Mechanisms and Translational Perspectives Raitis Peculis †, Helvijs Niedra † and Vita Rovite * Latvian Biomedical Research and Study Centre, Ratsupites Str. 1-k1, LV-1067 Riga, Latvia; [email protected] (R.P.); [email protected] (H.N.) * Correspondence: [email protected]; Tel.: +371-674-730-83 † Both authors contributed equally to this review. Simple Summary: Pituitary neuroendocrine tumors are non-cancerous tumors of the pituitary gland, that may overproduce hormones leading to serious health conditions or due to tumor size cause chronic headache, vertigo or visual impairment. In recent years pituitary neuroendocrine tumors are studied with the latest molecular biology methods that simultaneously investigate a large number of factors to understand the mechanisms of how these tumors develop and how they could be diagnosed or treated. In this review article, we have studied literature reports, compiled information and described molecular factors that could affect the development and clinical characteristics of pituitary neuroendocrine tumors, discovered factors that overlap between several studies using large scale molecular analysis and interpreted the potential involvement of these factors in pituitary tumor Citation: Peculis, R.; Niedra, H.; development. Overall, this study provides a valuable resource for understanding the biology of Rovite, V. Large Scale Molecular pituitary neuroendocrine tumors. Studies of Pituitary Neuroendocrine Tumors: Novel Markers, Mechanisms Abstract: Pituitary neuroendocrine tumors (PitNETs) are non-metastatic neoplasms of the pituitary, and Translational Perspectives. which overproduce hormones leading to systemic disorders, or tumor mass effects causing headaches, Cancers 2021, 13, 1395. https:// vertigo or visual impairment.