MAP17, a ROS-Dependent Oncogene
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Role of Cdep in the Embryonic Morphogenesis of Drosophila Melanogaster
The Role of Cdep in the Embryonic Morphogenesis of Drosophila melanogaster Dissertation zur Erlangung des akademischen Grades Doctor rerum naturalium (Dr. rer.nat.) vorgelegt der Fakultät Mathematik und Naturwissenschaften der Technischen Universität Dresden von Anne Morbach Geboren am 05. August 1984 in Nordhausen, Deutschland Eingereicht am 30. Oktober 2015 Die Dissertation wurde in der Zeit von April 2012 bis Oktober 2015 im Max-Planck-Institut für molekulare Zellbiologie und Genetik angefertigt Gutachter: Prof. Dr. Elisabeth Knust Prof. Dr. Christian Dahmann Now, here, you see, it takes all the running you can do, to keep in the same place. Lewis Carroll, Through the Looking-Glass I Contents List of Figures V List of Tables VII Abstract IX List of Abbreviations XI 1 Introduction 1 1.1 Epithelial cell polarity . 1 1.1.1 Cellularization and formation of the primary epithelium . 1 1.1.1.1 Establishment of epithelial polarity and adhesion . 2 1.1.2 The epithelial polarity network . 3 1.1.3 Cell-cell adhesion . 5 1.1.3.1 Adherens junctions . 5 1.1.3.2 Septate junctions . 6 1.2 Epithelial movements in Drosophila embryonic morphogenesis . 6 1.2.1 Epithelial tube formation during Drosophila embryogenesis . 7 1.2.2 Coordinated migration of epithelial sheets during Dros. embryogenesis 7 1.2.2.1 FERM domain proteins in epithelial migration . 9 1.2.2.2 Cdep . 10 1.3 Mutagenesis with the CRISPR/Cas9 system . 12 2 Aim of My PhD Thesis Work 15 3 Preliminary Work 17 4 Results 19 4.1 A screen for novel regulators in Drosophila embryonic morphogenesis . -

Modular and Distinct Plexin-A4/FARP2/Rac1 Signaling Controls Dendrite Morphogenesis
The Journal of Neuroscience, July 8, 2020 • 40(28):5413–5430 • 5413 Development/Plasticity/Repair Modular and Distinct Plexin-A4/FARP2/Rac1 Signaling Controls Dendrite Morphogenesis Victor Danelon,1* Ron Goldner,2* Edward Martinez,1 Irena Gokhman,2 Kimberly Wang,1 Avraham Yaron,2 and Tracy S. Tran1 1Department of Biological Sciences, Rutgers University, Newark, New Jersey 07102, and 2Department of Biomolecular Sciences, Weizmann Institute of Science, Rehovot, 76100, Israel Diverse neuronal populations with distinct cellular morphologies coordinate the complex function of the nervous system. Establishment of distinct neuronal morphologies critically depends on signaling pathways that control axonal and dendritic devel- opment. The Sema3A-Nrp1/PlxnA4 signaling pathway promotes cortical neuron basal dendrite arborization but also repels axons. However, the downstream signaling components underlying these disparate functions of Sema3A signaling are unclear. Using the KRK-AAA novel PlxnA4 knock-in male and female mice, generated by CRISPR/cas9, we show here that the KRK motif in the PlxnA4 cytoplasmic domain is required for Sema3A-mediated cortical neuron dendritic elaboration but is dispensable for inhibitory axon guidance. The RhoGEF FARP2, which binds to the KRK motif, shows identical functional specificity as the KRK motif in the PlxnA4 receptor. We find that Sema3A activates the small GTPase Rac1, and that Rac1 activity is required for dendrite elaboration but not axon growth cone collapse. This work identifies a novel Sema3A-Nrp1/PlxnA4/FARP2/Rac1 signaling pathway that specifi- cally controls dendritic morphogenesis but is dispensable for repulsive guidance events. Overall, our results demonstrate that the divergent signaling output from multifunctional receptor complexes critically depends on distinct signaling motifs, highlighting the modular nature of guidance cue receptors and its potential to regulate diverse cellular responses. -

A Draft Map of the Human Proteome
ARTICLE doi:10.1038/nature13302 A draft map of the human proteome Min-Sik Kim1,2, Sneha M. Pinto3, Derese Getnet1,4, Raja Sekhar Nirujogi3, Srikanth S. Manda3, Raghothama Chaerkady1,2, Anil K. Madugundu3, Dhanashree S. Kelkar3, Ruth Isserlin5, Shobhit Jain5, Joji K. Thomas3, Babylakshmi Muthusamy3, Pamela Leal-Rojas1,6, Praveen Kumar3, Nandini A. Sahasrabuddhe3, Lavanya Balakrishnan3, Jayshree Advani3, Bijesh George3, Santosh Renuse3, Lakshmi Dhevi N. Selvan3, Arun H. Patil3, Vishalakshi Nanjappa3, Aneesha Radhakrishnan3, Samarjeet Prasad1, Tejaswini Subbannayya3, Rajesh Raju3, Manish Kumar3, Sreelakshmi K. Sreenivasamurthy3, Arivusudar Marimuthu3, Gajanan J. Sathe3, Sandip Chavan3, Keshava K. Datta3, Yashwanth Subbannayya3, Apeksha Sahu3, Soujanya D. Yelamanchi3, Savita Jayaram3, Pavithra Rajagopalan3, Jyoti Sharma3, Krishna R. Murthy3, Nazia Syed3, Renu Goel3, Aafaque A. Khan3, Sartaj Ahmad3, Gourav Dey3, Keshav Mudgal7, Aditi Chatterjee3, Tai-Chung Huang1, Jun Zhong1, Xinyan Wu1,2, Patrick G. Shaw1, Donald Freed1, Muhammad S. Zahari2, Kanchan K. Mukherjee8, Subramanian Shankar9, Anita Mahadevan10,11, Henry Lam12, Christopher J. Mitchell1, Susarla Krishna Shankar10,11, Parthasarathy Satishchandra13, John T. Schroeder14, Ravi Sirdeshmukh3, Anirban Maitra15,16, Steven D. Leach1,17, Charles G. Drake16,18, Marc K. Halushka15, T. S. Keshava Prasad3, Ralph H. Hruban15,16, Candace L. Kerr19{, Gary D. Bader5, Christine A. Iacobuzio-Donahue15,16,17, Harsha Gowda3 & Akhilesh Pandey1,2,3,4,15,16,20 The availability of human genome sequence has transformed biomedical research over the past decade. However, an equiv- alent map for the human proteome with direct measurements of proteins and peptides does not exist yet. Here we present a draft map of the human proteome using high-resolution Fourier-transform mass spectrometry. -

MAP17's Up-Regulation, a Crosspoint in Cancer and Inflammatory
García-Heredia and Carnero Molecular Cancer (2018) 17:80 https://doi.org/10.1186/s12943-018-0828-7 REVIEW Open Access Dr. Jekyll and Mr. Hyde: MAP17’s up- regulation, a crosspoint in cancer and inflammatory diseases José M. García-Heredia1,2,3 and Amancio Carnero1,3* Abstract Inflammation is a common defensive response that is activated after different harmful stimuli. This chronic, or pathological, inflammation is also one of the causes of neoplastic transformation and cancer development. MAP17 is a small protein localized to membranes with a restricted pattern of expression in adult tissues. However, its expression is common in destabilized cells, as it is overexpressed both in inflammatory diseases and in cancer. MAP17 is overexpressed in most, if not all, carcinomas and in many tumors of mesenchymal origin, and correlates with higher grade and poorly differentiated tumors. This overexpression drives deep changes in cell homeostasis including increased oxidative stress, deregulation of signaling pathways and increased growth rates. Importantly, MAP17 is associated in tumors with inflammatory cells infiltration, not only in cancer but in various inflammatory diseases such as Barret’s esophagus, lupus, Crohn’s, psoriasis and COPD. Furthermore, MAP17 also modifies the expression of genes connected to inflammation, showing a clear induction of the inflammatory profile. Since MAP17 appears highly correlated with the infiltration of inflammatory cells in cancer, is MAP17 overexpression an important cellular event connecting tumorigenesis and inflammation? Keywords: MAP17, Cancer, Inflammatory diseases Background process ends when the activated cells undergo apoptosis in Inflammatory response is a common defensive process acti- a highly regulated process that finishes after pathogens and vated after different harmful stimuli, constituting a highly cell debris have been phagocytized [9]. -

Genome-Wide DNA Methylation Dynamics During Epigenetic
Gómez‑Redondo et al. Clin Epigenet (2021) 13:27 https://doi.org/10.1186/s13148‑021‑01003‑x RESEARCH Open Access Genome‑wide DNA methylation dynamics during epigenetic reprogramming in the porcine germline Isabel Gómez‑Redondo1*† , Benjamín Planells1†, Sebastián Cánovas2,3, Elena Ivanova4, Gavin Kelsey4,5 and Alfonso Gutiérrez‑Adán1 Abstract Background: Prior work in mice has shown that some retrotransposed elements remain substantially methylated during DNA methylation reprogramming of germ cells. In the pig, however, information about this process is scarce. The present study was designed to examine the methylation profles of porcine germ cells during the time course of epigenetic reprogramming. Results: Sows were artifcially inseminated, and their fetuses were collected 28, 32, 36, 39, and 42 days later. At each time point, genital ridges were dissected from the mesonephros and germ cells were isolated through magnetic‑ activated cell sorting using an anti‑SSEA‑1 antibody, and recovered germ cells were subjected to whole‑genome bisulphite sequencing. Methylation levels were quantifed using SeqMonk software by performing an unbiased analysis, and persistently methylated regions (PMRs) in each sex were determined to extract those regions showing 50% or more methylation. Most genomic elements underwent a dramatic loss of methylation from day 28 to day 36, when the lowest levels were shown. By day 42, there was evidence for the initiation of genomic re‑methylation. We identifed a total of 1456 and 1122 PMRs in male and female germ cells, respectively, and large numbers of transpos‑ able elements (SINEs, LINEs, and LTRs) were found to be located within these PMRs. Twenty‑one percent of the introns located in these PMRs were found to be the frst introns of a gene, suggesting their regulatory role in the expression of these genes. -

Genome-Wide Detection of Natural Selection in African Americans Pre- and Post-Admixture
Downloaded from genome.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Method Genome-wide detection of natural selection in African Americans pre- and post-admixture Wenfei Jin,1 Shuhua Xu,1,6 Haifeng Wang,2 Yongguo Yu,3 Yiping Shen,4,5 Bailin Wu,4,5 and Li Jin1,4,6 1Chinese Academy of Sciences Key Laboratory of Computational Biology, Chinese Academy of Sciences and Max Planck Society (CAS-MPG) Partner Institute for Computational Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China; 2Chinese National Human Genome Center, Shanghai 201203, China; 3Shanghai Children’s Medical Center, Shanghai Jiaotong University School of Medicine, Shanghai 200127, China; 4Ministry of Education (MOE) Key Laboratory of Contemporary Anthropology, School of Life Sciences and Institutes of Biomedical Sciences, Fudan University, Shanghai 200433, China; 5Children’s Hospital Boston, Harvard Medical School, Boston, Massachusetts 02115, USA It is particularly meaningful to investigate natural selection in African Americans (AfA) due to the high mortality their African ancestry has experienced in history. In this study, we examined 491,526 autosomal single nucleotide poly- morphisms (SNPs) genotyped in 5210 individuals and conducted a genome-wide search for selection signals in 1890 AfA. Several genomic regions showing an excess of African or European ancestry, which were considered the footprints of selection since population admixture, were detected based on a commonly used approach. However, we also developed a new strategy to detect natural selection both pre- and post-admixture by reconstructing an ancestral African population (AAF) from inferred African components of ancestry in AfA and comparing it with indigenous African populations (IAF). -

Egfr Activates a Taz-Driven Oncogenic Program in Glioblastoma
EGFR ACTIVATES A TAZ-DRIVEN ONCOGENIC PROGRAM IN GLIOBLASTOMA by Minling Gao A thesis submitted to Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland March 2020 ©2020 Minling Gao All rights reserved Abstract Hyperactivated EGFR signaling is associated with about 45% of Glioblastoma (GBM), the most aggressive and lethal primary brain tumor in humans. However, the oncogenic transcriptional events driven by EGFR are still incompletely understood. We studied the role of the transcription factor TAZ to better understand master transcriptional regulators in mediating the EGFR signaling pathway in GBM. The transcriptional coactivator with PDZ- binding motif (TAZ) and its paralog gene, the Yes-associated protein (YAP) are two transcriptional co-activators that play important roles in multiple cancer types and are regulated in a context-dependent manner by various upstream signaling pathways, e.g. the Hippo, WNT and GPCR signaling. In GBM cells, TAZ functions as an oncogene that drives mesenchymal transition and radioresistance. This thesis intends to broaden our understanding of EGFR signaling and TAZ regulation in GBM. In patient-derived GBM cell models, EGF induced TAZ and its known gene targets through EGFR and downstream tyrosine kinases (ERK1/2 and STAT3). In GBM cells with EGFRvIII, an EGF-independent and constitutively active mutation, TAZ showed EGF- independent hyperactivation when compared to EGFRvIII-negative cells. These results revealed a novel EGFR-TAZ signaling axis in GBM cells. The second contribution of this thesis is that we performed next-generation sequencing to establish the first genome-wide map of EGF-induced TAZ target genes. -
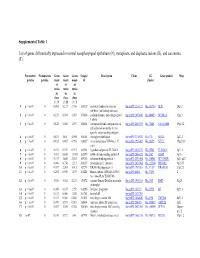
Supplemental Table 1 List of Genes Differentially Expressed In
Supplemental Table 1 List of genes differentially expressed in normal nasopharyngeal epithelium (N), metaplastic and displastic lesions (R), and carcinoma (T). Parametric Permutation Geom Geom Geom Unique Description Clone UG Gene symbol Map p-value p-value mean mean mean id cluster of of of ratios ratios ratios in in in class class class 1 : N 2 : R 3 : T 1 p < 1e-07 0 0.061 0.123 2.708 169329 secretory leukocyte protease IncytePD:2510171 Hs.251754 SLPI 20q12 inhibitor (antileukoproteinase) 2 p < 1e-07 0 0.125 0.394 1.863 163628 sodium channel, nonvoltage-gated IncytePD:1453049 Hs.446415 SCNN1A 12p13 1 alpha 3 p < 1e-07 0 0.122 0.046 1.497 160401 carcinoembryonic antigen-related IncytePD:2060355 Hs.73848 CEACAM6 19q13.2 cell adhesion molecule 6 (non- specific cross reacting antigen) 4 p < 1e-07 0 0.675 1.64 5.594 165101 monoglyceride lipase IncytePD:2174920 Hs.6721 MGLL 3q21.3 5 p < 1e-07 0 0.182 0.487 0.998 166827 nei endonuclease VIII-like 1 (E. IncytePD:1926409 Hs.28355 NEIL1 15q22.33 coli) 6 p < 1e-07 0 0.194 0.339 0.915 162931 hypothetical protein FLJ22418 IncytePD:2816379 Hs.36563 FLJ22418 1p11.1 7 p < 1e-07 0 1.313 0.645 13.593 162399 S100 calcium binding protein P IncytePD:2060823 Hs.2962 S100P 4p16 8 p < 1e-07 0 0.157 1.445 2.563 169315 selenium binding protein 1 IncytePD:2591494 Hs.334841 SELENBP1 1q21-q22 9 p < 1e-07 0 0.046 0.738 1.213 160115 prominin-like 1 (mouse) IncytePD:2070568 Hs.112360 PROML1 4p15.33 10 p < 1e-07 0 0.787 2.264 3.013 167294 HRAS-like suppressor 3 IncytePD:1969263 Hs.37189 HRASLS3 11q12.3 11 p < 1e-07 0 0.292 0.539 1.493 168221 Homo sapiens cDNA FLJ13510 IncytePD:64451 Hs.37896 2 fis, clone PLACE1005146. -

Prenatal Detection of TAR Syndrome in a Fetus with Compound Inheritance of an RBM8A SNP and a 334‑Kb Deletion: a Case Report
MOLECULAR MEDICINE REPORTS 9: 163-165, 2014 Prenatal detection of TAR syndrome in a fetus with compound inheritance of an RBM8A SNP and a 334‑kb deletion: A case report IOANNIS PAPOULIDIS1, EIRINI OIKONOMIDOU1, SANDRO ORRU2, ELISAVET SIOMOU1, MARIA KONTODIOU1, MAKARIOS ELEFTHERIADES3, VASILIOS BACOULAS4, JUAN C. CIGUDOSA5, JAVIER SUELA5, LORETTA THOMAIDIS6 and EMMANOUIL MANOLAKOS1,2 1Laboratory of Genetics, Eurogenetica S.A., Thessaloniki 55133, Greece; 2Department of Medical Genetics, University of Cagliari, Binaghi Hospital, Cagliari I‑09126, Italy; 3Embryocare, Fetal Medicine Unit, Athens 11522; 4Fetal Medicine Centre, Athens 10674, Greece; 5NIMGenetics, Madrid 28049, Spain; 6Department of Pediatrics, Aglaia Kyriakou Children's Hospital, University of Athens, Athens 11527, Greece Received May 30, 2013; Accepted October 16, 2013 DOI: 10.3892/mmr.2013.1788 Abstract. Thrombocytopenia-absent radius syndrome identified the presence of a minimally deleted 200‑kb region (TAR) is a rare genetic disorder that is characterized by the at chromosome band 1q21.1 in patients with TAR, but it is not absence of the radius bone in each forearm and a markedly sufficient to cause the phenotype (3,4). A study identified two reduced platelet count that results in life-threatening bleeding rare single nucleotide polymorphisms (SNPs) in the regulatory episodes (thrombocytopenia). Tar syndrome has been associ- region of the RBM8A gene that are involved in TAR syndrome ated with a deletion of a segment of 1q21.1 cytoband. The through the reduction of the expression of the RBM8A-encoded 1q21.1 deletion syndrome phenotype includes Tar and other Y14 protein (4). The first allele (rs139428292 G>A), which is features such as mental retardation, autism and microcephaly. -
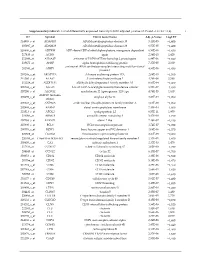
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Regulating Rac in the Nervous System: Molecular Function and Disease Implication of Rac Gefs and Gaps
Hindawi Publishing Corporation BioMed Research International Volume 2015, Article ID 632450, 17 pages http://dx.doi.org/10.1155/2015/632450 Review Article Regulating Rac in the Nervous System: Molecular Function and Disease Implication of Rac GEFs and GAPs Yanyang Bai, Xiaoliang Xiang, Chunmei Liang, and Lei Shi JNU-HKUST Joint Laboratory for Neuroscience and Innovative Drug Research, Jinan University, Guangzhou, Guangdong 510632, China Correspondence should be addressed to Lei Shi; [email protected] Received 23 January 2015; Accepted 6 March 2015 Academic Editor: Akito Tanoue Copyright © 2015 Yanyang Bai et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Rho family GTPases, including RhoA, Rac1, and Cdc42 as the most studied members, are master regulators of actin cytoskeletal organization. Rho GTPases control various aspects of the nervous system and are associated with a number of neuropsychiatric and neurodegenerative diseases. The activity of Rho GTPases is controlled by two families of regulators, guanine nucleotide exchange factors (GEFs) as the activators and GTPase-activating proteins (GAPs) as the inhibitors. Through coordinated regulation by GEFs and GAPs, Rho GTPases act as converging signaling molecules that convey different upstream signals in the nervous system. So far,morethan70membersofeitherGEFsorGAPsofRhoGTPaseshavebeenidentifiedinmammals,butonlyasmallsubset of them have well-known functions. Thus, characterization of important GEFs and GAPs in the nervous system is crucial for the understanding of spatiotemporal dynamics of Rho GTPase activity in different neuronal functions. In this review, we summarize the current understanding of GEFs and GAPs for Rac1, with emphasis on the molecular function and disease implication of these regulators in the nervous system.