Integrating Text Mining, Data Mining, and Network Analysis for Identifying
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Investigation of the Immune Receptors CEACAM3 and CEACAM4
Investigation of the human immune receptors CEACAM3 and CEACAM4 Dissertation Zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften (Dr. rer. nat.) vorgelegt von Julia Delgado Tascón An der Universität Konstanz des Fachbereichs Biologie Konstanz, Oktober 2015 Konstanzer Online-Publikations-System (KOPS) URL: http://nbn-resolving.de/urn:nbn:de:bsz:352-0-306516 Tag der mündlichen Prüfung: 05.11.2015 Vorsitzender und mündlicher Prüfer: Herr Professor Dr. Bürkle 1. Referent und und mündlicher Prüfer: Herr Professor Dr. Hauck 2. Referent und und mündlicher Prüfer: Herr Professor Dr. Tschan, Universität Bern A mi familia Acknowledgements I would like to express my special gratitude to my advisor Prof. Dr. Christof Hauck. His patient guidance and enthusiastic encouragement during these four years of PhD were a crucial aid to my process. I’m very thankful for his willingness and for granting me with his time in search for valuable and constructive suggestions during the planning and development of this research work. This certainly allowed me to grow as a person and as a scientist. I would also like to thank my committee members: to Prof. Dr. Mario Tschan for giving me his academic support at this last phase of my PhD thesis, and to Prof. M.Dr. Alexander Bürkle for his wise advices accompanied with Spanish greetings along this time. My thanks are extended to every member of the AG Hauck as well. To Anne, Susana, Petra and Claudia: thank you very much for your technical and personal guidance during these years. I’m also thankful to my fellow colleagues for countless ‘Kaffeepausen’ full of jokes, nice discussions, and delicious vegan cakes. -

IQSEC2 Antibody Cat
IQSEC2 Antibody Cat. No.: 8011 Western blot analysis of IQSEC2 in SK-N-SH cell lysate with IQSEC2 antibody at 1 μg/ml. Immunohistochemistry of IQSEC2 in mouse brain tissue with IQSEC2 antibody at 5 μg/ml. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human, Mouse, Rat IQSEC2 antibody was raised against an 18 amino acid peptide near the carboxy terminus of human IQSEC2. IMMUNOGEN: The immunogen is located within amino acids 1380 - 1430 of IQSEC2. TESTED APPLICATIONS: ELISA, IHC-P, WB IQSEC2 antibody can be used for detection of IQSEC2 by Western blot at 1 - 2 μg/ml. Antibody can also be used for immunohistochemistry starting at 5 μg/mL. APPLICATIONS: Antibody validated: Western Blot in human samples and Immunohistochemistry in mouse samples. All other applications and species not yet tested. September 27, 2021 1 https://www.prosci-inc.com/iqsec2-antibody-8011.html IQSEC2 antibody is human, mouse and rat reactive. At least two isoforms of IQSEC2 are SPECIFICITY: known to exist; this antibody will only detect the larger isoform. IQSEC2 antibody is predicted to not cross-react with IQSEC1. POSITIVE CONTROL: 1) Cat. No. 1220 - SK-N-SH Cell Lysate Predicted: 104, 133, 141, 164 kDa PREDICTED MOLECULAR WEIGHT: Observed: 140 kDa Properties PURIFICATION: IQSEC2 antibody is affinity chromatography purified via peptide column. CLONALITY: Polyclonal ISOTYPE: IgG CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: IQSEC2 antibody is supplied in PBS containing 0.02% sodium azide. CONCENTRATION: 1 mg/mL IQSEC2 antibody can be stored at 4˚C for three months and -20˚C, stable for up to one STORAGE CONDITIONS: year. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Integrative Differential Expression and Gene Set Enrichment Analysis Using Summary Statistics for Scrna-Seq Studies
ARTICLE https://doi.org/10.1038/s41467-020-15298-6 OPEN Integrative differential expression and gene set enrichment analysis using summary statistics for scRNA-seq studies ✉ Ying Ma 1,7, Shiquan Sun 1,7, Xuequn Shang2, Evan T. Keller 3, Mengjie Chen 4,5 & Xiang Zhou 1,6 Differential expression (DE) analysis and gene set enrichment (GSE) analysis are commonly applied in single cell RNA sequencing (scRNA-seq) studies. Here, we develop an integrative 1234567890():,; and scalable computational method, iDEA, to perform joint DE and GSE analysis through a hierarchical Bayesian framework. By integrating DE and GSE analyses, iDEA can improve the power and consistency of DE analysis and the accuracy of GSE analysis. Importantly, iDEA uses only DE summary statistics as input, enabling effective data modeling through com- plementing and pairing with various existing DE methods. We illustrate the benefits of iDEA with extensive simulations. We also apply iDEA to analyze three scRNA-seq data sets, where iDEA achieves up to five-fold power gain over existing GSE methods and up to 64% power gain over existing DE methods. The power gain brought by iDEA allows us to identify many pathways that would not be identified by existing approaches in these data. 1 Department of Biostatistics, University of Michigan, Ann Arbor, MI 48109, USA. 2 School of Computer Science, Northwestern Polytechnical University, Xi’an, Shaanxi 710072, P.R. China. 3 Department of Urology, University of Michigan, Ann Arbor, MI 48109, USA. 4 Department of Human Genetics, University of Chicago, Chicago, IL 60637, USA. 5 Section of Genetic Medicine, Department of Medicine, University of Chicago, Chicago, IL 60637, USA. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Product Datasheet
Antibody Technology Product Data Sheet anti-human CEACAM8 monoclonal antibody Product information Catalog Number: GM-0512 Clone: GM2H6 Description: purified monoclonal mouse antibody Specificity: anti-human CEACAM8 (CD66b/NCA-95) Isotype: IgG1 Purification: Protein G Storage: short term: 2°C - 8°C; long term: -20°C (avoid repeated freezing and thawing) Buffer : phosphate buffered saline, pH 7.2 Immunogen: genetic immunisation with cDNA encoding human CEACAM8 Selection: based on recognition of the complete native protein expressed on transfected mammalian cells Working dilutions Flow cytometry: 1.2 µg/106 cells CELISA: 1:200 - 1:400 For each application a titration should be performed to determine the optimal concentration. Specificity testing by flow cytometry CEACAM8 transfectant control transfectant Fig.1: FACS analysis of BOSC23 cells using GM2H6 Cat.# GM- 0512. BOSC23 cells were transiently transfected with an expression vector encoding either CEACAM8 (red curve) or an irrelevant protein (control transfectant). Binding of GM-0512 was detected with a PE -conjugated secondary antibody. A positive signal was obtained only with CEACAM8 transfected cells. For research use only. Not for diagnostic or therapeutic use. Aldevron Freiburg GmbH [email protected] Page 1 / 2 Waltershofener Str. 17 www.aldevron.com 79111 Freiburg, Germany Antibody Technology Antibody cross-reactivity with members of the CEA family CEACAM1 CEACAM3 CEACAM4 CEACAM5 CEACAM6 CEACAM7 CEACAM8 PSG1 Fig. 2: BOSC23 cells were transiently transfected with expression vectors containing either the cDNA of CEACAM1, CEACAM3-8 or PSG. The latter expressed as a membrane bound fusion protein. Expression of the constructs was tested with monoclonal antibodies known to recognize the corresponding proteins (CEACAM1,3,4,5 and 6: D14HD11; CEACAM7: BAC2; CEACAM8: Tet2; PSG: BAP3; green curves). -

CD Markers Are Routinely Used for the Immunophenotyping of Cells
ptglab.com 1 CD MARKER ANTIBODIES www.ptglab.com Introduction The cluster of differentiation (abbreviated as CD) is a protocol used for the identification and investigation of cell surface molecules. So-called CD markers are routinely used for the immunophenotyping of cells. Despite this use, they are not limited to roles in the immune system and perform a variety of roles in cell differentiation, adhesion, migration, blood clotting, gamete fertilization, amino acid transport and apoptosis, among many others. As such, Proteintech’s mini catalog featuring its antibodies targeting CD markers is applicable to a wide range of research disciplines. PRODUCT FOCUS PECAM1 Platelet endothelial cell adhesion of blood vessels – making up a large portion molecule-1 (PECAM1), also known as cluster of its intracellular junctions. PECAM-1 is also CD Number of differentiation 31 (CD31), is a member of present on the surface of hematopoietic the immunoglobulin gene superfamily of cell cells and immune cells including platelets, CD31 adhesion molecules. It is highly expressed monocytes, neutrophils, natural killer cells, on the surface of the endothelium – the thin megakaryocytes and some types of T-cell. Catalog Number layer of endothelial cells lining the interior 11256-1-AP Type Rabbit Polyclonal Applications ELISA, FC, IF, IHC, IP, WB 16 Publications Immunohistochemical of paraffin-embedded Figure 1: Immunofluorescence staining human hepatocirrhosis using PECAM1, CD31 of PECAM1 (11256-1-AP), Alexa 488 goat antibody (11265-1-AP) at a dilution of 1:50 anti-rabbit (green), and smooth muscle KD/KO Validated (40x objective). alpha-actin (red), courtesy of Nicola Smart. PECAM1: Customer Testimonial Nicola Smart, a cardiovascular researcher “As you can see [the immunostaining] is and a group leader at the University of extremely clean and specific [and] displays Oxford, has said of the PECAM1 antibody strong intercellular junction expression, (11265-1-AP) that it “worked beautifully as expected for a cell adhesion molecule.” on every occasion I’ve tried it.” Proteintech thanks Dr. -
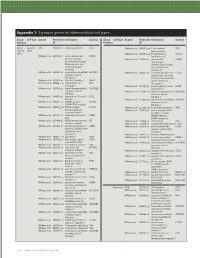
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Expressed Gene Fusions As Frequent Drivers of Poor Outcomes in Hormone Receptor–Positive Breast Cancer
Published OnlineFirst December 14, 2017; DOI: 10.1158/2159-8290.CD-17-0535 RESEARCH ARTICLE Expressed Gene Fusions as Frequent Drivers of Poor Outcomes in Hormone Receptor–Positive Breast Cancer Karina J. Matissek1,2, Maristela L. Onozato3, Sheng Sun1,2, Zongli Zheng2,3,4, Andrew Schultz1, Jesse Lee3, Kristofer Patel1, Piiha-Lotta Jerevall2,3, Srinivas Vinod Saladi1,2, Allison Macleay3, Mehrad Tavallai1,2, Tanja Badovinac-Crnjevic5, Carlos Barrios6, Nuran Beşe7, Arlene Chan8, Yanin Chavarri-Guerra9, Marcio Debiasi6, Elif Demirdögen10, Ünal Egeli10, Sahsuvar Gökgöz10, Henry Gomez11, Pedro Liedke6, Ismet Tasdelen10, Sahsine Tolunay10, Gustavo Werutsky6, Jessica St. Louis1, Nora Horick12, Dianne M. Finkelstein2,12, Long Phi Le2,3, Aditya Bardia1,2, Paul E. Goss1,2, Dennis C. Sgroi2,3, A. John Iafrate2,3, and Leif W. Ellisen1,2 ABSTRACT We sought to uncover genetic drivers of hormone receptor–positive (HR+) breast cancer, using a targeted next-generation sequencing approach for detecting expressed gene rearrangements without prior knowledge of the fusion partners. We identified inter- genic fusions involving driver genes, including PIK3CA, AKT3, RAF1, and ESR1, in 14% (24/173) of unselected patients with advanced HR+ breast cancer. FISH confirmed the corresponding chromo- somal rearrangements in both primary and metastatic tumors. Expression of novel kinase fusions in nontransformed cells deregulates phosphoprotein signaling, cell proliferation, and survival in three- dimensional culture, whereas expression in HR+ breast cancer models modulates estrogen-dependent growth and confers hormonal therapy resistance in vitro and in vivo. Strikingly, shorter overall survival was observed in patients with rearrangement-positive versus rearrangement-negative tumors. Cor- respondingly, fusions were uncommon (<5%) among 300 patients presenting with primary HR+ breast cancer. -

CEACAM3: an Innate Immune Receptor Directed Against Human-Restricted Bacterial Pathogens Stefan Pilsa, Dave T
ARTICLE IN PRESS International Journal of Medical Microbiology 298 (2008) 553–560 www.elsevier.de/ijmm CEACAM3: An innate immune receptor directed against human-restricted bacterial pathogens Stefan Pilsa, Dave T. Gerrardb,1, Axel Meyerb, Christof R. Haucka,Ã aLehrstuhl Zellbiologie, Fachbereich Biologie X908, Universita¨t Konstanz, Universita¨tsstrasse 10, 78457 Konstanz, Germany bLehrstuhl fu¨r Zoologie und Evolutionsbiologie, Fachbereich Biologie, Universita¨t Konstanz, Germany Received 15 January 2008; accepted 4 April 2008 Abstract Carcinoembryonic antigen-related cell adhesion molecule 3 (CEACAM3) is an immunoglobulin-related glycoprotein exclusively expressed on granulocytes. In contrast to other members of the CEACAM family, CEACAM3 does not support cell–cell adhesion, but rather mediates the opsonin-independent recognition and elimination of a restricted set of human-specific Gram-negative bacterial pathogens including Neisseria gonorrhoeae, Haemophilus influenzae,andMoraxella catarrhalis. Within the last 4 years, molecular determinants of CEACAM3 function and CEACAM3-initiated signaling pathways have been elucidated. Sequence comparison between CEACAM3 and other CEACAM family members points to a chimeric origin of this receptor with the bacteria- binding extracellular domain and the function-promoting intracellular domain derived from different genes. This review summarizes the current knowledge about the structure–function relationship of CEACAM3 and tries to combine these molecular aspects with a plausible scenario concerning the evolutionary origin of this phagocyte receptor in the light of host–pathogen adaptation. r 2008 Published by Elsevier GmbH. Keywords: ITAM; Signal transduction; Phagocytosis; Host–pathogen coevolution Introduction main (Beauchemin et al., 1999; Kuespert et al., 2006). In humans, the CEACAM family comprises 12 closely CEACAM3 belongs to the family of carcinoembry- related proteins, with several family members expressed onic antigen (CEA)-related cell adhesion molecules by epithelial cells (e.g. -

The Role of the Actin Cytoskeleton During Muscle Development In
THE ROLE OF THE ACTIN CYTOSKELETON DURING MUSCLE DEVELOPMENT IN DROSOPHILA AND MOUSE by Shannon Faye Yu A Dissertation Presented to the Faculty of the Louis V. Gerstner, Jr. Graduate School of the Biomedical Sciences in Partial Fulfillment of the Requirements of the Degree of Doctor of Philosophy New York, NY Oct, 2013 Mary K. Baylies, PhD! Date Dissertation Mentor Copyright by Shannon F. Yu 2013 ABSTRACT The actin cytoskeleton is essential for many processes within a developing organism. Unsurprisingly, actin and its regulators underpin many of the critical steps in the formation and function of muscle tissue. These include cell division during the specification of muscle progenitors, myoblast fusion, muscle elongation and attachment, and muscle maturation, including sarcomere assembly. Analysis in Drosophila has focused on regulators of actin polymerization particularly during myoblast fusion, and the conservation of many of the actin regulators required for muscle development has not yet been tested. In addition, dynamic actin processes also require the depolymerization of existing actin fibers to replenish the pool of actin monomers available for polymerization. Despite this, the role of actin depolymerization has not been described in depth in Drosophila or mammalian muscle development. ! Here, we first examine the role of the actin depolymerization factor Twinstar (Tsr) in muscle development in Drosophila. We show that Twinstar, the sole Drosophila member of the ADF/cofilin family of actin depolymerization proteins, is expressed in muscle where it is essential for development. tsr mutant embryos displayed a number of muscle defects, including muscle loss and muscle misattachment. Further, regulators of Tsr, including a Tsr-inactivating kinase, Center divider, a Tsr-activating phosphatase, Slingshot and a synergistic partner in depolymerization, Flare, are also required for embryonic muscle development.