JHG52-10.Pdf
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

A New Model for X-Linked Tremor/Ataxia
© 2016. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2016) 9, 553-562 doi:10.1242/dmm.022848 RESEARCH ARTICLE Spontaneous shaker rat mutant – a new model for X-linked tremor/ ataxia Karla P. Figueroa1, Sharan Paul1, Tito Calì2, Raffaele Lopreiato2, Sukanya Karan1, Martina Frizzarin2, Darren Ames3, Ginevra Zanni4, Marisa Brini5, Warunee Dansithong1, Brett Milash3, Daniel R. Scoles1, Ernesto Carafoli6 and Stefan M. Pulst1,* ABSTRACT mode of inheritance. Here, we describe the genetic analysis of the The shaker rat is an X-linked recessive spontaneous model of shaker rat, a model of Purkinje cell (PC) degeneration. This mutant progressive Purkinje cell (PC) degeneration exhibiting a shaking arose spontaneously and was observed in Sprague Dawley (SD) ataxia and wide stance. Generation of Wistar Furth (WF)/Brown outbred stock in 1991 at Saint Louis University, first described by Norwegian (BN) F1 hybrids and genetic mapping of F2 sib-sib La Regina et al. (1992), and the phenotype of whole-body tremor, ‘ ’ offspring using polymorphic markers narrowed the candidate gene ataxia and wide stance designated as shaker . The shaker trait was region to 26 Mbp denoted by the last recombinant genetic marker reported as an X-linked recessive trait. DXRat21 at 133 Mbp to qter (the end of the long arm). In the WF Various animal models of spontaneously occurring mutants that background, the shaker mutation has complete penetrance, results in parallel some aspects of human hereditary ataxia have been a stereotypic phenotype and there is a narrow window for age of reported; for example, weaver, lurcher, stumbler, tottering and disease onset; by contrast, the F2 hybrid phenotype was more varied, teetering mice (Chou et al., 1991; Frankel et al., 1994; Green and with a later age of onset and likely non-penetrance of the mutation. -

Metastatic Adrenocortical Carcinoma Displays Higher Mutation Rate and Tumor Heterogeneity Than Primary Tumors
ARTICLE DOI: 10.1038/s41467-018-06366-z OPEN Metastatic adrenocortical carcinoma displays higher mutation rate and tumor heterogeneity than primary tumors Sudheer Kumar Gara1, Justin Lack2, Lisa Zhang1, Emerson Harris1, Margaret Cam2 & Electron Kebebew1,3 Adrenocortical cancer (ACC) is a rare cancer with poor prognosis and high mortality due to metastatic disease. All reported genetic alterations have been in primary ACC, and it is 1234567890():,; unknown if there is molecular heterogeneity in ACC. Here, we report the genetic changes associated with metastatic ACC compared to primary ACCs and tumor heterogeneity. We performed whole-exome sequencing of 33 metastatic tumors. The overall mutation rate (per megabase) in metastatic tumors was 2.8-fold higher than primary ACC tumor samples. We found tumor heterogeneity among different metastatic sites in ACC and discovered recurrent mutations in several novel genes. We observed 37–57% overlap in genes that are mutated among different metastatic sites within the same patient. We also identified new therapeutic targets in recurrent and metastatic ACC not previously described in primary ACCs. 1 Endocrine Oncology Branch, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA. 2 Center for Cancer Research, Collaborative Bioinformatics Resource, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA. 3 Department of Surgery and Stanford Cancer Institute, Stanford University, Stanford, CA 94305, USA. Correspondence and requests for materials should be addressed to E.K. (email: [email protected]) NATURE COMMUNICATIONS | (2018) 9:4172 | DOI: 10.1038/s41467-018-06366-z | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/s41467-018-06366-z drenocortical carcinoma (ACC) is a rare malignancy with types including primary ACC from the TCGA to understand our A0.7–2 cases per million per year1,2. -

The Conserved DNMT1-Dependent Methylation Regions in Human Cells
Freeman et al. Epigenetics & Chromatin (2020) 13:17 https://doi.org/10.1186/s13072-020-00338-8 Epigenetics & Chromatin RESEARCH Open Access The conserved DNMT1-dependent methylation regions in human cells are vulnerable to neurotoxicant rotenone exposure Dana M. Freeman1 , Dan Lou1, Yanqiang Li1, Suzanne N. Martos1 and Zhibin Wang1,2,3* Abstract Background: Allele-specifc DNA methylation (ASM) describes genomic loci that maintain CpG methylation at only one inherited allele rather than having coordinated methylation across both alleles. The most prominent of these regions are germline ASMs (gASMs) that control the expression of imprinted genes in a parent of origin-dependent manner and are associated with disease. However, our recent report reveals numerous ASMs at non-imprinted genes. These non-germline ASMs are dependent on DNA methyltransferase 1 (DNMT1) and strikingly show the feature of random, switchable monoallelic methylation patterns in the mouse genome. The signifcance of these ASMs to human health has not been explored. Due to their shared allelicity with gASMs, herein, we propose that non-tradi- tional ASMs are sensitive to exposures in association with human disease. Results: We frst explore their conservancy in the human genome. Our data show that our putative non-germline ASMs were in conserved regions of the human genome and located adjacent to genes vital for neuronal develop- ment and maturation. We next tested the hypothesized vulnerability of these regions by exposing human embryonic kidney cell HEK293 with the neurotoxicant rotenone for 24 h. Indeed,14 genes adjacent to our identifed regions were diferentially expressed from RNA-sequencing. We analyzed the base-resolution methylation patterns of the predicted non-germline ASMs at two neurological genes, HCN2 and NEFM, with potential to increase the risk of neurodegenera- tion. -
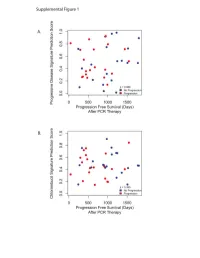
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Chapter 15: Disease Gene Prioritization
Education Chapter 15: Disease Gene Prioritization Yana Bromberg* Department of Biochemistry and Microbiology, School of Environmental and Biological Sciences, Rutgers University, New Brunswick, New Jersey, United States of America Abstract: Disease-causing aberra- chromosome 4 called huntigtin (HTT) 2. Background tions in the normal function of a [4]. Huntington’s became the first genetic The Merriam-Webster dictionary de- gene define that gene as a disease disease mapped using polymorphism fines the word ‘‘disease’’ as a ‘‘a condition gene. Proving a causal link be- information (G8 DNA probe/genetic tween a gene and a disease exper- marker), closely followed by the same of the living animal or plant body or of one imentally is expensive and time- year discovery of phenylketonuria associ- of its parts that impairs normal functioning consuming. Comprehensive priori- ation with polymorphisms in a hepatic and is typically manifested by distinguishing tization of candidate genes prior to enzyme phenylalanine hydroxylase [5]. signs and symptoms.’’ Thus, disease is experimental testing drastically re- These advances provided a route for defined with respect to normal function of said duces the associated costs. Com- predicting the likelihood of disease devel- body or body part. Note, that this definition putational gene prioritization is opment and even stirred some worries also describes the malfunction of individual based on various pieces of correl- regarding the possibility of the rise of cells or cell groups. In fact, many diseases ative evidence that associate each ‘‘medical eugenics’’ [6]. Interestingly, it can and should be defined on a cellular gene with the given disease and took another ten years for HTT’s se- level. -

Host Genome Analysis of Structural Variations by Optical
medRxiv preprint doi: https://doi.org/10.1101/2021.01.05.21249190; this version posted January 8, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Host genome analysis of structural variations by Optical Genome Mapping provides clinically valuable insights into genes implicated in critical immune, viral infection, and viral replication pathways in patients with severe COVID-19. Nikhil Shri Sahajpal1, Chi-Yu Jill Lai2, Alex Hastie2, Ashis K Mondal1, Siavash Raeisi Dehkordi3, Cas van der Made4, Olivier Fedrigo5, Farooq Al-Ajli5, Sawan Jalnapurkar6, Rashmi Kanagal- Shamanna7, Brynn Levy8, Silviu-Alin Bacanu9, Michael C Zody10, Catherine A. Brownstein11, Amyn M. Rojiani1, Alan H. Beggs11, Vineet Bafna3, Alexander Hoischen4, Erich D. Jarvis5,12,13, Alka Chaubey1,2, Ravindra Kolhe1* and the COVID19hostgenomesv consortium†. 1 Department of Pathology, Medical College of Georgia, Augusta University, GA, U.S.A. 2 Bionano Genomics, Inc., San Diego, CA, U.S.A. 3 Department of Computer Science and Engineering, University of California at San Diego, CA, U.S.A. 4 Department of Human Genetics, Radboud University Medical Center for Infectious Diseases (RCI), Department of Internal Medicine, Radboud Institute for Molecular Life Sciences, Radboud Expertise Center for Immunodeficiency and Autoinflammation, Radboud University Medical Center, Nijmegen, the Netherlands. 5 Vertebrate Genome Lab, The Rockefeller University, New York, NY, USA. 6 Department of Medicine, Medical College of Georgia, Augusta University, GA, U.S.A 7 Hematopathology, University of Texas M.D. -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

Genome-Scale Capture C Promoter Interactions Implicate Effector Genes at GWAS Loci for Bone Mineral Density
ARTICLE https://doi.org/10.1038/s41467-019-09302-x OPEN Genome-scale Capture C promoter interactions implicate effector genes at GWAS loci for bone mineral density Alessandra Chesi1, Yadav Wagley2, Matthew E. Johnson1, Elisabetta Manduchi1,3, Chun Su1, Sumei Lu1, Michelle E. Leonard1, Kenyaita M. Hodge1, James A. Pippin 1, Kurt D. Hankenson 2, Andrew D. Wells1,4 & Struan F.A. Grant 1,5,6 1234567890():,; Osteoporosis is a devastating disease with an essential genetic component. GWAS have discovered genetic signals robustly associated with bone mineral density (BMD), but not the precise localization of effector genes. Here, we carry out physical and direct variant to gene mapping in human mesenchymal progenitor cell-derived osteoblasts employing a massively parallel, high resolution Capture C based method in order to simultaneously characterize the genome-wide interactions of all human promoters. By intersecting our Capture C and ATAC- seq data, we observe consistent contacts between candidate causal variants and putative target gene promoters in open chromatin for ~ 17% of the 273 BMD loci investigated. Knockdown of two novel implicated genes, ING3 at ‘CPED1-WNT16’ and EPDR1 at ‘STARD3NL’, inhibits osteoblastogenesis, while promoting adipogenesis. This approach therefore aids target discovery in osteoporosis, here on the example of two relevant genes involved in the fate determination of mesenchymal progenitors, and can be applied to other common genetic diseases. 1 Center for Spatial and Functional Genomics, Children’s Hospital of Philadelphia, Philadelphia 19104 PA, USA. 2 Department of Orthopaedic Surgery, University of Michigan Medical School, Ann Arbor 48109 MI, USA. 3 Institute for Biomedical Informatics, University of Pennsylvania Perelman School of Medicine, Philadelphia 19104 PA, USA. -

Mitotic Checkpoints and Chromosome Instability Are Strong Predictors of Clinical Outcome in Gastrointestinal Stromal Tumors
MITOTIC CHECKPOINTS AND CHROMOSOME INSTABILITY ARE STRONG PREDICTORS OF CLINICAL OUTCOME IN GASTROINTESTINAL STROMAL TUMORS. Pauline Lagarde1,2, Gaëlle Pérot1, Audrey Kauffmann3, Céline Brulard1, Valérie Dapremont2, Isabelle Hostein2, Agnès Neuville1,2, Agnieszka Wozniak4, Raf Sciot5, Patrick Schöffski4, Alain Aurias1,6, Jean-Michel Coindre1,2,7 Maria Debiec-Rychter8, Frédéric Chibon1,2. Supplemental data NM cases deletion frequency. frequency. deletion NM cases Mand between difference the highest setswith of theprobe a view isdetailed panel Bottom frequently. sorted totheless deleted theprobe are frequently from more and thefrequency deletion represent Yaxes inblue. are cases (NM) metastatic for non- frequencies Corresponding inmetastatic (red). probe (M)cases sets figureSupplementary 1: 100 100 20 40 60 80 20 40 60 80 0 0 chr14 1 chr14 88 chr14 175 chr14 262 chr9 -MTAP 349 chr9 -MTAP 436 523 chr9-CDKN2A 610 Histogram presenting the 2000 more frequently deleted deleted frequently the 2000 more presenting Histogram chr9-CDKN2A 697 chr9-CDKN2A 784 chr9-CDKN2B 871 chr9-CDKN2B 958 chr9-CDKN2B 1045 chr22 1132 chr22 1219 chr22 1306 chr22 1393 1480 1567 M NM 1654 1741 1828 1915 M NM GIST14 GIST2 GIST16 GIST3 GIST19 GIST63 GIST9 GIST38 GIST61 GIST39 GIST56 GIST37 GIST47 GIST58 GIST28 GIST5 GIST17 GIST57 GIST47 GIST58 GIST28 GIST5 GIST17 GIST57 CDKN2A Supplementary figure 2: Chromosome 9 genomic profiles of the 18 metastatic GISTs (upper panel). Deletions and gains are indicated in green and red, respectively; and color intensity is proportional to copy number changes. A detailed view is given (bottom panel) for the 6 cases presenting a homozygous 9p21 deletion targeting CDKN2A locus (dark green). -

Cognitive Dysfunction in Spinocerebellar Ataxias
Dementia & Neuropsychologia 2009 September;3(3):180-187 Views & Reviews Cognitive dysfunction in spinocerebellar ataxias Helio Afonso Ghizoni Teive 1, Walter Oleschko Arruda1 Abstract – Spinocerebellar ataxias (SCAs) comprise a heterogeneous group of complex neurodegenerative diseases, characterized by the presence of progressive cerebellar ataxia, associated or otherwise with ophthalmoplegia, pyramidal signs, extrapyramidal features, pigmentary retinopathy, peripheral neuropathy, cognitive dysfunction and dementia. Objective: To verify the presence of cognitive dysfunction among the main types of SCA described in the literature. Methods: the review was conducted using the search system of the PUBMED and OMIM databases. Results: Cognitive dysfunction occurs in a considerable proportion of SCA, particularly in SCA 3, which is the most frequent form of SCA worldwide. Dementia has been described in several other types of SCA such as SCA 2, SCA 17 and DRPLA. Mental retardation is a specific clinical feature of SCA 13. Conclusions: The role of the cerebellum in cognitive functions has been observed in different types of SCAs which can manifest varying degrees of cognitive dysfunction, dementia and mental retardation. Key words: spinocerebellar ataxia, cognitive dysfunction, dementia. Disfunção cognitiva em ataxias espinocerebelares Resumo – Fundamento: As ataxias espinocerebelares (AECs) compreendem um grupo heterogêneo de enfermidades neurodegenerativas, que se caracterizam pela presença de ataxia cerebelar progressiva, associada de forma avariada com oftalmoplegia, sinais piramidais, extrapiramidais, retinopatia pigmentar, neuropatia periférica, disfunção cognitiva e demência. Objetivo: Revisar a presença de disfunção cognitiva entre os principais tipos de AECs publicados na literatura mundial. Métodos: A revisão foi realizada através da pesquisa pelo sistema do PUBMED e do OMIM. Resultados: Disfunção cognitiva ocorre em uma parcela considerável de AECs, particularmente na AEC 3, que é a forma de AEC mais encontrada em todo o mundo. -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein -

Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name pleckstrin homology domain containing, family G (with RhoGef domain) member 4 Gene Symbol PLEKHG4 Organism Human Gene Summary The protein encoded by this gene contains multiple domains suggestive of a role in intracellular signaling and cytoskeleton dynamics at the Golgi apparatus. Mutations in this gene are associated with spinocerebellar ataxia 16q22-linked. Several alternatively spliced transcript variants differing only in the 5' UTR or encoding a different isoform have been found for this gene. Gene Aliases ARHGEF44, DKFZp434I216, PRTPHN1, SCA4 RefSeq Accession No. NC_000016.9, NT_010498.15, NG_008439.1 UniGene ID Hs.188781 Ensembl Gene ID ENSG00000196155 Entrez Gene ID 25894 Assay Information Unique Assay ID qHsaCED0002921 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000360461, ENST00000427155, ENST00000379344, ENST00000450733, ENST00000393966 Amplicon Context Sequence TCTGAGCCCGGGACTGGACGAGCAGTAGATCCAGCAGCCTGCAGCTCCAAGGA ACATTGCCTCTCTGGATCTGCTGTGACCAGGGTGTGGC Amplicon Length (bp) 61 Chromosome Location 16:67322707-67322797 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 101 R2 0.9992 cDNA Cq 22.17 cDNA Tm (Celsius) 82 gDNA Cq 24.93 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report PLEKHG4, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak