Deleted Genes Associated with Digeorge/22Q11.2 Deletion Syndrome
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-
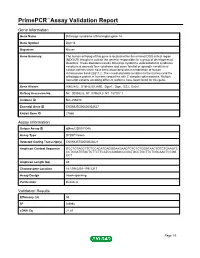
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name DiGeorge syndrome critical region gene 14 Gene Symbol Dgcr14 Organism Mouse Gene Summary The human ortholog of this gene is located within the minimal DGS critical region (MDGCR) thought to contain the gene(s) responsible for a group of developmental disorders. These disorders include DiGeorge syndrome velocardiofacial syndrome conotruncal anomaly face syndrome and some familial or sporadic conotruncal cardiac defects which have been associated with microdeletion of human chromosome band 22q11.2. The encoded protein localizes to the nucleus and the orthologous protein in humans co-purifies with C complex spliceosomes. Multiple transcript variants encoding different isoforms have been found for this gene. Gene Aliases AI462402, D16H22S1269E, Dgcr1, Dgsi, ES2, Es2el RefSeq Accession No. NC_000082.6, NT_039624.8, NT_187007.1 UniGene ID Mm.256480 Ensembl Gene ID ENSMUSG00000003527 Entrez Gene ID 27886 Assay Information Unique Assay ID qMmuCID0011048 Assay Type SYBR® Green Detected Coding Transcript(s) ENSMUST00000003621 Amplicon Context Sequence GCCTGTAGCTTCTCCACATCAGGGAAGAAGTCTCTCTGGATAACTGTCTGAAGTC CCTCGATGTACTCTTCTTCATCCAGGACCCGCTGCCTGCTTCTCGCAACTCCGG CCT Amplicon Length (bp) 82 Chromosome Location 16:17910201-17911217 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 95 R2 0.9994 cDNA Cq 21.87 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 79.5 gDNA Cq 23.94 Specificity (%) 100 Information to assist with data interpretation is provided -

CRNKL1 Is a Highly Selective Regulator of Intron-Retaining HIV-1 and Cellular Mrnas
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.04.934927; this version posted February 11, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 CRNKL1 is a highly selective regulator of intron-retaining HIV-1 and cellular mRNAs 2 3 4 Han Xiao1, Emanuel Wyler2#, Miha Milek2#, Bastian Grewe3, Philipp Kirchner4, Arif Ekici4, Ana Beatriz 5 Oliveira Villela Silva1, Doris Jungnickl1, Markus Landthaler2,5, Armin Ensser1, and Klaus Überla1* 6 7 1 Institute of Clinical and Molecular Virology, University Hospital Erlangen, Friedrich-Alexander 8 Universität Erlangen-Nürnberg, Erlangen, Germany 9 2 Berlin Institute for Medical Systems Biology, Max-Delbrück-Center for Molecular Medicine in the 10 Helmholtz Association, Robert-Rössle-Strasse 10, 13125, Berlin, Germany 11 3 Department of Molecular and Medical Virology, Ruhr-University, Bochum, Germany 12 4 Institute of Human Genetics, University Hospital Erlangen, Friedrich-Alexander Universität Erlangen- 13 Nürnberg, Erlangen, Germany 14 5 IRI Life Sciences, Institute für Biologie, Humboldt Universität zu Berlin, Philippstraße 13, 10115, Berlin, 15 Germany 16 # these two authors contributed equally 17 18 19 *Corresponding author: 20 Prof. Dr. Klaus Überla 21 Institute of Clinical and Molecular Virology, University Hospital Erlangen 22 Friedrich-Alexander Universität Erlangen-Nürnberg 23 Schlossgarten 4, 91054 Erlangen 24 Germany 25 Tel: (+49) 9131-8523563 26 e-mail: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.04.934927; this version posted February 11, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Protein Kinase A-Mediated Septin7 Phosphorylation Disrupts Septin Filaments and Ciliogenesis
cells Article Protein Kinase A-Mediated Septin7 Phosphorylation Disrupts Septin Filaments and Ciliogenesis Han-Yu Wang 1,2, Chun-Hsiang Lin 1, Yi-Ru Shen 1, Ting-Yu Chen 2,3, Chia-Yih Wang 2,3,* and Pao-Lin Kuo 1,2,4,* 1 Department of Obstetrics and Gynecology, College of Medicine, National Cheng Kung University, Tainan 701, Taiwan; [email protected] (H.-Y.W.); [email protected] (C.-H.L.); [email protected] (Y.-R.S.) 2 Institute of Basic Medical Sciences, College of Medicine, National Cheng Kung University, Tainan 701, Taiwan; [email protected] 3 Department of Cell Biology and Anatomy, College of Medicine, National Cheng Kung University, Tainan 701, Taiwan 4 Department of Obstetrics and Gynecology, National Cheng-Kung University Hospital, Tainan 704, Taiwan * Correspondence: [email protected] (C.-Y.W.); [email protected] (P.-L.K.); Tel.: +886-6-2353535 (ext. 5338); (C.-Y.W.)+886-6-2353535 (ext. 5262) (P.-L.K.) Abstract: Septins are GTP-binding proteins that form heteromeric filaments for proper cell growth and migration. Among the septins, septin7 (SEPT7) is an important component of all septin filaments. Here we show that protein kinase A (PKA) phosphorylates SEPT7 at Thr197, thus disrupting septin filament dynamics and ciliogenesis. The Thr197 residue of SEPT7, a PKA phosphorylating site, was conserved among different species. Treatment with cAMP or overexpression of PKA catalytic subunit (PKACA2) induced SEPT7 phosphorylation, followed by disruption of septin filament formation. Constitutive phosphorylation of SEPT7 at Thr197 reduced SEPT7-SEPT7 interaction, but did not affect SEPT7-SEPT6-SEPT2 or SEPT4 interaction. -

1 Supporting Information for a Microrna Network Regulates
Supporting Information for A microRNA Network Regulates Expression and Biosynthesis of CFTR and CFTR-ΔF508 Shyam Ramachandrana,b, Philip H. Karpc, Peng Jiangc, Lynda S. Ostedgaardc, Amy E. Walza, John T. Fishere, Shaf Keshavjeeh, Kim A. Lennoxi, Ashley M. Jacobii, Scott D. Rosei, Mark A. Behlkei, Michael J. Welshb,c,d,g, Yi Xingb,c,f, Paul B. McCray Jr.a,b,c Author Affiliations: Department of Pediatricsa, Interdisciplinary Program in Geneticsb, Departments of Internal Medicinec, Molecular Physiology and Biophysicsd, Anatomy and Cell Biologye, Biomedical Engineeringf, Howard Hughes Medical Instituteg, Carver College of Medicine, University of Iowa, Iowa City, IA-52242 Division of Thoracic Surgeryh, Toronto General Hospital, University Health Network, University of Toronto, Toronto, Canada-M5G 2C4 Integrated DNA Technologiesi, Coralville, IA-52241 To whom correspondence should be addressed: Email: [email protected] (M.J.W.); yi- [email protected] (Y.X.); Email: [email protected] (P.B.M.) This PDF file includes: Materials and Methods References Fig. S1. miR-138 regulates SIN3A in a dose-dependent and site-specific manner. Fig. S2. miR-138 regulates endogenous SIN3A protein expression. Fig. S3. miR-138 regulates endogenous CFTR protein expression in Calu-3 cells. Fig. S4. miR-138 regulates endogenous CFTR protein expression in primary human airway epithelia. Fig. S5. miR-138 regulates CFTR expression in HeLa cells. Fig. S6. miR-138 regulates CFTR expression in HEK293T cells. Fig. S7. HeLa cells exhibit CFTR channel activity. Fig. S8. miR-138 improves CFTR processing. Fig. S9. miR-138 improves CFTR-ΔF508 processing. Fig. S10. SIN3A inhibition yields partial rescue of Cl- transport in CF epithelia. -

Hira Peracha a Thesis
DIAGNOSIS METHOD FOR MUCOPOLYSACCHARIDOSES by Hira Peracha A thesis submitted to the Faculty of the University of Delaware in partial fulfillment of the requirements for the degree of Bachelor of Arts in Biological Sciences with Distinction Spring 2018 © 2018 Hira Peracha All Rights Reserved i DIAGNOSIS METHOD FOR MUCOPOLYSACCHARIDOSES by Hira Peracha Approved: __________________________________________________________ Shunji Tomatsu, MD, Ph.D. Professor in charge of thesis on behalf of the Advisory Committee Approved: __________________________________________________________ Deni Galileo, Ph.D. Professor in charge of thesis on behalf of the Advisory Committee Approved: __________________________________________________________ Jessica Tanis, Ph.D. Committee member from the Department of Department Name Approved: __________________________________________________________ Barbara Settles, Ph.D. Committee member from the Board of Senior Thesis Readers Approved: __________________________________________________________ Michael Chajes, Ph.D. Chair of the University Committee on Student and Faculty Honors ii “Education is the key to unlocking the world, a passport to freedom.” -Oprah Winfrey iii ACKNOWLEDGMENTS I dedicate my thesis to my late grandfather, who taught me the value of life, hard work and sacrifice- especially in sickness and in health. I would like to express my deepest acknowledgements to my family. I could not have achieved all of my dreams and goals in life without my parents; my mom and my dad, whose sacrifices, love and support have been endless and infinite. Thank you for raising me to believe that anything was possible, and for spending your entire lives to better mine. To my younger siblings, Saarah, Hafsah and Arif, who have been the light in the dark times in my life and have always brought joy and happiness in my life. -

Análise Integrativa De Perfis Transcricionais De Pacientes Com
UNIVERSIDADE DE SÃO PAULO FACULDADE DE MEDICINA DE RIBEIRÃO PRETO PROGRAMA DE PÓS-GRADUAÇÃO EM GENÉTICA ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Ribeirão Preto – 2012 ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Tese apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo para obtenção do título de Doutor em Ciências. Área de Concentração: Genética Orientador: Prof. Dr. Eduardo Antonio Donadi Co-orientador: Prof. Dr. Geraldo A. S. Passos Ribeirão Preto – 2012 AUTORIZO A REPRODUÇÃO E DIVULGAÇÃO TOTAL OU PARCIAL DESTE TRABALHO, POR QUALQUER MEIO CONVENCIONAL OU ELETRÔNICO, PARA FINS DE ESTUDO E PESQUISA, DESDE QUE CITADA A FONTE. FICHA CATALOGRÁFICA Evangelista, Adriane Feijó Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. Ribeirão Preto, 2012 192p. Tese de Doutorado apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo. Área de Concentração: Genética. Orientador: Donadi, Eduardo Antonio Co-orientador: Passos, Geraldo A. 1. Expressão gênica – microarrays 2. Análise bioinformática por module maps 3. Diabetes mellitus tipo 1 4. Diabetes mellitus tipo 2 5. Diabetes mellitus gestacional FOLHA DE APROVAÇÃO ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. -

Association of Gene Ontology Categories with Decay Rate for Hepg2 Experiments These Tables Show Details for All Gene Ontology Categories
Supplementary Table 1: Association of Gene Ontology Categories with Decay Rate for HepG2 Experiments These tables show details for all Gene Ontology categories. Inferences for manual classification scheme shown at the bottom. Those categories used in Figure 1A are highlighted in bold. Standard Deviations are shown in parentheses. P-values less than 1E-20 are indicated with a "0". Rate r (hour^-1) Half-life < 2hr. Decay % GO Number Category Name Probe Sets Group Non-Group Distribution p-value In-Group Non-Group Representation p-value GO:0006350 transcription 1523 0.221 (0.009) 0.127 (0.002) FASTER 0 13.1 (0.4) 4.5 (0.1) OVER 0 GO:0006351 transcription, DNA-dependent 1498 0.220 (0.009) 0.127 (0.002) FASTER 0 13.0 (0.4) 4.5 (0.1) OVER 0 GO:0006355 regulation of transcription, DNA-dependent 1163 0.230 (0.011) 0.128 (0.002) FASTER 5.00E-21 14.2 (0.5) 4.6 (0.1) OVER 0 GO:0006366 transcription from Pol II promoter 845 0.225 (0.012) 0.130 (0.002) FASTER 1.88E-14 13.0 (0.5) 4.8 (0.1) OVER 0 GO:0006139 nucleobase, nucleoside, nucleotide and nucleic acid metabolism3004 0.173 (0.006) 0.127 (0.002) FASTER 1.28E-12 8.4 (0.2) 4.5 (0.1) OVER 0 GO:0006357 regulation of transcription from Pol II promoter 487 0.231 (0.016) 0.132 (0.002) FASTER 6.05E-10 13.5 (0.6) 4.9 (0.1) OVER 0 GO:0008283 cell proliferation 625 0.189 (0.014) 0.132 (0.002) FASTER 1.95E-05 10.1 (0.6) 5.0 (0.1) OVER 1.50E-20 GO:0006513 monoubiquitination 36 0.305 (0.049) 0.134 (0.002) FASTER 2.69E-04 25.4 (4.4) 5.1 (0.1) OVER 2.04E-06 GO:0007050 cell cycle arrest 57 0.311 (0.054) 0.133 (0.002) -

Anti-Histone H2B Antibody (ARG41373)
Product datasheet [email protected] ARG41373 Package: 100 μl anti-Histone H2B antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes Histone H2B Tested Reactivity Hu, Ms, Rat Tested Application FACS, ICC/IF, IHC-P, IP, WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name Histone H2B Antigen Species Human Immunogen Synthetic peptide derived from Human Histone H2B. Conjugation Un-conjugated Alternate Names Histone H2B type 1-K; H2BFAiii; H2BFT; H2B/S; H2B K; H2BK; HIRA-interacting protein 1 Application Instructions Application table Application Dilution FACS 1:50 ICC/IF 1:50 - 1:200 IHC-P 1:50 - 1:200 IP 1:30 WB 1:5000 - 1:20000 Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 14 kDa Observed Size ~ 14 kDa Properties Form Liquid Purification Affinity purified. Buffer PBS (pH 7.4), 150 mM NaCl, 0.02% Sodium azide and 50% Glycerol. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot www.arigobio.com 1/2 and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation Gene Symbol HIST1H2BK Gene Full Name histone cluster 1, H2bk Background Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. -

Protein Interactions in the Cancer Proteome† Cite This: Mol
Molecular BioSystems View Article Online PAPER View Journal | View Issue Small-molecule binding sites to explore protein– protein interactions in the cancer proteome† Cite this: Mol. BioSyst., 2016, 12,3067 David Xu,ab Shadia I. Jalal,c George W. Sledge Jr.d and Samy O. Meroueh*aef The Cancer Genome Atlas (TCGA) offers an unprecedented opportunity to identify small-molecule binding sites on proteins with overexpressed mRNA levels that correlate with poor survival. Here, we analyze RNA-seq and clinical data for 10 tumor types to identify genes that are both overexpressed and correlate with patient survival. Protein products of these genes were scanned for binding sites that possess shape and physicochemical properties that can accommodate small-molecule probes or therapeutic agents (druggable). These binding sites were classified as enzyme active sites (ENZ), protein–protein interaction sites (PPI), or other sites whose function is unknown (OTH). Interestingly, the overwhelming majority of binding sites were classified as OTH. We find that ENZ, PPI, and OTH binding sites often occurred on the same structure suggesting that many of these OTH cavities can be used for allosteric modulation of Creative Commons Attribution 3.0 Unported Licence. enzyme activity or protein–protein interactions with small molecules. We discovered several ENZ (PYCR1, QPRT,andHSPA6)andPPI(CASC5, ZBTB32,andCSAD) binding sites on proteins that have been seldom explored in cancer. We also found proteins that have been extensively studied in cancer that have not been previously explored with small molecules that harbor ENZ (PKMYT1, STEAP3,andNNMT) and PPI (HNF4A, MEF2B,andCBX2) binding sites. All binding sites were classified by the signaling pathways to Received 29th March 2016, which the protein that harbors them belongs using KEGG. -

Análise Correlacional Entre a Expressão Dos Fatores De Splicing E a Ocorrência De Splicing Alternativo Em Tecidos Humanos E De Camundongos
ANÁLISE CORRELACIONAL ENTRE A EXPRESSÃO DOS FATORES DE SPLICING E A OCORRÊNCIA DE SPLICING ALTERNATIVO EM TECIDOS HUMANOS E DE CAMUNDONGOS JULIO CÉSAR NUNES Dissertação apresentada à Fundação Antônio Prudente para a obtenção do título de Mestre em Ciências Área de Concentração: Oncologia Orientador: Dr. Sandro José de Souza São Paulo 2008 Livros Grátis http://www.livrosgratis.com.br Milhares de livros grátis para download. FICHA CATALOGRÁFICA Preparada pela Biblioteca da Fundação Antônio Prudente Nunes, Julio César Análise correlacional entre a expressão dos fatores de splicing e a ocorrência de splicing alternativo em tecidos humanos e de camundongos / Julio César Nunes – São Paulo, 2008. 79p. Dissertação (Mestrado) - Fundação Antônio Prudente. Curso de Pós-Graduação em Ciências - Área de concentração: Oncologia. Orientador: Sandro José Souza Descritores: 1. SPLICING ALTERNATIVO 2. BIOLOGIA MOLECULAR COMPUTACIONAL 3. CÂNCER 4. GENOMICA. AGRADECIMENTOS Agradeço à FAPESP e CAPES pela bolsa de Mestrado. Ao Sandro José de Souza agradeço toda orientação e conhecimento oferecido. Meus especiais agradecimentos ao Pedro Alexandre Favoretto Galante que dedicou atenção a minha formação no processo de Pós-Graduação na Fundação Antônio Prudente, bem como pela sua oficiosa co-orientação ao projeto de pesquisa. À grande família e amigos pela dedicação e incentivo a minha formação acadêmica. À Fundação Antônio Prudente, Hospital do Câncer e Instituto Ludwig de Pesquisa sobre o Câncer dedico os meus nobres agradecimentos finais. RESUMO Nunes JC. Análise correlacional entre a expressão dos fatores de splicing e a ocorrência de splicing alternativo em tecidos humanos e de camundongos. São Paulo; 2007. [Dissertacão de Mestrado - Fundação Antônio Prudente] Splicing alternativo desempenha uma significante função no aumento da complexidade genômica, produzindo um extenso número de mRNA e isoformas protéicas. -

Genomic and Functional Analysis Identifies CRKL As an Oncogene
Oncogene (2010) 29, 1421–1430 & 2010 Macmillan Publishers Limited All rights reserved 0950-9232/10 $32.00 www.nature.com/onc ORIGINAL ARTICLE Genomic and functional analysis identifies CRKL as an oncogene amplified in lung cancer YH Kim1,7, KA Kwei1,7, L Girard2, K Salari1,3, J Kao1, M Pacyna-Gengelbach4, P Wang5, T Hernandez-Boussard3, AF Gazdar2, I Petersen6, JD Minna2 and JR Pollack1 1Department of Pathology, Stanford University, Stanford, CA, USA; 2Hamon Center for Therapeutic Oncology Research, University of Texas Southwestern Medical Center, Dallas, TX, USA; 3Department of Genetics, Stanford University, Stanford, CA, USA; 4Institute of Pathology, University Hospital Charite´, Berlin, Germany; 5Division of Public Health Sciences, Fred Hutchinson Cancer Research Center, Seattle, WA, USA and 6Institute of Pathology, Universita¨tsklinikum Jena, Jena, Germany DNA amplifications, leading to the overexpression of related mortality (Jemal et al., 2008). Nearly 80% oncogenes, are a cardinal feature of lung cancer and of lung cancers diagnosed are non-small-cell lung directly contribute to its pathogenesis. To uncover such cancers (NSCLCs), which are classified into three main novel alterations, we performed an array-based compara- histological subtypes: adenocarcinoma, squamous cell tive genomic hybridization survey of 128 non-small-cell carcinoma and large cell carcinoma. Despite the lung cancer cell lines and tumors. Prominent among our advancement of surgical, cytotoxic and radiological findings, we identified recurrent high-level amplification at treatment options over the years, lung cancer therapy cytoband 22q11.21 in 3% of lung cancer specimens, with remains largely ineffective from a clinical standpoint, another 11% of specimens exhibiting low-level gain spanning as evidenced by a low 5-year survival rate (o15%), that locus. -

DGCR6 at the Proximal Part of the Digeorge Critical Region Is Involved in Conotruncal Heart Defects
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Institutional Repository : the EHIME area OPEN Citation: Human Genome Variation (2015) 2, 15004; doi:10.1038/hgv.2015.4 © 2015 The Japan Society of Human Genetics All rights reserved 2054-345X/15 www.nature.com/hgv ARTICLE DGCR6 at the proximal part of the DiGeorge critical region is involved in conotruncal heart defects Wenming Gao1, Takashi Higaki1, Minenori Eguchi-Ishimae1, Hidehiko Iwabuki1, Zhouying Wu1, Eiichi Yamamoto2, Hidemi Takata1, Masaaki Ohta1, Issei Imoto3, Eiichi Ishii1 and Mariko Eguchi1 Cardiac anomaly is one of the hallmarks of DiGeorge syndrome (DGS), observed in approximately 80% of patients. It often shows a characteristic morphology, termed as conotruncal heart defects. In many cases showing only the conotruncal heart defect, deletion of 22q11.2 region cannot be detected by fluorescence in situ hybridization (FISH), which is used to detect deletion in DGS. We investigated the presence of genomic aberrations in six patients with congenital conotruncal heart defects, who show no deletion at 22q11.2 in an initial screening by FISH. In these patients, no abnormalities were identified in the coding region of the TBX1 gene, one of the key genes responsible for the phenotype of DGS. However, when copy number alteration was analyzed by high-resolution array analysis, a small deletion or duplication in the proximal end of DiGeorge critical region was detected in two patients. The affected region contains the DGCR6 and PRODH genes. DGCR6 has been reported to affect the expression of the TBX1 gene. Our results suggest that altered dosage of gene(s) other than TBX1, possibly DGCR6, may also be responsible for the development of conotruncal heart defects observed in patients with DGS and, in particular, in those with stand-alone conotruncal heart defects.