Bayesian Linear Regression
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Logistic Regression, Dependencies, Non-Linear Data and Model Reduction
COMP6237 – Logistic Regression, Dependencies, Non-linear Data and Model Reduction Markus Brede [email protected] Lecture slides available here: http://users.ecs.soton.ac.uk/mb8/stats/datamining.html (Thanks to Jason Noble and Cosma Shalizi whose lecture materials I used to prepare) COMP6237: Logistic Regression ● Outline: – Introduction – Basic ideas of logistic regression – Logistic regression using R – Some underlying maths and MLE – The multinomial case – How to deal with non-linear data ● Model reduction and AIC – How to deal with dependent data – Summary – Problems Introduction ● Previous lecture: Linear regression – tried to predict a continuous variable from variation in another continuous variable (E.g. basketball ability from height) ● Here: Logistic regression – Try to predict results of a binary (or categorical) outcome variable Y from a predictor variable X – This is a classification problem: classify X as belonging to one of two classes – Occurs quite often in science … e.g. medical trials (will a patient live or die dependent on medication?) Dependent variable Y Predictor Variables X The Oscars Example ● A fictional data set that looks at what it takes for a movie to win an Oscar ● Outcome variable: Oscar win, yes or no? ● Predictor variables: – Box office takings in millions of dollars – Budget in millions of dollars – Country of origin: US, UK, Europe, India, other – Critical reception (scores 0 … 100) – Length of film in minutes – This (fictitious) data set is available here: https://www.southampton.ac.uk/~mb1a10/stats/filmData.txt Predicting Oscar Success ● Let's start simple and look at only one of the predictor variables ● Do big box office takings make Oscar success more likely? ● Could use same techniques as below to look at budget size, film length, etc. -

Naïve Bayes Classifier
CS 60050 Machine Learning Naïve Bayes Classifier Some slides taken from course materials of Tan, Steinbach, Kumar Bayes Classifier ● A probabilistic framework for solving classification problems ● Approach for modeling probabilistic relationships between the attribute set and the class variable – May not be possible to certainly predict class label of a test record even if it has identical attributes to some training records – Reason: noisy data or presence of certain factors that are not included in the analysis Probability Basics ● P(A = a, C = c): joint probability that random variables A and C will take values a and c respectively ● P(A = a | C = c): conditional probability that A will take the value a, given that C has taken value c P(A,C) P(C | A) = P(A) P(A,C) P(A | C) = P(C) Bayes Theorem ● Bayes theorem: P(A | C)P(C) P(C | A) = P(A) ● P(C) known as the prior probability ● P(C | A) known as the posterior probability Example of Bayes Theorem ● Given: – A doctor knows that meningitis causes stiff neck 50% of the time – Prior probability of any patient having meningitis is 1/50,000 – Prior probability of any patient having stiff neck is 1/20 ● If a patient has stiff neck, what’s the probability he/she has meningitis? P(S | M )P(M ) 0.5×1/50000 P(M | S) = = = 0.0002 P(S) 1/ 20 Bayesian Classifiers ● Consider each attribute and class label as random variables ● Given a record with attributes (A1, A2,…,An) – Goal is to predict class C – Specifically, we want to find the value of C that maximizes P(C| A1, A2,…,An ) ● Can we estimate -
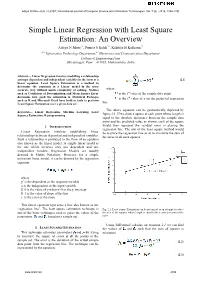
Simple Linear Regression with Least Square Estimation: an Overview
Aditya N More et al, / (IJCSIT) International Journal of Computer Science and Information Technologies, Vol. 7 (6) , 2016, 2394-2396 Simple Linear Regression with Least Square Estimation: An Overview Aditya N More#1, Puneet S Kohli*2, Kshitija H Kulkarni#3 #1-2Information Technology Department,#3 Electronics and Communication Department College of Engineering Pune Shivajinagar, Pune – 411005, Maharashtra, India Abstract— Linear Regression involves modelling a relationship amongst dependent and independent variables in the form of a (2.1) linear equation. Least Square Estimation is a method to determine the constants in a Linear model in the most accurate way without much complexity of solving. Metrics where such as Coefficient of Determination and Mean Square Error is the ith value of the sample data point determine how good the estimation is. Statistical Packages is the ith value of y on the predicted regression such as R and Microsoft Excel have built in tools to perform Least Square Estimation over a given data set. line The above equation can be geometrically depicted by Keywords— Linear Regression, Machine Learning, Least Squares Estimation, R programming figure 2.1. If we draw a square at each point whose length is equal to the absolute difference between the sample data point and the predicted value as shown, each of the square would then represent the residual error in placing the I. INTRODUCTION regression line. The aim of the least square method would Linear Regression involves establishing linear be to place the regression line so as to minimize the sum of relationships between dependent and independent variables. -

The Composite Marginal Likelihood (CML) Inference Approach with Applications to Discrete and Mixed Dependent Variable Models
Technical Report 101 The Composite Marginal Likelihood (CML) Inference Approach with Applications to Discrete and Mixed Dependent Variable Models Chandra R. Bhat Center for Transportation Research September 2014 Data-Supported Transportation Operations & Planning Center (D-STOP) A Tier 1 USDOT University Transportation Center at The University of Texas at Austin D-STOP is a collaborative initiative by researchers at the Center for Transportation Research and the Wireless Networking and Communications Group at The University of Texas at Austin. DISCLAIMER The contents of this report reflect the views of the authors, who are responsible for the facts and the accuracy of the information presented herein. This document is disseminated under the sponsorship of the U.S. Department of Transportation’s University Transportation Centers Program, in the interest of information exchange. The U.S. Government assumes no liability for the contents or use thereof. Technical Report Documentation Page 1. Report No. 2. Government Accession No. 3. Recipient's Catalog No. D-STOP/2016/101 4. Title and Subtitle 5. Report Date The Composite Marginal Likelihood (CML) Inference Approach September 2014 with Applications to Discrete and Mixed Dependent Variable 6. Performing Organization Code Models 7. Author(s) 8. Performing Organization Report No. Chandra R. Bhat Report 101 9. Performing Organization Name and Address 10. Work Unit No. (TRAIS) Data-Supported Transportation Operations & Planning Center (D- STOP) 11. Contract or Grant No. The University of Texas at Austin DTRT13-G-UTC58 1616 Guadalupe Street, Suite 4.202 Austin, Texas 78701 12. Sponsoring Agency Name and Address 13. Type of Report and Period Covered Data-Supported Transportation Operations & Planning Center (D- STOP) The University of Texas at Austin 14. -

The Bayesian Approach to Statistics
THE BAYESIAN APPROACH TO STATISTICS ANTHONY O’HAGAN INTRODUCTION the true nature of scientific reasoning. The fi- nal section addresses various features of modern By far the most widely taught and used statisti- Bayesian methods that provide some explanation for the rapid increase in their adoption since the cal methods in practice are those of the frequen- 1980s. tist school. The ideas of frequentist inference, as set out in Chapter 5 of this book, rest on the frequency definition of probability (Chapter 2), BAYESIAN INFERENCE and were developed in the first half of the 20th century. This chapter concerns a radically differ- We first present the basic procedures of Bayesian ent approach to statistics, the Bayesian approach, inference. which depends instead on the subjective defini- tion of probability (Chapter 3). In some respects, Bayesian methods are older than frequentist ones, Bayes’s Theorem and the Nature of Learning having been the basis of very early statistical rea- Bayesian inference is a process of learning soning as far back as the 18th century. Bayesian from data. To give substance to this statement, statistics as it is now understood, however, dates we need to identify who is doing the learning and back to the 1950s, with subsequent development what they are learning about. in the second half of the 20th century. Over that time, the Bayesian approach has steadily gained Terms and Notation ground, and is now recognized as a legitimate al- ternative to the frequentist approach. The person doing the learning is an individual This chapter is organized into three sections. -

Scalable and Robust Bayesian Inference Via the Median Posterior
Scalable and Robust Bayesian Inference via the Median Posterior CS 584: Big Data Analytics Material adapted from David Dunson’s talk (http://bayesian.org/sites/default/files/Dunson.pdf) & Lizhen Lin’s ICML talk (http://techtalks.tv/talks/scalable-and-robust-bayesian-inference-via-the-median-posterior/61140/) Big Data Analytics • Large (big N) and complex (big P with interactions) data are collected routinely • Both speed & generality of data analysis methods are important • Bayesian approaches offer an attractive general approach for modeling the complexity of big data • Computational intractability of posterior sampling is a major impediment to application of flexible Bayesian methods CS 584 [Spring 2016] - Ho Existing Frequentist Approaches: The Positives • Optimization-based approaches, such as ADMM or glmnet, are currently most popular for analyzing big data • General and computationally efficient • Used orders of magnitude more than Bayes methods • Can exploit distributed & cloud computing platforms • Can borrow some advantages of Bayes methods through penalties and regularization CS 584 [Spring 2016] - Ho Existing Frequentist Approaches: The Drawbacks • Such optimization-based methods do not provide measure of uncertainty • Uncertainty quantification is crucial for most applications • Scalable penalization methods focus primarily on convex optimization — greatly limits scope and puts ceiling on performance • For non-convex problems and data with complex structure, existing optimization algorithms can fail badly CS 584 [Spring 2016] - -

LINEAR REGRESSION MODELS Likelihood and Reference Bayesian
{ LINEAR REGRESSION MODELS { Likeliho o d and Reference Bayesian Analysis Mike West May 3, 1999 1 Straight Line Regression Mo dels We b egin with the simple straight line regression mo del y = + x + i i i where the design p oints x are xed in advance, and the measurement/sampling i 2 errors are indep endent and normally distributed, N 0; for each i = i i 1;::: ;n: In this context, wehavelooked at general mo delling questions, data and the tting of least squares estimates of and : Now we turn to more formal likeliho o d and Bayesian inference. 1.1 Likeliho o d and MLEs The formal parametric inference problem is a multi-parameter problem: we 2 require inferences on the three parameters ; ; : The likeliho o d function has a simple enough form, as wenow show. Throughout, we do not indicate the design p oints in conditioning statements, though they are implicitly conditioned up on. Write Y = fy ;::: ;y g and X = fx ;::: ;x g: Given X and the mo del 1 n 1 n parameters, each y is the corresp onding zero-mean normal random quantity i plus the term + x ; so that y is normal with this term as its mean and i i i 2 variance : Also, since the are indep endent, so are the y : Thus i i 2 2 y j ; ; N + x ; i i with conditional density function 2 2 2 2 1=2 py j ; ; = expfy x =2 g=2 i i i for each i: Also, by indep endence, the joint density function is n Y 2 2 : py j ; ; = pY j ; ; i i=1 1 Given the observed resp onse values Y this provides the likeliho o d function for the three parameters: the joint density ab ove evaluated at the observed and 2 hence now xed values of Y; now viewed as a function of ; ; as they vary across p ossible parameter values. -

1 Estimation and Beyond in the Bayes Universe
ISyE8843A, Brani Vidakovic Handout 7 1 Estimation and Beyond in the Bayes Universe. 1.1 Estimation No Bayes estimate can be unbiased but Bayesians are not upset! No Bayes estimate with respect to the squared error loss can be unbiased, except in a trivial case when its Bayes’ risk is 0. Suppose that for a proper prior ¼ the Bayes estimator ±¼(X) is unbiased, Xjθ (8θ)E ±¼(X) = θ: This implies that the Bayes risk is 0. The Bayes risk of ±¼(X) can be calculated as repeated expectation in two ways, θ Xjθ 2 X θjX 2 r(¼; ±¼) = E E (θ ¡ ±¼(X)) = E E (θ ¡ ±¼(X)) : Thus, conveniently choosing either EθEXjθ or EX EθjX and using the properties of conditional expectation we have, θ Xjθ 2 θ Xjθ X θjX X θjX 2 r(¼; ±¼) = E E θ ¡ E E θ±¼(X) ¡ E E θ±¼(X) + E E ±¼(X) θ Xjθ 2 θ Xjθ X θjX X θjX 2 = E E θ ¡ E θ[E ±¼(X)] ¡ E ±¼(X)E θ + E E ±¼(X) θ Xjθ 2 θ X X θjX 2 = E E θ ¡ E θ ¢ θ ¡ E ±¼(X)±¼(X) + E E ±¼(X) = 0: Bayesians are not upset. To check for its unbiasedness, the Bayes estimator is averaged with respect to the model measure (Xjθ), and one of the Bayesian commandments is: Thou shall not average with respect to sample space, unless you have Bayesian design in mind. Even frequentist agree that insisting on unbiasedness can lead to bad estimators, and that in their quest to minimize the risk by trading off between variance and bias-squared a small dosage of bias can help. -

Logistic Regression Maths and Statistics Help Centre
Logistic regression Maths and Statistics Help Centre Many statistical tests require the dependent (response) variable to be continuous so a different set of tests are needed when the dependent variable is categorical. One of the most commonly used tests for categorical variables is the Chi-squared test which looks at whether or not there is a relationship between two categorical variables but this doesn’t make an allowance for the potential influence of other explanatory variables on that relationship. For continuous outcome variables, Multiple regression can be used for a) controlling for other explanatory variables when assessing relationships between a dependent variable and several independent variables b) predicting outcomes of a dependent variable using a linear combination of explanatory (independent) variables The maths: For multiple regression a model of the following form can be used to predict the value of a response variable y using the values of a number of explanatory variables: y 0 1x1 2 x2 ..... q xq 0 Constant/ intercept , 1 q co efficients for q explanatory variables x1 xq The regression process finds the co-efficients which minimise the squared differences between the observed and expected values of y (the residuals). As the outcome of logistic regression is binary, y needs to be transformed so that the regression process can be used. The logit transformation gives the following: p ln 0 1x1 2 x2 ..... q xq 1 p p p probabilty of event occuring e.g. person dies following heart attack, odds ratio 1- p If probabilities of the event of interest happening for individuals are needed, the logistic regression equation exp x x .... -

Generalized Linear Models
CHAPTER 6 Generalized linear models 6.1 Introduction Generalized linear modeling is a framework for statistical analysis that includes linear and logistic regression as special cases. Linear regression directly predicts continuous data y from a linear predictor Xβ = β0 + X1β1 + + Xkβk.Logistic regression predicts Pr(y =1)forbinarydatafromalinearpredictorwithaninverse-··· logit transformation. A generalized linear model involves: 1. A data vector y =(y1,...,yn) 2. Predictors X and coefficients β,formingalinearpredictorXβ 1 3. A link function g,yieldingavectoroftransformeddataˆy = g− (Xβ)thatare used to model the data 4. A data distribution, p(y yˆ) | 5. Possibly other parameters, such as variances, overdispersions, and cutpoints, involved in the predictors, link function, and data distribution. The options in a generalized linear model are the transformation g and the data distribution p. In linear regression,thetransformationistheidentity(thatis,g(u) u)and • the data distribution is normal, with standard deviation σ estimated from≡ data. 1 1 In logistic regression,thetransformationistheinverse-logit,g− (u)=logit− (u) • (see Figure 5.2a on page 80) and the data distribution is defined by the proba- bility for binary data: Pr(y =1)=y ˆ. This chapter discusses several other classes of generalized linear model, which we list here for convenience: The Poisson model (Section 6.2) is used for count data; that is, where each • data point yi can equal 0, 1, 2, ....Theusualtransformationg used here is the logarithmic, so that g(u)=exp(u)transformsacontinuouslinearpredictorXiβ to a positivey ˆi.ThedatadistributionisPoisson. It is usually a good idea to add a parameter to this model to capture overdis- persion,thatis,variationinthedatabeyondwhatwouldbepredictedfromthe Poisson distribution alone. -

Generalized Linear Models
Generalized Linear Models Advanced Methods for Data Analysis (36-402/36-608) Spring 2014 1 Generalized linear models 1.1 Introduction: two regressions • So far we've seen two canonical settings for regression. Let X 2 Rp be a vector of predictors. In linear regression, we observe Y 2 R, and assume a linear model: T E(Y jX) = β X; for some coefficients β 2 Rp. In logistic regression, we observe Y 2 f0; 1g, and we assume a logistic model (Y = 1jX) log P = βT X: 1 − P(Y = 1jX) • What's the similarity here? Note that in the logistic regression setting, P(Y = 1jX) = E(Y jX). Therefore, in both settings, we are assuming that a transformation of the conditional expec- tation E(Y jX) is a linear function of X, i.e., T g E(Y jX) = β X; for some function g. In linear regression, this transformation was the identity transformation g(u) = u; in logistic regression, it was the logit transformation g(u) = log(u=(1 − u)) • Different transformations might be appropriate for different types of data. E.g., the identity transformation g(u) = u is not really appropriate for logistic regression (why?), and the logit transformation g(u) = log(u=(1 − u)) not appropriate for linear regression (why?), but each is appropriate in their own intended domain • For a third data type, it is entirely possible that transformation neither is really appropriate. What to do then? We think of another transformation g that is in fact appropriate, and this is the basic idea behind a generalized linear model 1.2 Generalized linear models • Given predictors X 2 Rp and an outcome Y , a generalized linear model is defined by three components: a random component, that specifies a distribution for Y jX; a systematic compo- nent, that relates a parameter η to the predictors X; and a link function, that connects the random and systematic components • The random component specifies a distribution for the outcome variable (conditional on X). -

Introduction to Bayesian Inference and Modeling Edps 590BAY
Introduction to Bayesian Inference and Modeling Edps 590BAY Carolyn J. Anderson Department of Educational Psychology c Board of Trustees, University of Illinois Fall 2019 Introduction What Why Probability Steps Example History Practice Overview ◮ What is Bayes theorem ◮ Why Bayesian analysis ◮ What is probability? ◮ Basic Steps ◮ An little example ◮ History (not all of the 705+ people that influenced development of Bayesian approach) ◮ In class work with probabilities Depending on the book that you select for this course, read either Gelman et al. Chapter 1 or Kruschke Chapters 1 & 2. C.J. Anderson (Illinois) Introduction Fall 2019 2.2/ 29 Introduction What Why Probability Steps Example History Practice Main References for Course Throughout the coures, I will take material from ◮ Gelman, A., Carlin, J.B., Stern, H.S., Dunson, D.B., Vehtari, A., & Rubin, D.B. (20114). Bayesian Data Analysis, 3rd Edition. Boco Raton, FL, CRC/Taylor & Francis.** ◮ Hoff, P.D., (2009). A First Course in Bayesian Statistical Methods. NY: Sringer.** ◮ McElreath, R.M. (2016). Statistical Rethinking: A Bayesian Course with Examples in R and Stan. Boco Raton, FL, CRC/Taylor & Francis. ◮ Kruschke, J.K. (2015). Doing Bayesian Data Analysis: A Tutorial with JAGS and Stan. NY: Academic Press.** ** There are e-versions these of from the UofI library. There is a verson of McElreath, but I couldn’t get if from UofI e-collection. C.J. Anderson (Illinois) Introduction Fall 2019 3.3/ 29 Introduction What Why Probability Steps Example History Practice Bayes Theorem A whole semester on this? p(y|θ)p(θ) p(θ|y)= p(y) where ◮ y is data, sample from some population.