Functional Investigation of the Potential Therapeutic Target Gene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Chr21 Protein-Protein Interactions: Enrichment in Products Involved in Intellectual Disabilities, Autism and Late Onset Alzheimer Disease
bioRxiv preprint doi: https://doi.org/10.1101/2019.12.11.872606; this version posted December 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Chr21 protein-protein interactions: enrichment in products involved in intellectual disabilities, autism and Late Onset Alzheimer Disease Julia Viard1,2*, Yann Loe-Mie1*, Rachel Daudin1, Malik Khelfaoui1, Christine Plancon2, Anne Boland2, Francisco Tejedor3, Richard L. Huganir4, Eunjoon Kim5, Makoto Kinoshita6, Guofa Liu7, Volker Haucke8, Thomas Moncion9, Eugene Yu10, Valérie Hindie9, Henri Bléhaut11, Clotilde Mircher12, Yann Herault13,14,15,16,17, Jean-François Deleuze2, Jean- Christophe Rain9, Michel Simonneau1, 18, 19, 20** and Aude-Marie Lepagnol- Bestel1** 1 Centre Psychiatrie & Neurosciences, INSERM U894, 75014 Paris, France 2 Laboratoire de génomique fonctionnelle, CNG, CEA, Evry 3 Instituto de Neurociencias CSIC-UMH, Universidad Miguel Hernandez-Campus de San Juan 03550 San Juan (Alicante), Spain 4 Department of Neuroscience, The Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA 5 Center for Synaptic Brain Dysfunctions, Institute for Basic Science, Daejeon 34141, Republic of Korea 6 Department of Molecular Biology, Division of Biological Science, Nagoya University Graduate School of Science, Furo, Chikusa, Nagoya, Japan 7 Department of Biological Sciences, University of Toledo, Toledo, OH, 43606, USA 8 Leibniz Forschungsinstitut für Molekulare Pharmakologie -

Neural Stem Cell-Derived Exosomes Revert HFD-Dependent Memory Impairment Via CREB-BDNF Signalling
International Journal of Molecular Sciences Article Neural Stem Cell-Derived Exosomes Revert HFD-Dependent Memory Impairment via CREB-BDNF Signalling Matteo Spinelli 1, Francesca Natale 1,2, Marco Rinaudo 1, Lucia Leone 1,2, Daniele Mezzogori 1, Salvatore Fusco 1,2,* and Claudio Grassi 1,2 1 Department of Neuroscience, Università Cattolica del Sacro Cuore, 00168 Rome, Italy; [email protected] (M.S.); [email protected] (F.N.); [email protected] (M.R.); [email protected] (L.L.); [email protected] (D.M.); [email protected] (C.G.) 2 Fondazione Policlinico Universitario Agostino Gemelli IRCCS, 00168 Rome, Italy * Correspondence: [email protected] Received: 6 October 2020; Accepted: 25 November 2020; Published: 26 November 2020 Abstract: Overnutrition and metabolic disorders impair cognitive functions through molecular mechanisms still poorly understood. In mice fed with a high fat diet (HFD) we analysed the expression of synaptic plasticity-related genes and the activation of cAMP response element-binding protein (CREB)-brain-derived neurotrophic factor (BDNF)-tropomyosin receptor kinase B (TrkB) signalling. We found that a HFD inhibited both CREB phosphorylation and the expression of a set of CREB target genes in the hippocampus. The intranasal administration of neural stem cell (NSC)-derived exosomes (exo-NSC) epigenetically restored the transcription of Bdnf, nNOS, Sirt1, Egr3, and RelA genes by inducing the recruitment of CREB on their regulatory sequences. Finally, exo-NSC administration rescued both BDNF signalling and memory in HFD mice. Collectively, our findings highlight novel mechanisms underlying HFD-related memory impairment and provide evidence of the potential therapeutic effect of exo-NSC against metabolic disease-related cognitive decline. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Genetic and Neurodevelopmental Spectrum Of
Cognitive and behavioural genetics J Med Genet: first published as 10.1136/jmedgenet-2015-103451 on 17 March 2016. Downloaded from ORIGINAL ARTICLE Genetic and neurodevelopmental spectrum of SYNGAP1-associated intellectual disability and epilepsy Cyril Mignot,1,2,3 Celina von Stülpnagel,4,5 Caroline Nava,1,6 Dorothée Ville,7 Damien Sanlaville,8,9,10 Gaetan Lesca,8,9,10 Agnès Rastetter,6 Benoit Gachet,6 Yannick Marie,6 G Christoph Korenke,11 Ingo Borggraefe,12 Dorota Hoffmann-Zacharska,13 Elżbieta Szczepanik,14 Mariola Rudzka-Dybała,14 Uluç Yiş,15 Hande Çağlayan,16 Arnaud Isapof,17 Isabelle Marey,1 Eleni Panagiotakaki,18 Christian Korff,19 Eva Rossier,20 Angelika Riess,21 Stefanie Beck-Woedl,21 Anita Rauch,22 Christiane Zweier,23 Juliane Hoyer,23 André Reis,23 Mikhail Mironov,24 Maria Bobylova,24 Konstantin Mukhin,24 Laura Hernandez-Hernandez,25 Bridget Maher,25 Sanjay Sisodiya,25 Marius Kuhn,26 Dieter Glaeser,26 Sarah Weckhuysen,6,27 Candace T Myers,28 Heather C Mefford,28 Konstanze Hörtnagel,29 Saskia Biskup,29 EuroEPINOMICS-RES MAE working group, Johannes R Lemke,30 Delphine Héron,1,2,3,4 Gerhard Kluger,4,5 Christel Depienne1,6 ▸ Additional material is ABSTRACT INTRODUCTION published online only. To view Objective We aimed to delineate the neurodevelopmental The human SYNGAP1 gene on chromosome please visit the journal online (http://dx.doi.org/10.1136/ spectrum associated with SYNGAP1 mutations and to 6p21.3 encodes the synaptic RAS-GTPase-activating jmedgenet-2015-103451). investigate genotype–phenotype correlations. protein 1, a protein of the post-synaptic density Methods We sequenced the exome or screened the exons (PSD) of glutamatergic neurons.12SYNGAP1 inter- For numbered affiliations see end of article. -

Downloads/ (Accessed on 17 January 2020)
cells Review Novel Approaches for Identifying the Molecular Background of Schizophrenia Arkadiy K. Golov 1,2,*, Nikolay V. Kondratyev 1 , George P. Kostyuk 3 and Vera E. Golimbet 1 1 Mental Health Research Center, 34 Kashirskoye shosse, 115522 Moscow, Russian; [email protected] (N.V.K.); [email protected] (V.E.G.) 2 Institute of Gene Biology, Russian Academy of Sciences, 34/5 Vavilova Street, 119334 Moscow, Russian 3 Alekseev Psychiatric Clinical Hospital No. 1, 2 Zagorodnoye shosse, 115191 Moscow, Russian; [email protected] * Correspondence: [email protected] Received: 5 November 2019; Accepted: 16 January 2020; Published: 18 January 2020 Abstract: Recent advances in psychiatric genetics have led to the discovery of dozens of genomic loci associated with schizophrenia. However, a gap exists between the detection of genetic associations and understanding the underlying molecular mechanisms. This review describes the basic approaches used in the so-called post-GWAS studies to generate biological interpretation of the existing population genetic data, including both molecular (creation and analysis of knockout animals, exploration of the transcriptional effects of common variants in human brain cells) and computational (fine-mapping of causal variability, gene set enrichment analysis, partitioned heritability analysis) methods. The results of the crucial studies, in which these approaches were used to uncover the molecular and neurobiological basis of the disease, are also reported. Keywords: schizophrenia; GWAS; causal genetic variants; enhancers; brain epigenomics; genome/epigenome editing 1. Introduction Schizophrenia is a severe mental illness that affects between 0.5% and 0.7% of the human population [1]. Both environmental and genetic factors are thought to be involved in its pathogenesis, with genetic factors playing a key role in disease risk, as the heritability of schizophrenia is estimated to be 70–85% [2,3]. -

BDNF Integrity in Ageing and Stress
MOJ Gerontology & Geriatrics Editorial Open Access BDNF integrity in ageing and stress Editorial Volume 1 Issue 6 - 2017 Alzheimer’s disease (AD) presents a progressive, stage-dependent Trevor Archer age-related neurodegenerative disorder that is characterized by Department of Psychology, University of Gothenburg, Sweden aggregation of toxic forms of amyloid β peptide (Aβ). There is much support for the notion that that brain-derived neurotrophic factor Correspondence: Trevor Archer, Department of Psychology, (BDNF) mediates beneficial effects of exercise on neuroplasticity and University of Gothenburg, Sweden, cellular stress resistance.1‒3 BDNF is associated with neuroplasticity Email [email protected] changes promoting health and well-being whereas these influences Received: July 16, 2017 | Published: August 01, 2017 may be opposed by pro-inflammatory cytokines, key factors in neurodegenerative processes.4 Lower serum levels are linked to greater symptom, cognitive, impairment.5 Thus, interventions, as for example physical exercise, whether acute or chronic, endurance or resistance, promote the mobilization of BDNF and other neurotrophic factors6,7 increase hippocampal and other brain regional resilience8 and therewith progression under conditions of stress when confronted critical life protect against cell death by inducing cell proliferation and maturation episodes or periods, appears to be modulated by the availability and with enhanced neuronal reparation, neurogenesis and the growth and accessibility of BDNF.20‒22 Over a wide range of neurologic and functioning of neurons in neurodegenerative disorders.9,10 Sampaio et psychiatric disease states it is found that the integrity and concentrations al.11 in a review of the therapeutic applications of neurotrophic factors, of brain, serum and plasma BDNF is compromised, particularly under particularly BDNF, postulate that they are essential for survival, stressful conditions. -
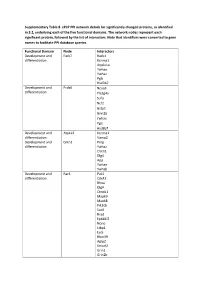
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Artificial Intelligence-Based Computational Framework for Drug
Tsuji et al. Alzheimer’s Research & Therapy (2021) 13:92 https://doi.org/10.1186/s13195-021-00826-3 RESEARCH Open Access Artificial intelligence-based computational framework for drug-target prioritization and inference of novel repositionable drugs for Alzheimer’s disease Shingo Tsuji1*† , Takeshi Hase2,3,4,5†, Ayako Yachie-Kinoshita2,4, Taiko Nishino2,SamikGhosh2, Masataka Kikuchi6, Kazuro Shimokawa7, Hiroyuki Aburatani1, Hiroaki Kitano2 and Hiroshi Tanaka3 Abstract Background: Identifying novel therapeutic targets is crucial for the successful development of drugs. However, the cost to experimentally identify therapeutic targets is huge and only approximately 400 genes are targets for FDA-approved drugs. As a result, it is inevitable to develop powerful computational tools that can identify potential novel therapeutic targets. Fortunately, the human protein-protein interaction network (PIN) could be a useful resource to achieve this objective. Methods: In this study, we developed a deep learning-based computational framework that extracts low-dimensional representations of high-dimensional PIN data. Our computational framework uses latent features and state-of-the-art machine learning techniques to infer potential drug target genes. Results: We applied our computational framework to prioritize novel putative target genes for Alzheimer’s disease and successfully identified key genes that may serve as novel therapeutic targets (e.g., DLG4, EGFR, RAC1, SYK, PTK2B, SOCS1). Furthermore, based on these putative targets, we could infer repositionable -

47421 SAP102 (A7R8L) Rabbit Mab
Revision 1 C 0 2 - t SAP102 (A7R8L) Rabbit mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] 1 Support: 877-678-TECH (8324) 2 4 Web: [email protected] 7 www.cellsignal.com 4 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IP H M R Endogenous 102 Rabbit Q92796 1741 Product Usage Information 1. Cuthbert, P.C. et al. (2007) J Neurosci 27, 2673-82. 2. Makino, K. et al. (1997) Oncogene 14, 2425-33. Application Dilution 3. Masuko, N. et al. (1999) J Biol Chem 274, 5782-90. 4. Lau, L.F. et al. (1996) J Biol Chem 271, 21622-8. Western Blotting 1:1000 5. Yan, J. et al. (2005) Mol Psychiatry 10, 329-32. Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity SAP102 (A7R8L) Rabbit mAb recognizes endogenous levels of total SAP102 protein. Species Reactivity: Human, Mouse, Rat Source / Purification Monoclonal antibody is produced by immunizing animals with a synthetic peptide corresponding to residues near the carboxy terminus of human SAP102 protein. Background Synapse-associated protein 102 (SAP102, DLG3) belongs to the membrane-associated guanylate kinase (MAGUK) protein family and is a homolog of the Drosophila disc large (dlg) tumor suppressor protein. SAP102 consists of three PDZ domains, a Src homology 3 (SH3) domain, and a guanylate kinase (GK) domain (1). -

Loss of RET Promotes Mesenchymal Identity in Neuroblastoma Cells
cancers Article Loss of RET Promotes Mesenchymal Identity in Neuroblastoma Cells Joachim T. Siaw 1,†, Jonatan L. Gabre 1,2,† , Ezgi Uçkun 1 , Marc Vigny 3, Wancun Zhang 4, Jimmy Van den Eynden 2 , Bengt Hallberg 1 , Ruth H. Palmer 1 and Jikui Guan 1,4,* 1 Department of Medical Biochemistry and Cell Biology, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg, SE-40530 Gothenburg, Sweden; [email protected] (J.T.S.); [email protected] (J.L.G.); [email protected] (E.U.); [email protected] (B.H.); [email protected] (R.H.P.) 2 Anatomy and Embryology Unit, Department of Human Structure and Repair, Ghent University, 9000 Ghent, Belgium; [email protected] 3 Université Pierre et Marie Curie, UPMC, INSERM UMRS-839, 75005 Paris, France; [email protected] 4 Department of Pediatric Oncology Surgery, Children’s Hospital Affiliated to Zhengzhou University, Zhengzhou 450018, China; [email protected] * Correspondence: [email protected] † These authors contributed equally to this work. Simple Summary: The anaplastic lymphoma kinase (ALK) and rearranged during transfection (RET) receptor tyrosine kinases (RTKs) are expressed in both the developing neural crest and the pediatric cancer neuroblastoma. Moreover, ALK is mutated in approximately 10% of neuroblastomas. Here, we investigated ALK and RET in neuroblastoma, with the aim of better understanding their respective contributions. Using neuroblastoma cell lines, we show that ALK modulates RET signaling at the level of RET phosphorylation, as well as at the level of transcription. Using CRISPR/Cas9, we generated Citation: Siaw, J.T.; Gabre, J.L.; Uçkun, E.; Vigny, M.; Zhang, W.; Van RET knockout neuroblastoma cell lines and performed a multi-omics approach, combining RNA-Seq den Eynden, J.; Hallberg, B.; Palmer, and proteomics to characterize the effect of deleting RET in a neuroblastoma context. -

LGI1–ADAM22–MAGUK Configures Transsynaptic Nanoalignment for Synaptic Transmission and Epilepsy Prevention
LGI1–ADAM22–MAGUK configures transsynaptic nanoalignment for synaptic transmission and epilepsy prevention Yuko Fukataa,b,1, Xiumin Chenc,d,1, Satomi Chikenb,e, Yoko Hiranoa,f, Atsushi Yamagatag, Hiroki Inahashia, Makoto Sanboh, Hiromi Sanob,e, Teppei Gotoh, Masumi Hirabayashib,h, Hans-Christian Kornaui,j, Harald Prüssi,k, Atsushi Nambub,e, Shuya Fukail, Roger A. Nicollc,d,2, and Masaki Fukataa,b,2 aDivision of Membrane Physiology, Department of Molecular and Cellular Physiology, National Institute for Physiological Sciences, National Institutes of Natural Sciences, Aichi 444-8787, Japan; bDepartment of Physiological Sciences, School of Life Science, SOKENDAI (The Graduate University for Advanced Studies), Aichi 444-8585, Japan; cDepartment of Cellular and Molecular Pharmacology, University of California, San Francisco, CA 94158; dDepartment of Physiology, University of California, San Francisco, CA 94158; eDivision of System Neurophysiology, Department of System Neuroscience, National Institute for Physiological Sciences, National Institutes of Natural Sciences, Aichi 444-8585, Japan; fDepartment of Pediatrics, Graduate School of Medicine, The University of Tokyo, Tokyo 113-8655, Japan; gLaboratory for Protein Functional and Structural Biology, RIKEN Center for Biosystems Dynamics Research, Kanagawa 230-0045, Japan; hCenter for Genetic Analysis of Behavior, National Institute for Physiological Sciences, National Institutes of Natural Sciences, Okazaki 444-8787, Japan; iGerman Center for Neurodegenerative Diseases (DZNE), 10117 Berlin, Germany; jNeuroscience Research Center, Cluster NeuroCure, Charité-Universitätsmedizin Berlin, 10117 Berlin, Germany; kDepartment of Neurology and Experimental Neurology, Charité-Universitätsmedizin Berlin, 10117 Berlin, Germany; and lDepartment of Chemistry, Graduate School of Science, Kyoto University, 606-8502 Kyoto, Japan Contributed by Roger A. Nicoll, December 1, 2020 (sent for review October 29, 2020; reviewed by David S. -

Host Cell Factors Necessary for Influenza a Infection: Meta-Analysis of Genome Wide Studies
Host Cell Factors Necessary for Influenza A Infection: Meta-Analysis of Genome Wide Studies Juliana S. Capitanio and Richard W. Wozniak Department of Cell Biology, Faculty of Medicine and Dentistry, University of Alberta Abstract: The Influenza A virus belongs to the Orthomyxoviridae family. Influenza virus infection occurs yearly in all countries of the world. It usually kills between 250,000 and 500,000 people and causes severe illness in millions more. Over the last century alone we have seen 3 global influenza pandemics. The great human and financial cost of this disease has made it the second most studied virus today, behind HIV. Recently, several genome-wide RNA interference studies have focused on identifying host molecules that participate in Influen- za infection. We used nine of these studies for this meta-analysis. Even though the overlap among genes identified in multiple screens was small, network analysis indicates that similar protein complexes and biological functions of the host were present. As a result, several host gene complexes important for the Influenza virus life cycle were identified. The biological function and the relevance of each identified protein complex in the Influenza virus life cycle is further detailed in this paper. Background and PA bound to the viral genome via nucleoprotein (NP). The viral core is enveloped by a lipid membrane derived from Influenza virus the host cell. The viral protein M1 underlies the membrane and anchors NEP/NS2. Hemagglutinin (HA), neuraminidase Viruses are the simplest life form on earth. They parasite host (NA), and M2 proteins are inserted into the envelope, facing organisms and subvert the host cellular machinery for differ- the viral exterior.