A Machine Learning Workflow for Constructing and Comparing Transcriptome-Wide Gene Regulatory Networks from Single- Cell Data
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Ck1δ Over-Expressing Mice Display ADHD-Like Behaviors, Frontostriatal Neuronal Abnormalities and Altered Expressions of ADHD-Candidate Genes
Molecular Psychiatry (2020) 25:3322–3336 https://doi.org/10.1038/s41380-018-0233-z ARTICLE CK1δ over-expressing mice display ADHD-like behaviors, frontostriatal neuronal abnormalities and altered expressions of ADHD-candidate genes 1 1 1 2 1 1 1 Mingming Zhou ● Jodi Gresack ● Jia Cheng ● Kunihiro Uryu ● Lars Brichta ● Paul Greengard ● Marc Flajolet Received: 8 November 2017 / Revised: 4 July 2018 / Accepted: 18 July 2018 / Published online: 19 October 2018 © Springer Nature Limited 2018 Abstract The cognitive mechanisms underlying attention-deficit hyperactivity disorder (ADHD), a highly heritable disorder with an array of candidate genes and unclear genetic architecture, remain poorly understood. We previously demonstrated that mice overexpressing CK1δ (CK1δ OE) in the forebrain show hyperactivity and ADHD-like pharmacological responses to D- amphetamine. Here, we demonstrate that CK1δ OE mice exhibit impaired visual attention and a lack of D-amphetamine- induced place preference, indicating a disruption of the dopamine-dependent reward pathway. We also demonstrate the presence of abnormalities in the frontostriatal circuitry, differences in synaptic ultra-structures by electron microscopy, as 1234567890();,: 1234567890();,: well as electrophysiological perturbations of both glutamatergic and GABAergic transmission, as observed by altered frequency and amplitude of mEPSCs and mIPSCs. Furthermore, gene expression profiling by next-generation sequencing alone, or in combination with bacTRAP technology to study specifically Drd1a versus Drd2 medium spiny neurons, revealed that developmental CK1δ OE alters transcriptional homeostasis in the striatum, including specific alterations in Drd1a versus Drd2 neurons. These results led us to perform a fine molecular characterization of targeted gene networks and pathway analysis. Importantly, a large fraction of 92 genes identified by GWAS studies as associated with ADHD in humans are significantly altered in our mouse model. -

Signal Transduction in L-DOPA-Induced Dyskinesia: from Receptor Sensitization to Abnormal… Et Al
Journal of Neural Transmission https://doi.org/10.1007/s00702-018-1847-7 NEUROLOGY AND PRECLINICAL NEUROLOGICAL STUDIES - REVIEW ARTICLE Signal transduction in L‑DOPA‑induced dyskinesia: from receptor sensitization to abnormal gene expression Giada Spigolon1 · Gilberto Fisone1 Received: 1 November 2017 / Accepted: 23 January 2018 © The Author(s) 2018. This article is an open access publication Abstract A large number of signaling abnormalities have been implicated in the emergence and expression of L-DOPA-induced dyskinesia (LID). The primary cause for many of these changes is the development of sensitization at dopamine receptors located on striatal projection neurons (SPN). This initial priming, which is particularly evident at the level of dopamine D1 receptors (D1R), can be viewed as a homeostatic response to dopamine depletion and is further exacerbated by chronic administration of L-DOPA, through a variety of mechanisms afecting various components of the G-protein-coupled receptor machinery. Sensitization of dopamine receptors in combination with pulsatile administration of L-DOPA leads to intermit- tent and coordinated hyperactivation of signal transduction cascades, ultimately resulting in long-term modifcations of gene expression and protein synthesis. A detailed mapping of these pathological changes and of their involvement in LID has been produced during the last decade. According to this emerging picture, activation of sensitized D1R results in the stimulation of cAMP-dependent protein kinase and of the dopamine- and cAMP-regulated phosphoprotein of 32 kDa. This, in turn, activates the extracellular signal-regulated kinases 1 and 2 (ERK), leading to chromatin remodeling and aberrant gene transcription. Dysregulated ERK results also in the stimulation of the mammalian target of rapamycin complex 1, which promotes protein synthesis. -
Supplemental Data Olig2-Regulated Lineage-Restricted Pathway Controls Replication Competence in Neural Stem Cells and Malignant
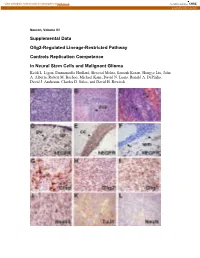
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Caltech Authors Neuron, Volume 53 Supplemental Data Olig2-Regulated Lineage-Restricted Pathway Controls Replication Competence in Neural Stem Cells and Malignant Glioma Keith L. Ligon, Emmanuelle Huillard, Shwetal Mehta, Santosh Kesari, Hongye Liu, John A. Alberta, Robert M. Bachoo, Michael Kane, David N. Louis, Ronald A. DePinho, David J. Anderson, Charles D. Stiles, and David H. Rowitch Figure S1. Olig1/2+/- Ink4a/Arf-/-EGFRvIII Gliomas Express Characteristic Morphologic and Immunophenotypic Features of Human Malignant Gliomas (A) Tumors exhibit dense cellularity and atypia. (B) Characteristic infiltration of host SVZ stem cell niche. (C) Although occasional tumors exhibited pseudopalisading necrosis (n, arrowhead) and hemorrhage (h, arrowhead) similar to human GBM (Astrocytoma WHO Grade IV), most tumors lacked these features. (D) IHC for hEGFR highlights hallmark feature of human gliomas, including perivascular (pv) and subpial (sp) accumulation of tumor cells, (E) striking white matter tropism of tumor cells in corpus callosum (cc), and (F) distant single cell infiltration of cerebellar white matter (wm, arrows). (G-J) Immunohistochemical markers characteristic of human tumors (Gfap, Olig2, Olig1, Nestin) are present. (K and L) Weak staining for the early neuronal marker TuJ1 and absence of the differentiated neuronal marker, NeuN similar to human glioma and consistent with heterogeneous lines of partial differentiation. Figure S2. Olig1/2+/- Ink4a/Arf-/-EGFRvIII Neurospheres Are Multipotent and Olig1/2-/- Ink4a/Arf-/-EGFRvIII Neurospheres Are Bipotent Olig1/2+/- or Olig1/2-/- Ink4a/Arf-/-EGFRvIII neurospheres were allowed to differentiate for 6 days in medium without EGF. -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

I FOUR JOINTED BOX ONE, a NOVEL PRO-ANGIOGENIC PROTEIN IN
FOUR JOINTED BOX ONE, A NOVEL PRO-ANGIOGENIC PROTEIN IN COLORECTAL CARCINOMA. BY Nicole Theresa Al-Greene Dissertation Submitted to the Faculty of the Graduate School of Vanderbilt University in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY In Cell and Developmental Biology. December, 2013 Nashville Tennessee Approved: R. Daniel Beauchamp Susan Wente James Goldenring Albert Reynolds i DEDICATION To my parents, Karen and John, who have helped me in every way possible, every single day. ii ACKNOWLEDGMENTS. Funding for this work was supported by grants DK052334, CA069457, The GI Cancer SPORE, GM088822, the VICC, the Clinical and Translational Science Award (NCRR/NIH UL1RR024975), the DDRC (P30DK058404), and the Cooperative Human Tissue Network (UO1CA094664) and U01CA094664. I was lucky enough to be allowed to perform research as an undergraduate in the lab of Ken Belanger. I will be forever grateful for that opportunity that sparked my love of research. Equally important was my time as a technician in Len Zon’s lab where I confirmed the fact that I needed to go graduate school and earn my degree. During my time at Vanderbilt I have been helped by so many individuals, and the collaborative nature of everyone I have met with is truly an amazing aspect of the research community here. I am especially thankful for all the technical help and insightful conversations I have had with Natasha Deane, Anna Means, Claudia Andl, Tanner Freeman, Connie Weaver, Keeli Lewis, Jalal Hamaamen, Jenny Zi, John Neff, Christian Kis, Andries Zjistra, Trennis Palmer, Joseph Roland, and Lynn LaPierre. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Suppression of Rgsz1 Function Optimizes the Actions of Opioid
Suppression of RGSz1 function optimizes the actions PNAS PLUS of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway Sevasti Gasparia,b,c, Immanuel Purushothamana,b, Valeria Cogliania,b, Farhana Saklotha,b, Rachael L. Neved, David Howlande, Robert H. Ringf, Elliott M. Rossg,h, Li Shena,b, and Venetia Zacharioua,b,1 aFishberg Department of Neuroscience, Icahn School of Medicine at Mount Sinai, New York, NY 10029; bFriedman Brain Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029; cDepartment of Basic Sciences, University of Crete Medical School, 71003 Heraklion, Greece; dGene Delivery Technology Core, Massachusetts General Hospital, MA 01239; eCure Huntington’s Disease Initiative (CHDI) Foundation, Princeton, NJ 08540; fDepartment of Psychiatry, Icahn School of Medicine at Mount Sinai, New York, NY 10029; gDepartment of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390; and hGreen Center for Systems Biology, University of Texas Southwestern Medical Center, Dallas, TX 75390 Edited by Solomon H. Snyder, Johns Hopkins University School of Medicine, Baltimore, MD, and approved January 5, 2018 (received for review June 23, 2017) Regulator of G protein signaling z1 (RGSz1), a member of the RGS studies have documented brain region-specific effects of other family of proteins, is present in several networks expressing mu RGS proteins (RGS9-2, RGS7, and RGS4) (29–33) on the opioid receptors (MOPRs). By using genetic mouse models for global functional responses of MOPRs, the role of RGSz1 in opioid or brain region-targeted manipulations of RGSz1 expression, we actions is not known. We hypothesized that the expression and demonstrated that the suppression of RGSz1 function increases the activity of RGSz1 are altered by chronic opioid use and that the analgesic efficacy of MOPR agonists in male and female mice and manipulation of RGSz1 expression will impact the development delays the development of morphine tolerance while decreasing the of tolerance to the drug and its analgesic and rewarding effects. -

Small Molecule Protein–Protein Interaction Inhibitors As CNS Therapeutic Agents: Current Progress and Future Hurdles
Neuropsychopharmacology REVIEWS (2009) 34, 126–141 & 2009 Nature Publishing Group All rights reserved 0893-133X/09 $30.00 ............................................................................................................................................................... REVIEW 126 www.neuropsychopharmacology.org Small Molecule Protein–Protein Interaction Inhibitors as CNS Therapeutic Agents: Current Progress and Future Hurdles 1 ,1,2,3 Levi L Blazer and Richard R Neubig* 1Department of Pharmacology, University of Michigan Medical School, Ann Arbor, MI, USA; 2Department of Internal Medicine, University of Michigan Medical School, Ann Arbor, MI, USA; 3Center for Chemical Genomics, University of Michigan Medical School, Ann Arbor, MI, USA Protein–protein interactions are a crucial element in cellular function. The wealth of information currently available on intracellular-signaling pathways has led many to appreciate the untapped pool of potential drug targets that reside downstream of the commonly targeted receptors. Over the last two decades, there has been significant interest in developing therapeutics and chemical probes that inhibit specific protein–protein interactions. Although it has been a challenge to develop small molecules that are capable of occluding the large, often relatively featureless protein–protein interaction interface, there are increasing numbers of examples of small molecules that function in this manner with reasonable potency. This article will highlight the current progress in the development of small -

Regulators of G-Protein Signaling (RGS) Proteins: Region-Specific Expression of Nine Subtypes in Rat Brain
The Journal of Neuroscience, October 15, 1997, 17(20):8024–8037 Regulators of G-Protein Signaling (RGS) Proteins: Region-Specific Expression of Nine Subtypes in Rat Brain Stephen J. Gold,1,2 YanG.Ni,1,2 Henrik G. Dohlman,3 and Eric J. Nestler1,2,3 1Laboratory of Molecular Psychiatry and Departments of 2Psychiatry and 3Pharmacology, Yale University School of Medicine, New Haven, Connecticut 06508 The recently discovered regulators of G-protein signaling (RGS) richment in striatal regions. RGS10 mRNA is densely expressed proteins potently modulate the functioning of heterotrimeric in dentate gyrus granule cells, superficial layers of neocortex, G-proteins by stimulating the GTPase activity of G-protein a and dorsal raphe. To assess whether the expression of RGS subunits. The mRNAs for numerous subtypes of putative RGS mRNAs can be regulated, we examined the effect of an acute proteins have been identified in mammalian tissues, but little is seizure on levels of RGS7, RGS8, and RGS10 mRNAs in hip- known about their expression in brain. We performed a system- pocampus. Of the three subtypes, changes in RGS10 levels atic survey of the localization of mRNAs encoding nine of these were most pronounced, decreasing by ;40% in a time- RGSs, RGS3–RGS11, in brain by in situ hybridization. Striking dependent manner in response to a single seizure. These re- region-specific patterns of expression were observed. Five sults, which document highly specific patterns of RGS mRNA subtypes, RGS4, RGS7, RGS8, RGS9, and RGS10 mRNAs, are expression in brain and their ability to be regulated in a dynamic densely expressed in brain, whereas the other subtypes (RGS3, manner, support the view that RGS proteins may play an im- RGS5, RGS6, and RGS11) are expressed at lower density and portant role in determining the intensity and specificity of sig- in more restricted regions. -

1 Novel Expression Signatures Identified by Transcriptional Analysis
ARD Online First, published on October 7, 2009 as 10.1136/ard.2009.108043 Ann Rheum Dis: first published as 10.1136/ard.2009.108043 on 7 October 2009. Downloaded from Novel expression signatures identified by transcriptional analysis of separated leukocyte subsets in SLE and vasculitis 1Paul A Lyons, 1Eoin F McKinney, 1Tim F Rayner, 1Alexander Hatton, 1Hayley B Woffendin, 1Maria Koukoulaki, 2Thomas C Freeman, 1David RW Jayne, 1Afzal N Chaudhry, and 1Kenneth GC Smith. 1Cambridge Institute for Medical Research and Department of Medicine, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK 2Roslin Institute, University of Edinburgh, Roslin, Midlothian, EH25 9PS, UK Correspondence should be addressed to Dr Paul Lyons or Prof Kenneth Smith, Department of Medicine, Cambridge Institute for Medical Research, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK. Telephone: +44 1223 762642, Fax: +44 1223 762640, E-mail: [email protected] or [email protected] Key words: Gene expression, autoimmune disease, SLE, vasculitis Word count: 2,906 The Corresponding Author has the right to grant on behalf of all authors and does grant on behalf of all authors, an exclusive licence (or non-exclusive for government employees) on a worldwide basis to the BMJ Publishing Group Ltd and its Licensees to permit this article (if accepted) to be published in Annals of the Rheumatic Diseases and any other BMJPGL products to exploit all subsidiary rights, as set out in their licence (http://ard.bmj.com/ifora/licence.pdf). http://ard.bmj.com/ on September 29, 2021 by guest. Protected copyright. 1 Copyright Article author (or their employer) 2009.