Complete Genome Sequence of the Phenanthrene-Degrading Soil Bacterium Delftia Acidovorans Cs1-4 Ameesha R
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The 2014 Golden Gate National Parks Bioblitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event
National Park Service U.S. Department of the Interior Natural Resource Stewardship and Science The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 ON THIS PAGE Photograph of BioBlitz participants conducting data entry into iNaturalist. Photograph courtesy of the National Park Service. ON THE COVER Photograph of BioBlitz participants collecting aquatic species data in the Presidio of San Francisco. Photograph courtesy of National Park Service. The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 Elizabeth Edson1, Michelle O’Herron1, Alison Forrestel2, Daniel George3 1Golden Gate Parks Conservancy Building 201 Fort Mason San Francisco, CA 94129 2National Park Service. Golden Gate National Recreation Area Fort Cronkhite, Bldg. 1061 Sausalito, CA 94965 3National Park Service. San Francisco Bay Area Network Inventory & Monitoring Program Manager Fort Cronkhite, Bldg. 1063 Sausalito, CA 94965 March 2016 U.S. Department of the Interior National Park Service Natural Resource Stewardship and Science Fort Collins, Colorado The National Park Service, Natural Resource Stewardship and Science office in Fort Collins, Colorado, publishes a range of reports that address natural resource topics. These reports are of interest and applicability to a broad audience in the National Park Service and others in natural resource management, including scientists, conservation and environmental constituencies, and the public. The Natural Resource Report Series is used to disseminate comprehensive information and analysis about natural resources and related topics concerning lands managed by the National Park Service. -

Downloaded 10/5/2021 9:43:05 AM
Chemical Science View Article Online EDGE ARTICLE View Journal | View Issue Aromatic side-chain flips orchestrate the conformational sampling of functional loops in Cite this: Chem. Sci.,2021,12,9318 † All publication charges for this article human histone deacetylase 8 have been paid for by the Royal Society a a bcd of Chemistry Vaibhav Kumar Shukla, ‡ Lucas Siemons, ‡ Francesco L. Gervasio and D. Flemming Hansen *a Human histone deacetylase 8 (HDAC8) is a key hydrolase in gene regulation and an important drug-target. High-resolution structures of HDAC8 in complex with substrates or inhibitors are available, which have provided insights into the bound state of HDAC8 and its function. Here, using long all-atom unbiased molecular dynamics simulations and Markov state modelling, we show a strong correlation between the conformation of aromatic side chains near the active site and opening and closing of the surrounding functional loops of HDAC8. We also investigated two mutants known to allosterically downregulate the enzymatic activity of HDAC8. Based on experimental data, we hypothesise that I19S-HDAC8 is unable to Received 6th April 2021 Creative Commons Attribution 3.0 Unported Licence. release the product, whereas both product release and substrate binding are impaired in the S39E- Accepted 27th May 2021 HDAC8 mutant. The presented results deliver detailed insights into the functional dynamics of HDAC8 DOI: 10.1039/d1sc01929e and provide a mechanism for the substantial downregulation caused by allosteric mutations, including rsc.li/chemical-science a disease causing one. Introduction II (HDAC-4, -5, -6, -7, -9, and HDAC10), and class III (SIRT1-7) have sequence similarity to yeast Rpd3, Hda1, and Sir2, Acetylation of lysine side chains occurs as a co-translation or respectively, whereas class IV (HDAC11) shares sequence simi- 2 This article is licensed under a post-translational modication of proteins and was rst iden- larity with both class I and II proteins. -
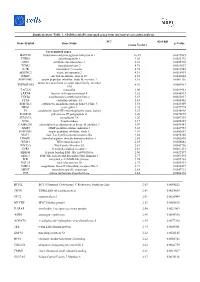
Supplementary Table 1. All Differentially Expressed Genes from Microarray Screening Analysis
Supplementary Table 1. All differentially expressed genes from microarray screening analysis. FCa (Gal-KD Gene Symbol Gene Name p-Value versus Vector) Up-regulated genes HAPLN1 hyaluronan and proteoglycan link protein 1 10.49 0.0027085 THBS1 thrombospondin 1 9.20 0.0022192 ODC1 ornithine decarboxylase 1 6.38 0.0055776 TGM2 transglutaminase 2 4.76 0.0015627 IL7R interleukin 7 receptor 4.75 0.0017245 SERINC2 serine incorporator 2 4.51 0.0014919 ITM2C integral membrane protein 2C 4.32 0.0044644 SERPINB7 serpin peptidase inhibitor, clade B, member 7 4.18 0.0081136 tumor necrosis factor receptor superfamily, member TNFRSF10D 4.01 0.0085561 10d TAGLN transgelin 3.90 0.0099963 LRRN4 leucine rich repeat neuronal 4 3.82 0.0046513 TGFB2 transforming growth factor beta 2 3.51 0.0035017 CPA4 carboxypeptidase A4 3.43 0.0008452 EPB41L3 erythrocyte membrane protein band 4.1-like 3 3.34 0.0025309 NRG1 neuregulin 1 3.28 0.0079724 F3 coagulation factor III (thromboplastin, tissue factor) 3.27 0.0038968 POLR3G polymerase III polypeptide G 3.26 0.0070675 SEMA7A semaphorin 7A 3.20 0.0087335 NT5E 5-nucleotidase 3.17 0.0036353 CAMK2N1 calmodulin-dependent protein kinase II inhibitor 1 3.07 0.0090141 TIMP3 TIMP metallopeptidase inhibitor 3 3.03 0.0047953 SERPINE1 serpin peptidase inhibitor, clade E 2.97 0.0053652 MALL mal, T-cell differentiation protein-like 2.88 0.0078205 DDAH1 dimethylarginine dimethylaminohydrolase 1 2.86 0.0002895 WDR3 WD repeat domain 3 2.85 0.0058842 WNT5A Wnt Family Member 5A 2.81 0.0043796 GPR1 G protein-coupled receptor 1 2.81 0.0021313 -

Which Organisms Are Used for Anti-Biofouling Studies
Table S1. Semi-systematic review raw data answering: Which organisms are used for anti-biofouling studies? Antifoulant Method Organism(s) Model Bacteria Type of Biofilm Source (Y if mentioned) Detection Method composite membranes E. coli ATCC25922 Y LIVE/DEAD baclight [1] stain S. aureus ATCC255923 composite membranes E. coli ATCC25922 Y colony counting [2] S. aureus RSKK 1009 graphene oxide Saccharomycetes colony counting [3] methyl p-hydroxybenzoate L. monocytogenes [4] potassium sorbate P. putida Y. enterocolitica A. hydrophila composite membranes E. coli Y FESEM [5] (unspecified/unique sample type) S. aureus (unspecified/unique sample type) K. pneumonia ATCC13883 P. aeruginosa BAA-1744 composite membranes E. coli Y SEM [6] (unspecified/unique sample type) S. aureus (unspecified/unique sample type) graphene oxide E. coli ATCC25922 Y colony counting [7] S. aureus ATCC9144 P. aeruginosa ATCCPAO1 composite membranes E. coli Y measuring flux [8] (unspecified/unique sample type) graphene oxide E. coli Y colony counting [9] (unspecified/unique SEM sample type) LIVE/DEAD baclight S. aureus stain (unspecified/unique sample type) modified membrane P. aeruginosa P60 Y DAPI [10] Bacillus sp. G-84 LIVE/DEAD baclight stain bacteriophages E. coli (K12) Y measuring flux [11] ATCC11303-B4 quorum quenching P. aeruginosa KCTC LIVE/DEAD baclight [12] 2513 stain modified membrane E. coli colony counting [13] (unspecified/unique colony counting sample type) measuring flux S. aureus (unspecified/unique sample type) modified membrane E. coli BW26437 Y measuring flux [14] graphene oxide Klebsiella colony counting [15] (unspecified/unique sample type) P. aeruginosa (unspecified/unique sample type) graphene oxide P. aeruginosa measuring flux [16] (unspecified/unique sample type) composite membranes E. -

Biochemical Characterization and Comparison of Aspartylglucosaminidases Secreted in Venom of the Parasitoid Wasps Asobara Tabida and Leptopilina Heterotoma
RESEARCH ARTICLE Biochemical characterization and comparison of aspartylglucosaminidases secreted in venom of the parasitoid wasps Asobara tabida and Leptopilina heterotoma Quentin Coulette1, SeÂverine Lemauf2, Dominique Colinet2, Geneviève PreÂvost1, Caroline Anselme1, Marylène Poirie 2, Jean-Luc Gatti2* a1111111111 1 Unite ªEcologie et Dynamique des Systèmes AnthropiseÂsº (EDYSAN, FRE 3498 CNRS-UPJV), Universite de Picardie Jules Verne, Amiens, France, 2 Universite CoÃte d'Azur, INRA, CNRS, ISA, Sophia Antipolis, a1111111111 France a1111111111 a1111111111 * [email protected] a1111111111 Abstract Aspartylglucosaminidase (AGA) is a low-abundance intracellular enzyme that plays a key OPEN ACCESS role in the last stage of glycoproteins degradation, and whose deficiency leads to human Citation: Coulette Q, Lemauf S, Colinet D, PreÂvost aspartylglucosaminuria, a lysosomal storage disease. Surprisingly, high amounts of AGA- G, Anselme C, Poirie M, et al. (2017) Biochemical characterization and comparison of like proteins are secreted in the venom of two phylogenetically distant hymenopteran para- aspartylglucosaminidases secreted in venom of the sitoid wasp species, Asobara tabida (Braconidae) and Leptopilina heterotoma (Cynipidae). parasitoid wasps Asobara tabida and Leptopilina These venom AGAs have a similar domain organization as mammalian AGAs. They share heterotoma. PLoS ONE 12(7): e0181940. https:// with them key residues for autocatalysis and activity, and the mature - and -subunits also doi.org/10.1371/journal.pone.0181940 α β form an (αβ)2 structure in solution. Interestingly, only one of these AGAs subunits (α for Editor: Erjun Ling, Institute of Plant Physiology and AtAGA and for LhAGA) is glycosylated instead of the two subunits for lysosomal human Ecology Shanghai Institutes for Biological β Sciences, CHINA AGA (hAGA), and these glycosylations are partially resistant to PGNase F treatment. -

Biosynthesis of Natural Products Containing Β-Amino Acids
Natural Product Reports Biosynthesis of natural products containing β -amino acids Journal: Natural Product Reports Manuscript ID: NP-REV-01-2014-000007.R1 Article Type: Review Article Date Submitted by the Author: 21-Apr-2014 Complete List of Authors: Kudo, Fumitaka; Tokyo Institute Of Technology, Department of Chemistry Miyanaga, Akimasa; Tokyo Institute Of Technology, Department of Chemistry Eguchi, T; Tokyo Institute Of Technology, Department of Chemistry and Materials Science Page 1 of 20 Natural Product Reports NPR RSC Publishing REVIEW Biosynthesis of natural products containing βββ- amino acids Cite this: DOI: 10.1039/x0xx00000x Fumitaka Kudo, a Akimasa Miyanaga, a and Tadashi Eguchi *b Received 00th January 2014, We focus here on β-amino acids as components of complex natural products because the presence of β-amino acids Accepted 00th January 2014 produces structural diversity in natural products and provides characteristic architectures beyond that of ordinary DOI: 10.1039/x0xx00000x α-L-amino acids, thus generating significant and unique biological functions in nature. In this review, we first survey the known bioactive β-amino acid-containing natural products including nonribosomal peptides, www.rsc.org/ macrolactam polyketides, and nucleoside-β-amino acid hybrids. Next, the biosynthetic enzymes that form β-amino acids from α-amino acids and de novo synthesis of β-amino acids are summarized. Then, the mechanisms of β- amino acid incorporation into natural products are reviewed. Because it is anticipated that the rational swapping of the β-amino acid moieties with various side chains and stereochemistries by biosynthetic engineering should lead to the creation of novel architectures and bioactive compounds, the accumulation of knowledge regarding β- amino acid-containing natural product biosynthetic machinery could have a significant impact in this field. -

The Study on the Cultivable Microbiome of the Aquatic Fern Azolla Filiculoides L
applied sciences Article The Study on the Cultivable Microbiome of the Aquatic Fern Azolla Filiculoides L. as New Source of Beneficial Microorganisms Artur Banach 1,* , Agnieszka Ku´zniar 1, Radosław Mencfel 2 and Agnieszka Woli ´nska 1 1 Department of Biochemistry and Environmental Chemistry, The John Paul II Catholic University of Lublin, 20-708 Lublin, Poland; [email protected] (A.K.); [email protected] (A.W.) 2 Department of Animal Physiology and Toxicology, The John Paul II Catholic University of Lublin, 20-708 Lublin, Poland; [email protected] * Correspondence: [email protected]; Tel.: +48-81-454-5442 Received: 6 May 2019; Accepted: 24 May 2019; Published: 26 May 2019 Abstract: The aim of the study was to determine the still not completely described microbiome associated with the aquatic fern Azolla filiculoides. During the experiment, 58 microbial isolates (43 epiphytes and 15 endophytes) with different morphologies were obtained. We successfully identified 85% of microorganisms and assigned them to 9 bacterial genera: Achromobacter, Bacillus, Microbacterium, Delftia, Agrobacterium, and Alcaligenes (epiphytes) as well as Bacillus, Staphylococcus, Micrococcus, and Acinetobacter (endophytes). We also studied an A. filiculoides cyanobiont originally classified as Anabaena azollae; however, the analysis of its morphological traits suggests that this should be renamed as Trichormus azollae. Finally, the potential of the representatives of the identified microbial genera to synthesize plant growth-promoting substances such as indole-3-acetic acid (IAA), cellulase and protease enzymes, siderophores and phosphorus (P) and their potential of utilization thereof were checked. Delftia sp. AzoEpi7 was the only one from all the identified genera exhibiting the ability to synthesize all the studied growth promoters; thus, it was recommended as the most beneficial bacteria in the studied microbiome. -

Active Site Tyrosine Is Essential for Amidohydrolase but Not for Esterase
Active site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme Kristin Moreth, Daniel Riester, Christian Hildmann, René Hempel, Dennis Wegener, Andreas Schober, Andreas Schwienhorst To cite this version: Kristin Moreth, Daniel Riester, Christian Hildmann, René Hempel, Dennis Wegener, et al.. Ac- tive site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme. Biochemical Journal, Portland Press, 2006, 401 (3), pp.659-665. 10.1042/BJ20061239. hal-00478649 HAL Id: hal-00478649 https://hal.archives-ouvertes.fr/hal-00478649 Submitted on 30 Apr 2010 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Biochemical Journal Immediate Publication. Published on 12 Oct 2006 as manuscript BJ20061239 Active site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme Kristin Moreth¶, Daniel Riester¶, Christian Hildmann¶, René Hempel¶, Dennis Wegener¶,§,‡, -

Analyzing the Del Gene Cluster for Gold Biomineralization Across Delftia Spp
Analyzing the del Gene Cluster for Gold Biomineralization across Delftia spp. Rose Krebs ([email protected]), Dr. Carlos Goller Phyla Unassigned Nitrospirae Bacteroidetes Results Introduction Viruses Chloroflexi Planctomycetes A+3 Average Nucleotide Identity Tail Ignavibacteriae A+3 80% Acidobacteria Relative Abundance Relative A+3 Query Genome Reference Genome ANI Proteobacteria Euryarchaeota Actinobacteria 60% Firmicutes Delftia acidovorans NBRC 14950 Delftia acidovorans 2167 99.998 A+3 Figure 4. Analysis of the raw reads shows Proteobacteria is +3 D. tsuruhatensis NBRC 16741 D. lacustris LMG 24775 98.3373 A 40% the primary phylum represented in three of the four samples. Delftia acidovorans was Firmicutes is the primary phylum present in the hydraulic identified as one of the Gold ions are toxic to Delftia spp. ZNC008 D. lacustris LMG 24775 98.1463 fracture fluid sample. main species of biofilms on bacteria, yet D. acidovorans 20% 1) Subsurface gold mine gold nuggets1. tolerates high concentrations D. acidovorans NBRC 14950 D. acidovorans SPH-1 97.5302 of gold ions1. 2) Rice root iron plaque 3) Subsurface gas well sediment D. acidovorans 2167 D. acidovorans SPH-1 97.5098 0% 1 2 3 4 4) Hydraulic fracture fluid D. tsuruhatensis NBRC 16741 D. acidovorans SPH-1 95.5255 Conclusions D. lacustris LMG 24775 D. acidovorans SPH-1 95.391 ● Delftia species are very closely related and could potentially be The del cluster (delA-delP) Burkholderia cenocepacia D. acidovorans SPH-1 76.81 considered the same species since they have average nucleotide Delftibactin forms solid gold produces delftibactin. identities of over 95%. nanoparticles from gold ions. Whole genome sequencing Table 1. -

Fish Bacterial Flora Identification Via Rapid Cellular Fatty Acid Analysis
Fish bacterial flora identification via rapid cellular fatty acid analysis Item Type Thesis Authors Morey, Amit Download date 09/10/2021 08:41:29 Link to Item http://hdl.handle.net/11122/4939 FISH BACTERIAL FLORA IDENTIFICATION VIA RAPID CELLULAR FATTY ACID ANALYSIS By Amit Morey /V RECOMMENDED: $ Advisory Committe/ Chair < r Head, Interdisciplinary iProgram in Seafood Science and Nutrition /-■ x ? APPROVED: Dean, SchooLof Fisheries and Ocfcan Sciences de3n of the Graduate School Date FISH BACTERIAL FLORA IDENTIFICATION VIA RAPID CELLULAR FATTY ACID ANALYSIS A THESIS Presented to the Faculty of the University of Alaska Fairbanks in Partial Fulfillment of the Requirements for the Degree of MASTER OF SCIENCE By Amit Morey, M.F.Sc. Fairbanks, Alaska h r A Q t ■ ^% 0 /v AlA s ((0 August 2007 ^>c0^b Abstract Seafood quality can be assessed by determining the bacterial load and flora composition, although classical taxonomic methods are time-consuming and subjective to interpretation bias. A two-prong approach was used to assess a commercially available microbial identification system: confirmation of known cultures and fish spoilage experiments to isolate unknowns for identification. Bacterial isolates from the Fishery Industrial Technology Center Culture Collection (FITCCC) and the American Type Culture Collection (ATCC) were used to test the identification ability of the Sherlock Microbial Identification System (MIS). Twelve ATCC and 21 FITCCC strains were identified to species with the exception of Pseudomonas fluorescens and P. putida which could not be distinguished by cellular fatty acid analysis. The bacterial flora changes that occurred in iced Alaska pink salmon ( Oncorhynchus gorbuscha) were determined by the rapid method. -

Phenotypic and Genetic Diversity of Pseudomonads
PHENOTYPIC AND GENETIC DIVERSITY OF PSEUDOMONADS ASSOCIATED WITH THE ROOTS OF FIELD-GROWN CANOLA A Thesis Submitted to the College of Graduate Studies and Research In Partial Fulfillment of the Requirements For the Degree of Doctor of Philosophy In the Department of Applied Microbiology and Food Science University of Saskatchewan Saskatoon By Danielle Lynn Marie Hirkala © Copyright Danielle Lynn Marie Hirkala, November 2006. All rights reserved. PERMISSION TO USE In presenting this thesis in partial fulfilment of the requirements for a Postgraduate degree from the University of Saskatchewan, I agree that the Libraries of this University may make it freely available for inspection. I further agree that permission for copying of this thesis in any manner, in whole or in part, for scholarly purposes may be granted by the professor or professors who supervised my thesis work or, in their absence, by the Head of the Department or the Dean of the College in which my thesis work was done. It is understood that any copying or publication or use of this thesis or parts thereof for financial gain shall not be allowed without my written permission. It is also understood that due recognition shall be given to me and to the University of Saskatchewan in any scholarly use which may be made of any material in my thesis. Requests for permission to copy or to make other use of material in this thesis in whole or part should be addressed to: Head of the Department of Applied Microbiology and Food Science University of Saskatchewan Saskatoon, Saskatchewan, S7N 5A8 i ABSTRACT Pseudomonads, particularly the fluorescent pseudomonads, are common rhizosphere bacteria accounting for a significant portion of the culturable rhizosphere bacteria. -

Delftia Rhizosphaerae Sp. Nov. Isolated from the Rhizosphere of Cistus Ladanifer
TAXONOMIC DESCRIPTION Carro et al., Int J Syst Evol Microbiol 2017;67:1957–1960 DOI 10.1099/ijsem.0.001892 Delftia rhizosphaerae sp. nov. isolated from the rhizosphere of Cistus ladanifer Lorena Carro,1† Rebeca Mulas,2 Raquel Pastor-Bueis,2 Daniel Blanco,3 Arsenio Terrón,4 Fernando Gonzalez-Andr es, 2 Alvaro Peix5,6 and Encarna Velazquez 1,6,* Abstract A bacterial strain, designated RA6T, was isolated from the rhizosphere of Cistus ladanifer. Phylogenetic analyses based on 16S rRNA gene sequence placed the isolate into the genus Delftia within a cluster encompassing the type strains of Delftia lacustris, Delftia tsuruhatensis, Delftia acidovorans and Delftia litopenaei, which presented greater than 97 % sequence similarity with respect to strain RA6T. DNA–DNA hybridization studies showed average relatedness ranging from of 11 to 18 % between these species of the genus Delftia and strain RA6T. Catalase and oxidase were positive. Casein was hydrolysed but gelatin and starch were not. Ubiquinone 8 was the major respiratory quinone detected in strain RA6T together with low amounts of ubiquinones 7 and 9. The major fatty acids were those from summed feature 3 (C16 : 1!7c/C16 : 1 !6c) and C16 : 0. The predominant polar lipids were diphosphatidylglycerol, phosphatidylglycerol and phosphatidylethanolamine. Phylogenetic, chemotaxonomic and phenotypic analyses showed that strain RA6T should be considered as a representative of a novel species of genus Delftia, for which the name Delftia rhizosphaerae sp. nov. is proposed. The type strain is RA6T (=LMG 29737T= CECT 9171T). The genus Delftia comprises Gram-stain-negative, non- The strain was grown on nutrient agar (NA; Sigma) for 48 h sporulating, strictly aerobic rods, motile by polar or bipolar at 22 C to check for motility by phase-contrast microscopy flagella.