FUNCTIONAL ANNOTATION of PROTOCADHERIN BETA GENES HYPER- IJCRR Section: Healthcare METHYLATION and THEIR SIGNIFICANCE in Sci
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genomic Correlates of Relationship QTL Involved in Fore- Versus Hind Limb Divergence in Mice
Loyola University Chicago Loyola eCommons Biology: Faculty Publications and Other Works Faculty Publications 2013 Genomic Correlates of Relationship QTL Involved in Fore- Versus Hind Limb Divergence in Mice Mihaela Palicev Gunter P. Wagner James P. Noonan Benedikt Hallgrimsson James M. Cheverud Loyola University Chicago, [email protected] Follow this and additional works at: https://ecommons.luc.edu/biology_facpubs Part of the Biology Commons Recommended Citation Palicev, M, GP Wagner, JP Noonan, B Hallgrimsson, and JM Cheverud. "Genomic Correlates of Relationship QTL Involved in Fore- Versus Hind Limb Divergence in Mice." Genome Biology and Evolution 5(10), 2013. This Article is brought to you for free and open access by the Faculty Publications at Loyola eCommons. It has been accepted for inclusion in Biology: Faculty Publications and Other Works by an authorized administrator of Loyola eCommons. For more information, please contact [email protected]. This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 3.0 License. © Palicev et al., 2013. GBE Genomic Correlates of Relationship QTL Involved in Fore- versus Hind Limb Divergence in Mice Mihaela Pavlicev1,2,*, Gu¨ nter P. Wagner3, James P. Noonan4, Benedikt Hallgrı´msson5,and James M. Cheverud6 1Konrad Lorenz Institute for Evolution and Cognition Research, Altenberg, Austria 2Department of Pediatrics, Cincinnati Children‘s Hospital Medical Center, Cincinnati, Ohio 3Yale Systems Biology Institute and Department of Ecology and Evolutionary Biology, Yale University 4Department of Genetics, Yale University School of Medicine 5Department of Cell Biology and Anatomy, The McCaig Institute for Bone and Joint Health and the Alberta Children’s Hospital Research Institute for Child and Maternal Health, University of Calgary, Calgary, Canada 6Department of Anatomy and Neurobiology, Washington University *Corresponding author: E-mail: [email protected]. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Novel Transcriptional Targets of the SRY-HMG Box Transcription Factor SOX4 Link Its Expression to the Development of Small Cell Lung Cancer
Published OnlineFirst November 14, 2011; DOI: 10.1158/0008-5472.CAN-11-3506 Cancer Molecular and Cellular Pathobiology Research Novel Transcriptional Targets of the SRY-HMG Box Transcription Factor SOX4 Link Its Expression to the Development of Small Cell Lung Cancer Sandra D. Castillo1, Ander Matheu2, Niccolo Mariani1, Julian Carretero3, Fernando Lopez-Rios4, Robin Lovell-Badge2, and Montse Sanchez-Cespedes1 Abstract The HMG box transcription factor SOX4 involved in neuronal development is amplified and overexpressed in a subset of lung cancers, suggesting that it may be a driver oncogene. In this study, we sought to develop this hypothesis including by defining targets of SOX4 that may mediate its involvement in lung cancer. Ablating SOX4 expression in SOX4-amplified lung cancer cells revealed a gene expression signature that included genes involved in neuronal development such as PCDHB, MYB, RBP1, and TEAD2. Direct recruitment of SOX4 to gene promoters was associated with their upregulation upon ectopic overexpression of SOX4. We confirmed upregulation of the SOX4 expression signature in a panel of primary lung tumors, validating their specific response by a comparison using embryonic fibroblasts from Sox4-deficient mice. Interestingly, we found that small cell lung cancer (SCLC), a subtype of lung cancer with neuroendocrine characteristics, was generally characterized by high levels of SOX2, SOX4, and SOX11 along with the SOX4-specific gene expression signature identified. Taken together, our findings identify a functional role for SOX genes in SCLC, particularly for SOX4 and several novel targets defined in this study. Cancer Res; 72(1); 1–11. Ó2011 AACR. Introduction on chromosome 6p22, containing the SOX4 gene, which was expressed at very high levels and was the best candidate for an As with other types of cancer, lung cancer is undergoing a oncogene in the amplicon (2). -

Supplementary Table 1. List of Genes Up-Regulated in Abiraterone-Resistant Vcap Xenograft Samples PIK3IP1 Phosphoinositide-3-Kin
Supplementary Table 1. List of genes up-regulated in abiraterone-resistant VCaP xenograft samples PIK3IP1 phosphoinositide-3-kinase interacting protein 1 TMEM45A transmembrane protein 45A THBS1 thrombospondin 1 C7orf63 chromosome 7 open reading frame 63 OPTN optineurin FAM49A family with sequence similarity 49, member A APOL4 apolipoprotein L, 4 C17orf108|LOC201229 chromosome 17 open reading frame 108 | hypothetical protein LOC201229 SNORD94 small nucleolar RNA, C/D box 94 PCDHB11 protocadherin beta 11 RBM11 RNA binding motif protein 11 C6orf225 chromosome 6 open reading frame 225 KIAA1984|C9orf86|TMEM14 1 KIAA1984 | chromosome 9 open reading frame 86 | transmembrane protein 141 KIAA1107 TLR3 toll-like receptor 3 LPAR6 lysophosphatidic acid receptor 6 KIAA1683 GRB10 growth factor receptor-bound protein 10 TIMP2 TIMP metallopeptidase inhibitor 2 CCDC28A coiled-coil domain containing 28A FBXL2 F-box and leucine-rich repeat protein 2 NOV nephroblastoma overexpressed gene TSPAN31 tetraspanin 31 NR3C2 nuclear receptor subfamily 3, group C, member 2 DYNC2LI1 dynein, cytoplasmic 2, light intermediate chain 1 C15orf51 dynamin 1 pseudogene SAMD13 sterile alpha motif domain containing 13 RASSF6 Ras association (RalGDS/AF-6) domain family member 6 ZNF167 zinc finger protein 167 GATA2 GATA binding protein 2 NUDT7 nudix (nucleoside diphosphate linked moiety X)-type motif 7 DNAJC18 DnaJ (Hsp40) homolog, subfamily C, member 18 SNORA57 small nucleolar RNA, H/ACA box 57 CALCOCO1 calcium binding and coiled-coil domain 1 RLN2 relaxin 2 ING4 inhibitor of -

The Pdx1 Bound Swi/Snf Chromatin Remodeling Complex Regulates Pancreatic Progenitor Cell Proliferation and Mature Islet Β Cell
Page 1 of 125 Diabetes The Pdx1 bound Swi/Snf chromatin remodeling complex regulates pancreatic progenitor cell proliferation and mature islet β cell function Jason M. Spaeth1,2, Jin-Hua Liu1, Daniel Peters3, Min Guo1, Anna B. Osipovich1, Fardin Mohammadi3, Nilotpal Roy4, Anil Bhushan4, Mark A. Magnuson1, Matthias Hebrok4, Christopher V. E. Wright3, Roland Stein1,5 1 Department of Molecular Physiology and Biophysics, Vanderbilt University, Nashville, TN 2 Present address: Department of Pediatrics, Indiana University School of Medicine, Indianapolis, IN 3 Department of Cell and Developmental Biology, Vanderbilt University, Nashville, TN 4 Diabetes Center, Department of Medicine, UCSF, San Francisco, California 5 Corresponding author: [email protected]; (615)322-7026 1 Diabetes Publish Ahead of Print, published online June 14, 2019 Diabetes Page 2 of 125 Abstract Transcription factors positively and/or negatively impact gene expression by recruiting coregulatory factors, which interact through protein-protein binding. Here we demonstrate that mouse pancreas size and islet β cell function are controlled by the ATP-dependent Swi/Snf chromatin remodeling coregulatory complex that physically associates with Pdx1, a diabetes- linked transcription factor essential to pancreatic morphogenesis and adult islet-cell function and maintenance. Early embryonic deletion of just the Swi/Snf Brg1 ATPase subunit reduced multipotent pancreatic progenitor cell proliferation and resulted in pancreas hypoplasia. In contrast, removal of both Swi/Snf ATPase subunits, Brg1 and Brm, was necessary to compromise adult islet β cell activity, which included whole animal glucose intolerance, hyperglycemia and impaired insulin secretion. Notably, lineage-tracing analysis revealed Swi/Snf-deficient β cells lost the ability to produce the mRNAs for insulin and other key metabolic genes without effecting the expression of many essential islet-enriched transcription factors. -

Functional Test of PCDHB11, the Most Human-Specific Neuronal Surface Protein Guilherme Braga De Freitas, Rafaella Araújo Gonçalves and Matthias Gralle*
de Freitas et al. BMC Evolutionary Biology (2016) 16:75 DOI 10.1186/s12862-016-0652-x RESEARCHARTICLE Open Access Functional test of PCDHB11, the most human-specific neuronal surface protein Guilherme Braga de Freitas, Rafaella Araújo Gonçalves and Matthias Gralle* Abstract Background: Brain-expressed proteins that have undergone functional change during human evolution may contribute to human cognitive capacities, and may also leave us vulnerable to specifically human diseases, such as schizophrenia, autism or Alzheimer’s disease. In order to search systematically for those proteins that have changed the most during human evolution and that might contribute to brain function and pathology, all proteins with orthologs in chimpanzee, orangutan and rhesus macaque and annotated as being expressed on the surface of cells in the human central nervous system were ordered by the number of human-specific amino acid differences that are fixed in modern populations. Results: PCDHB11, a beta-protocadherin homologous to murine cell adhesion proteins, stood out with 12 substitutions and maintained its lead after normalizing for protein size and applying weights for amino acid exchange probabilities. Human PCDHB11 was found to cause homophilic cell adhesion, but at lower levels than shown for other clustered protocadherins. Homophilic adhesion caused by a PCDHB11 with reversion of human- specific changes was as low as for modern human PCDHB11; while neither human nor reverted PCDHB11 adhered to controls, they did adhere to each other. A loss of function in PCDHB11 is unlikely because intra-human variability did not increase relative to the other human beta-protocadherins. Conclusions: The brain-expressed protein with the highest number of human-specific substitutions is PCDHB11. -

Thousands of Cpgs Show DNA Methylation Differences in ACPA-Positive Individuals
G C A T T A C G G C A T genes Article Thousands of CpGs Show DNA Methylation Differences in ACPA-Positive Individuals Yixiao Zeng 1,2, Kaiqiong Zhao 2,3, Kathleen Oros Klein 2, Xiaojian Shao 4, Marvin J. Fritzler 5, Marie Hudson 2,6,7, Inés Colmegna 6,8, Tomi Pastinen 9,10, Sasha Bernatsky 6,8 and Celia M. T. Greenwood 1,2,3,9,11,* 1 PhD Program in Quantitative Life Sciences, Interfaculty Studies, McGill University, Montréal, QC H3A 1E3, Canada; [email protected] 2 Lady Davis Institute for Medical Research, Jewish General Hospital, Montréal, QC H3T 1E2, Canada; [email protected] (K.Z.); [email protected] (K.O.K.); [email protected] (M.H.) 3 Department of Epidemiology, Biostatistics and Occupational Health, McGill University, Montréal, QC H3A 1A2, Canada 4 Digital Technologies Research Centre, National Research Council Canada, Ottawa, ON K1A 0R6, Canada; [email protected] 5 Cumming School of Medicine, University of Calgary, Calgary, AB T2N 1N4, Canada; [email protected] 6 Department of Medicine, McGill University, Montréal, QC H4A 3J1, Canada; [email protected] (I.C.); [email protected] (S.B.) 7 Division of Rheumatology, Jewish General Hospital, Montréal, QC H3T 1E2, Canada 8 Division of Rheumatology, McGill University, Montréal, QC H3G 1A4, Canada 9 Department of Human Genetics, McGill University, Montréal, QC H3A 0C7, Canada; [email protected] 10 Center for Pediatric Genomic Medicine, Children’s Mercy, Kansas City, MO 64108, USA 11 Gerald Bronfman Department of Oncology, McGill University, Montréal, QC H4A 3T2, Canada * Correspondence: [email protected] Citation: Zeng, Y.; Zhao, K.; Oros Abstract: High levels of anti-citrullinated protein antibodies (ACPA) are often observed prior to Klein, K.; Shao, X.; Fritzler, M.J.; Hudson, M.; Colmegna, I.; Pastinen, a diagnosis of rheumatoid arthritis (RA). -

Downloaded by Clicking “Download Results”
bioRxiv preprint doi: https://doi.org/10.1101/562199; this version posted February 27, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. EpiAlignment: alignment with both DNA sequence and epigenomic data Jia Lu1, Xiaoyi Cao1 and Sheng Zhong1,* 1 Department of Bioengineering, University of California San Diego, La Jolla, California, 92093, USA * To whom correspondence should be addressed. Tel: 858-246-1118; Fax: 858-244-4543; Email: [email protected] ABSTRACT Comparative epigenomics, by subjecting both epigenome and genome to interspecies comparison, has become a powerful approach to reveal regulatory features of the genome. Thus elucidated regulatory features surpassed the information derived from comparison of genomic sequences alone. Here, we present EpiAlignment, a web-based tool to align genomic regions with both DNA sequence and epigenomic data. EpiAlignment takes DNA sequence and epigenomic profiles derived by ChIP- seq, DNase-seq, or ATAC-seq from two species as input data, and outputs the best semi-global alignments. These alignments are based on EpiAlignment scores, computed by a dynamic programming algorithm that accounts for both sequence alignment and epigenome similarity. For timely response, the EpiAlignment web server automatically initiates up to 140 computing threads depending on the size of user input data. For users’ convenience, we have pre-compiled the comparable human and mouse epigenome datasets in matched cell types and tissues from the Roadmap Epigenomics and ENCODE consortia. -

The Biological Significance of Methylome Differences in Human Papilloma Virus Associated Head and Neck Cancer
ONCOLOGY LETTERS 12: 4949-4956, 2016 The biological significance of methylome differences in human papilloma virus associated head and neck cancer MARIA J. WORSHAM1, KANG MEI CHEN1, INDRANI DATTA2, JOSENA K. STEPHEN1, DHANANJAY CHITALE3, ALEXANDRA GOTHARD4 and GEORGE DIVINE2 Departments of 1Otolaryngology/Head and Neck Research, 2Public Health Sciences and 3Pathology, Henry Ford Hospital, Detroit, MI 48202; 4Department of Biology, Kalamazoo College, Kalamazoo, MI 49006, USA Received May 31, 2016; Accepted August 26, 2016 DOI: 10.3892/ol.2016.5303 Abstract. In recent years, studies have suggested that Introduction promoter methylation in human papilloma virus (HPV) posi- tive head and neck squamous cell carcinoma (HNSCC) has The results from a number of studies involving next‑generation a mechanistic role and has the potential to improve patient sequencing sequence analysis, gene expression microarrays, survival. The present study aimed to replicate key molecular whole‑exome sequencing and genome‑wide methylation findings from previous analyses of the methylomes of HPV studies support the observation that human papilloma virus positive and HPV negative HNSCC in an independent cohort, (HPV) positive squamous head and neck cancer (HNSCC) to assess the reliability of differentially methylated markers in is a distinct entity and therefore exhibits a particular set of HPV-associated tumors. HPV was measured using real-time somatic alterations (1-6). Data from epidemiological and quantitative PCR and the biological significance of methyla- clinical studies have also indicated that HPV positive HNSCC tion differences was assessed by Ingenuity Pathway Analysis is clinically different from HPV negative HNSCC in terms of (IPA). Using an identical experimental design of a 450K patient characteristics, sensitivity to treatment and molecular methylation platform, 7 of the 11 genes were detected to be biology (7,8). -
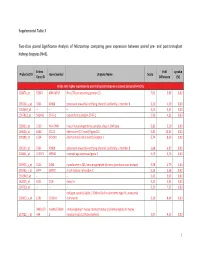
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Prioritizing Candidate Genes Post-GWAS Using Multiple Sources
Cai et al. BMC Genomics (2018) 19:656 https://doi.org/10.1186/s12864-018-5050-x RESEARCHARTICLE Open Access Prioritizing candidate genes post-GWAS using multiple sources of data for mastitis resistance in dairy cattle Zexi Cai* , Bernt Guldbrandtsen, Mogens Sandø Lund and Goutam Sahana Abstract Background: Improving resistance to mastitis, one of the costliest diseases in dairy production, has become an important objective in dairy cattle breeding. However, mastitis resistance is influenced by many genes involved in multiple processes, including the response to infection, inflammation, and post-infection healing. Low genetic heritability, environmental variations, and farm management differences further complicate the identification of links between genetic variants and mastitis resistance. Consequently, studies of the genetics of variation in mastitis resistance in dairy cattle lack agreement about the responsible genes. Results: We associated 15,552,968 imputed whole-genome sequencing markers for 5147 Nordic Holstein cattle with mastitis resistance in a genome-wide association study (GWAS). Next, we augmented P-values for markers in genes in the associated regions using Gene Ontology terms, Kyoto Encyclopedia of Genes and Genomes pathway analysis, and mammalian phenotype database. To confirm results of gene-based analyses, we used gene expression data from E. coli-challenged cow udders. We identified 22 independent quantitative trait loci (QTL) that collectively explained 14% of the variance in breeding values for resistance to clinical mastitis (CM). Using association test statistics with multiple pieces of independent information on gene function and differential expression during bacterial infection, we suggested putative causal genes with biological relevance for 12 QTL affecting resistance to CM in dairy cattle. -

Ncomms7336.Pdf
ARTICLE Received 18 Jul 2014 | Accepted 21 Jan 2015 | Published 19 Mar 2015 DOI: 10.1038/ncomms7336 OPEN Recurrent chromosomal gains and heterogeneous driver mutations characterise papillary renal cancer evolution Michal Kovac1,2,*, Carolina Navas3,*, Stuart Horswell4,*, Max Salm4,*, Chiara Bardella1,*, Andrew Rowan3, Mark Stares3, Francesc Castro-Giner1, Rosalie Fisher3, Elza C. de Bruin5, Monika Kovacova6, Maggie Gorman1, Seiko Makino1, Jennet Williams1, Emma Jaeger1, Angela Jones1, Kimberley Howarth1, James Larkin7, Lisa Pickering7, Martin Gore7, David L. Nicol8,9, Steven Hazell10, Gordon Stamp11, Tim O’Brien12, Ben Challacombe12, Nik Matthews13, Benjamin Phillimore13, Sharmin Begum13, Adam Rabinowitz13, Ignacio Varela14, Ashish Chandra15, Catherine Horsfield15, Alexander Polson15, Maxine Tran16, Rupesh Bhatt17, Luigi Terracciano18, Serenella Eppenberger-Castori18, Andrew Protheroe19, Eamonn Maher20, Mona El Bahrawy21, Stewart Fleming22, Peter Ratcliffe23, Karl Heinimann2, Charles Swanton3,5 & Ian Tomlinson1,24 Papillary renal cell carcinoma (pRCC) is an important subtype of kidney cancer with a problematic pathological classification and highly variable clinical behaviour. Here we sequence the genomes or exomes of 31 pRCCs, and in four tumours, multi-region sequencing is undertaken. We identify BAP1, SETD2, ARID2 and Nrf2 pathway genes (KEAP1, NHE2L2 and CUL3) as probable drivers, together with at least eight other possible drivers. However, only B10% of tumours harbour detectable pathogenic changes in any one driver gene, and where present, the mutations are often predicted to be present within cancer sub-clones. We specifically detect parallel evolution of multiple SETD2 mutations within different sub-regions of the same tumour. By contrast, large copy number gains of chromosomes 7, 12, 16 and 17 are usually early, monoclonal changes in pRCC evolution.