Download (8MB)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Information
doi: 10.1038/nature08795 SUPPLEMENTARY INFORMATION Supplementary Discussion Population naming In some contexts, the indigenous hunter-gatherer and pastoralist peoples of southern Africa are referred to collectively as the Khoisan (Khoi-San) or more recently Khoesan (Khoe-San) people. This grouping is based on the unique linguistic use of click-consonants1. Many names, often country-specific, have been used by Bantu pastoralists and European settlers to describe the hunter-gatherers, including San, Saan, Sonqua, Soaqua, Souqua, Sanqua, Kwankhala, Basarwa, Batwa, Abathwa, Baroa, Bushmen, Bossiesmans, Bosjemans, or Bosquimanos. In addition, group-specific names such as !Kung and Khwe are often used for the broader population. The two most commonly used names, “San” and “Bushmen”, have both been associated with much controversy due to derogatory connotations2. “San” has become the more popular term used in Western literature, although “Bushmen” is arguably the more commonly recognized term within the communities. Since they have no collective name for themselves, the term Bushmen was selected for use in this paper as the term most familiar to the participants themselves. Regarding identification of individuals The five men identified in this study have all elected to have their identity made public knowledge. Thus we present two complete personal genomes (KB1 and ABT), a low-coverage personal genome (NB1), and personal exomes for all five men. On a scientific level, identification allows for current and future correlation of genetic data with demographic and medical histories. On a social level, identification allows for maximizing community benefit. For !Gubi, G/aq’o, D#kgao and !Aî, their name represents not only themselves, but importantly their extended family unit and a way of life severely under threat. -

Supplemental Table 3 Site ID Intron Poly(A) Site Type NM/KG Inum
Supplemental Table 3 Site ID Intron Poly(A) site Type NM/KG Inum Region Gene ID Gene Symbol Gene Annotation Hs.120277.1.10 chr3:170997234:170996860 170996950 b NM_153353 7 CDS 151827 LRRC34 leucine rich repeat containing 34 Hs.134470.1.27 chr17:53059664:53084458 53065543 b NM_138962 10 CDS 124540 MSI2 musashi homolog 2 (Drosophila) Hs.162889.1.18 chr14:80367239:80329208 80366262 b NM_152446 12 CDS 145508 C14orf145 chromosome 14 open reading frame 145 Hs.187898.1.27 chr22:28403623:28415294 28404458 b NM_181832 16 3UTR 4771 NF2 neurofibromin 2 (bilateral acoustic neuroma) Hs.228320.1.6 chr10:115527009:115530350 115527470 b BC036365 5 CDS 79949 C10orf81 chromosome 10 open reading frame 81 Hs.266308.1.2 chr11:117279579:117278191 117278967 b NM_032046 12 CDS 84000 TMPRSS13 transmembrane protease, serine 13 Hs.266308.1.4 chr11:117284536:117281662 117283722 b NM_032046 9 CDS 84000 TMPRSS13 transmembrane protease, serine 13 Hs.2689.1.4 chr10:53492398:53563605 53492622 b NM_006258 7 CDS 5592 PRKG1 protein kinase, cGMP-dependent, type I Hs.280781.1.6 chr18:64715646:64829150 64715837 b NM_024781 4 CDS 79839 C18orf14 chromosome 18 open reading frame 14 Hs.305985.2.25 chr12:8983686:8984438 8983942 b BX640639 17 3UTR NA NA NA Hs.312098.1.36 chr1:151843991:151844258 151844232 b NM_003815 15 CDS 8751 ADAM15 a disintegrin and metalloproteinase domain 15 (metargidin) Hs.314338.1.11 chr21:39490293:39481214 39487623 b NM_018963 41 CDS 54014 BRWD1 bromodomain and WD repeat domain containing 1 Hs.33368.1.3 chr15:92685158:92689361 92688314 b NM_018349 6 CDS 55784 MCTP2 multiple C2-domains with two transmembrane regions 2 Hs.346736.1.21 chr2:99270738:99281614 99272414 b AK126402 10 3UTR 51263 MRPL30 mitochondrial ribosomal protein L30 Hs.445061.1.19 chr16:69322898:69290216 69322712 b NM_018052 14 CDS 55697 VAC14 Vac14 homolog (S. -

Chromosomal Microarray Analysis of Patients with Duane Retraction Syndrome
Int Ophthalmol (2019) 39:2057–2067 https://doi.org/10.1007/s10792-018-1042-8 (0123456789().,-volV)(0123456789().,-volV) ORIGINAL PAPER Chromosomal microarray analysis of patients with Duane retraction syndrome Leyla Niyaz . Sengul Tural . Ozlem Eski Yucel . Ertug˘rul Can . Nursen Ariturk . Zulfinaz B. Celik . Esra Tekcan . Nurten Kara Received: 11 April 2018 / Accepted: 9 November 2018 / Published online: 26 November 2018 Ó Springer Nature B.V. 2018 Abstract Conclusions This study emphasized that CNVs in Purpose Duane retraction syndrome (DS) is a rare several chromosomal regions are known to be congenital strabismus with genetic heterogeneity. The involved in DS. We also underscore the genetic genetic causes of DS are not always of monogenic heterogeneity of DS. Our suggestion is that genes origin; various chromosomal copy number variations located in the most frequently affected regions should (CNVs) have also been reported. The objective of our be focused on in the following candidate gene studies. study was to characterize the CNVs, including gains and losses detected by high-resolution chromosomal Keywords Duane retraction syndrome Á microarray in patients with DS. Chromosomal microarray (CMA) Á Copy number Methods Twenty patients with DS were investigated variation (CNV) using high-resolution chromosomal microarray anal- ysis (CMA) (Affymetrix CytoScan Array 750 K). Conventional cytogenetic analysis was also performed. Introduction Results All samples revealed normal karyotype by cytogenetic analysis. However, in all our patients, Duane syndrome (DS) refers to a spectrum of multiple CNVs, including gains and losses, were congenital ocular motility disorders associated with detected using the high-resolution CMA method. the abnormal innervation of extraocular muscles. -

Partial Gene Duplication and the Formation of Novel Genes
6 Partial Gene Duplication and the Formation of Novel Genes Macarena Toll-Riera1, Steve Laurie1, Núria Radó-Trilla1 and M.Mar Albà1,2 1Evolutionary Genomics Group, Biomedical Informatics Programme, Universitat Pompeu Fabra (UPF) - Institut Municipal d’Investigació Mèdica (IMIM) 2Catalan Institution for Research and Advanced Studies (ICREA), Barcelona Spain 1. Introduction The publication of the first fully sequenced genomes represented a landmark in the biological sciences. The comparison of genomes from different organisms provides us with unprecedented opportunities to address many long-standing evolutionary questions in a more comprehensive way. 1.1 Lineage-specific genes The availability of several genomes from related organisms permits the identification of newly evolved genes in different lineages or species, the study of their mechanisms of formation and the investigation of their role in adapting to new environments or physiological conditions (Domazet-Loso & Tautz, 2003; Guo et al., 2007; Khalturin et al., 2009; Kuo & Kissinger, 2008; Siepel, 2009; Toll-Riera et al., 2009a; Zhou et al., 2008). Recently formed genes give us the opportunity to study the action of natural selection in recent times and to investigate the processes associated with gene creation (Zhou & Wang, 2008). The number of species-specific genes, or orphan genes, is not insignificant. They represent around 14% of the genes in 60 fully sequenced microbial genomes (Siew & Fischer, 2003) and between 20-30% in Drosophila species (Domazet-Loso & Tautz, 2003; Drosophila 12 Genomes Consortium, 2007). Genes restricted to particular lineages include vomeronasal receptors and casein milk proteins in mammals, which are known to be involved in specific physiological adaptations in this lineage (International Chicken Genome Sequencing Consortium, 2004). -

SUPPLEMENTAL METHODS Flow Cytometry Analysis Phycoerythrin
SUPPLEMENTAL METHODS Flow cytometry analysis Phycoerythrin (PE)-conjugated monoclonal antibodies (mAb) to CD3 (clone UCHT1), CD10 (clone ALB1), CD19 (clone J4.119), CD34 (clone 581), CD45 (clone J33), CD54 (clone 84H10), CD86 (clone HA5.2B7), HLA-DR (clone Immu-357), CD105 (clone 1G2), and CD120a (clone H398); Fluorescein IsoThioCyanate (FITC)- conjugated mAb to CD19 (clone J4.119), CD20 (clone B9E9), and CD44 (clone J- 173); and Phycocyanin (PC)7- and PC5-conjugated mAb to CD14 (clone RMO52), CD19 (clone J4.119), and CD20 (clone B9E9) were provided by Beckman Coulter (Villepinte, France). PE-conjugated mAb to CD73 (clone AD2), CD90 (clone 5E10), LTβR (clone hTNF-RP-M12), and CD120b (clone hTNFR-M1) were provided by Becton Dickinson (Le Pont de Claix, France) whereas PE-conjugated mAb anti- CD106 (clone 1.G11B1) was from Chemicon (Temecula, CA). Proliferation assay was performed using the allophycocyanin (APC)-BrdU Flow kit and apoptosis evaluation using PE-conjugated active caspase-3 apoptosis kit (Becton Dickinson). Isotype-matched mAb were used as negative control and, unless otherwise specified, all analyses were performed on TOPRO-3neg viable cells (Invitrogen) using a FACSCalibur (Becton Dickinson) or a Gallios (Beckman Coulter) flow cytometer. Cell-sorting was performed on DAPIneg viable cells (Sigma, St Louis, MO) using a FACSAria cell sorter (Becton Dickinson). Microarray and bioinformatics analysis Scanned Affymetrix raw data were uploaded at https://webpub.chu- rennes.fr/extranetdim/echantadn/acces/editer.php Expression signal values and P- values were obtained for each probeset (PS) using the Partek Genomics Suite software (Partek, St Louis, MO) by the Robust Multichip Averaging algorithm using GC content of probes in normalization (GC-RMA). -

There Are 1393 Genes Significant by SAM with 90Th Percentile Confidence, the False Discovery Rate Among the 1393 Significant Genes Is 0.09968
Class 1 (NPM1 wild-type); Class 2 (NPM1 mutation). There are 1393 genes significant by SAM With 90th percentile confidence, the false discovery rate among the 1393 significant genes is 0.09968 . The delta value used to identify the significant genes is 0.78224 . The fudge factor for standard deviation is computed as 0.06115 . if ratio is >1 gene expression lower in NPM1mut 307 overlapping genes (overlap of genes lower expressed in NPM1mut and putative target genes) if ratio is <1 gene expression higher in NPM1mut rank, based Geom mean of Geom mean of Ratio of on p- ratios in class ratios in class geom Map value 1=wt 2=mut means Clone gene symbol_1 gene symbol_2 Description UG cluster Location 1 1,750 0,395 4,427 IMAGE:382787 TRH TRH TRH || Thyrotropin-releasing hormone || Hs.182231 || AA069596 || || 7200 || 98655 Hs.182231 3 MLLT3 || Myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 || 2 1,689 0,551 3,068 IMAGE:783998 MLLT3 MLLT3 Hs.493585 || AA443284 || || 4300 || 116155 Hs.591085 9 P2RY5 || Purinergic receptor P2Y, G-protein coupled, 5 || Hs.123464 || R91539 || purinergic receptor 3 1,866 0,497 3,754 IMAGE:196488 P2RY5 P2RY5 P2Y5=RB intorn-encoded putative G-protei || 10161 || 112029 Hs.123464 13 4 1,678 0,497 3,375 IMAGE:49923 BAALC BAALC BAALC || Brain and acute leukemia, cytoplasmic || Hs.533446 || H28986 || || 79870 || 311379 Hs.533446 8 5 1,611 0,612 2,634 IMAGE:416374 TSPAN13 TM4SF13 TM4SF13 || Tetraspanin 13 || Hs.364544 || W86201 || || 27075 || 226071 Hs.364544 7 6 1,770 0,557 3,179 -

Table of Contents
Table of Contents Comparison Page Numbers RF vs Static 2-231 HSS vs Static 232-507 LSS vs Static 508-715 RF vs HSS 716-746 RF vs LSS 747-751 HSS vs LSS 752-764 Fold change Regulation ([RF] vs ([RF] vs Probe Name [Static]) [Static]) Common name Gene Symbol Description Genbank Accession Homo sapiens apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3B (APOBEC3B), A_24_P66027 2.741304 down NM_004900 APOBEC3B mRNA [NM_004900] NM_004900 Homo sapiens vasohibin 1 (VASH1), mRNA A_23_P77000 3.932838 down NM_014909 VASH1 [NM_014909] NM_014909 Homo sapiens transmembrane protein 97 A_24_P151920 2.292691 down NM_014573 TMEM97 (TMEM97), mRNA [NM_014573] NM_014573 CN479126 UI-CF-FN0-afv-i-22-0-UI.s1 UI-CF-FN0 Homo sapiens cDNA clone UI-CF-FN0-afv-i-22-0-UI A_32_P149011 3.661964 up CN479126 CN479126 3', mRNA sequence [CN479126] CN479126 Homo sapiens notchless homolog 1 (Drosophila) (NLE1), transcript variant 2, mRNA A_23_P141315 1.785076 down NM_001014445 NLE1 [NM_001014445] NM_001014445 Homo sapiens ribonucleotide reductase M1 A_23_P87351 1.519893 down NM_001033 RRM1 polypeptide (RRM1), mRNA [NM_001033] NM_001033 AT_ssH_PC_3 2.387579 down AT_ssH_PC_3 AT_ssH_PC_3 Homo sapiens huntingtin interacting protein 2 A_23_P155677 1.640733 up NM_005339 HIP2 (HIP2), mRNA [NM_005339] NM_005339 Homo sapiens chromosome 1 open reading frame A_24_P131173 3.527582 down NM_024709 C1orf115 115 (C1orf115), mRNA [NM_024709] NM_024709 Homo sapiens CCR4-NOT transcription complex, A_23_P110846 1.616577 up NM_004779 CNOT8 subunit 8 (CNOT8), mRNA [NM_004779] NM_004779 Homo sapiens protein kinase, AMP-activated, alpha A_23_P60811 3.629454 down NM_006252 PRKAA2 2 catalytic subunit (PRKAA2), mRNA [NM_006252] NM_006252 Homo sapiens pannexin 1 (PANX1), mRNA A_23_P47155 2.378446 up NM_015368 PANX1 [NM_015368] NM_015368 Homo sapiens chemokine (C-X-C motif) ligand 2 A_23_P315364 11.12551 down NM_002089 CXCL2 (CXCL2), mRNA [NM_002089] NM_002089 Centromere protein P (CENP-P). -

Method for a Genome Wide Identification of Expression
(19) TZZ ___T (11) EP 2 177 615 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 21.04.2010 Bulletin 2010/16 C12N 15/11 (2006.01) (21) Application number: 08075816.2 (22) Date of filing: 10.10.2008 (84) Designated Contracting States: • Weltmeier, Fridtjof, Dr. AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 30627 Hannover (DE) HR HU IE IS IT LI LT LU LV MC MT NL NO PL PT RO SE SI SK TR (74) Representative: Baumbach, Friedrich Designated Extension States: Patentanwalt AL BA MK RS Robert-Rössle-Strasse 10 13125 Berlin (DE) (71) Applicant: Fraunhofer-Gesellschaft zur Förderung der Remarks: angewandten Forschung e.V. Claims 16-27,29,31,33-44 and 46-47 are deemed to 80686 München (DE) be abandoned due to non-payment of the claims fees (Rule 45(3) EPC). (72) Inventors: • Borlak, Jürgen, Prof. Dr. 31275 Lehrte OT Immensen (DE) (54) Method for a genome wide identification of expression regulatory sequences and use of genes and molecules derived thereof for the diagnosis and therapy of metabolic and/or tumorous diseases (57) The invention is directed to the use of particular located within the chromosomal position specified by par- human genes, nucleic acids hybridizing to said genes, ticular start and end sites. and gene products encoded thereby in the context of the The invention further relates to a method for a diagnosis and/or therapy of metabolic and/or cancerous genomewide identification of functional binding sites at diseases, preferably of diabetes mellitus and/or colorec- specifically targeted DNA sequences with high resolu- tal cancer, wherein the gene is selected from the group tion, wherein the method comprises, or preferably con- of the human chromosomal genes having at least one sists of, the steps of: expression regulatory sequence according to matrix 1 a. -
12.2% 122000 135M Top 1% 154 4800
CORE Metadata, citation and similar papers at core.ac.uk Provided by IntechOpen We are IntechOpen, the world’s leading publisher of Open Access books Built by scientists, for scientists 4,800 122,000 135M Open access books available International authors and editors Downloads Our authors are among the 154 TOP 1% 12.2% Countries delivered to most cited scientists Contributors from top 500 universities Selection of our books indexed in the Book Citation Index in Web of Science™ Core Collection (BKCI) Interested in publishing with us? Contact [email protected] Numbers displayed above are based on latest data collected. For more information visit www.intechopen.com 6 Partial Gene Duplication and the Formation of Novel Genes Macarena Toll-Riera1, Steve Laurie1, Núria Radó-Trilla1 and M.Mar Albà1,2 1Evolutionary Genomics Group, Biomedical Informatics Programme, Universitat Pompeu Fabra (UPF) - Institut Municipal d’Investigació Mèdica (IMIM) 2Catalan Institution for Research and Advanced Studies (ICREA), Barcelona Spain 1. Introduction The publication of the first fully sequenced genomes represented a landmark in the biological sciences. The comparison of genomes from different organisms provides us with unprecedented opportunities to address many long-standing evolutionary questions in a more comprehensive way. 1.1 Lineage-specific genes The availability of several genomes from related organisms permits the identification of newly evolved genes in different lineages or species, the study of their mechanisms of formation and the investigation of their role in adapting to new environments or physiological conditions (Domazet-Loso & Tautz, 2003; Guo et al., 2007; Khalturin et al., 2009; Kuo & Kissinger, 2008; Siepel, 2009; Toll-Riera et al., 2009a; Zhou et al., 2008). -
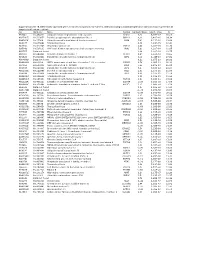
Supplementary File 1B Revised-Förslag
Supplementary file 1B. Differentially expressed genes in cancers compared to non-cancers, ranked according to statistical significance based on features present in all samples in at least one replicate. Acc UGCluster Name Symbol log Fold Changet Adj. P - Value B T57841 Hs.474213 Ubiquitin fusion degradation 1 like (yeast) UFD1L 5,02 5,77E-17 36,61 AA279072 Hs.523875 Inositol polyphosphate phosphatase-like 1 INPPL1 4,29 3,40E-16 34,34 AA454143 Hs.77500 Ubiquitin specific peptidase 4 (proto-oncogene) USP4 5,47 9,11E-16 33,04 N68327 Hs.675604 Transcribed locus 5,16 1,33E-15 32,42 T47230 Hs.522885 Zinc finger protein 32 ZNF32 4,96 3,20E-15 31,39 N95249 Hs.505033 V-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog KRAS 5,90 1,30E-14 29,85 R41787 Data not found 3,94 1,30E-14 29,75 H51262 Hs.658489 Chloride channel CLIC-like 1 CLCC1 5,67 3,80E-14 28,60 N74623 Hs.523414 Insulin-like growth factor 2 (somatomedin A) IGF2 8,04 1,04E-13 27,52 AA001614 Data not found 6,13 1,97E-13 26,81 AA488626 Hs.81424 SMT3 suppressor of mif two 3 homolog 1 (S. cerevisiae) SUMO1 6,74 2,96E-13 26,33 N50854 Hs.534334 Nucleolar protein 1, 120kDa NOL1 7,55 4,20E-13 25,91 N54596 Hs.523414 Insulin-like growth factor 2 (somatomedin A) IGF2 6,47 5,33E-12 23,37 AA129397 Hs.522868 Deleted in azoospermia 1 DAZ1 2,09 2,37E-11 21,85 N54596 Hs.523414 Insulin-like growth factor 2 (somatomedin A) IGF2 6,58 2,37E-11 21,78 AA424871 Hs.236646 Transcribed locus 3,00 2,11E-10 19,59 AA446876 Hs.165950 Fibroblast growth factor receptor 4 FGFR4 1,93 5,48E-10 18,59 AA152347 Hs.485557 Glutathione -

GLRX5 Deficiency Causes Sideroblastic Anemia by Specifically
GLRX5 deficiency causes sideroblastic anemia by specifically impairing heme biosynthesis and depleting cytosolic iron in erythroblasts Hong Ye,1 Suh Young Jeong,1 Manik C. Ghosh,1 Gennadiy Kovtunovych,1 Laura Silvestri,2 Danilo Ortillo,3 Naoya Uchida,4 John Tisdale,4 Clara Camaschella,2 and Tracey A. Rouault1,* Supplemental data Supplemental Figure 1 In situ hybridization of GLRX5 in adult mouse tissues reveals high expression in cerebellum. The hybridization by anti-sense probe indicates the expression of GLRX5 in vivo, whereas hybridization by sense probe is negative control. It shows a higher magnification of the cerebellum, where arrows point to deep cerebellar nuclei. Supplemental Figure 2 GLRX5 has an additional role in erythropoiesis. (A) GLRX5 is highly expressed in erythroblast cells in mouse bone marrow. Mouse bone marrow cells were sorted using Ter119 (for erythroid) and CD11b (for macrophage) coated beads (ref. 50). Ferroportin transcript levels were quantified by qRT-PCR. The order of cell sorting did not affect the results, comparing the two panels. (B) GLRX5 protein levels were upregulated (as shown by western blot) during DMSO-induced differentiation of MEL cells. Samples were collected at 24-72 hours post the induction by 2% DMSO. (C) Hemoglobin level was increased (as shown by Ponceau S staining) after DMSO induction. SOD2 was used as loading control in the western blot. (D) The GLRX5 deficiency could potentially impair the formation of hemoglobin. The GLRX5 oligo-1, which is complementary to the mouse sequence, was used to knockdown mouse GLRX5 in MEL cells. WT, mock and neg were used as negative RNAi controls (as described in Fig. -

The Endothelial Genomic Response to Chronic Hypoxia
CHAPTER 7 The endothelial genomic response to chronic hypoxia Robert E. Verloop1 Anton J. G. Horrevoets2 Marten A. Engelse1 Oscar L. Volger3 Sophie Nadaud4 Pieter Koolwijk1 Victor W.M. van Hinsbergh1 1Laboratory for Physiology and 2Department of Molecular Cell Biology and Immunolo- gy, Institute for Cardiovascular Research, VU University Medical Center, Amsterdam, the Netherland 3Department of Biochemistry, Academic Medical Center, Amsterdam, the Netherlands 4UPMC University Paris 06 and INSERM, UMR_S 525, F-75005, Pa- ris, France Abstract: Exposure of endothelial cells to hypoxia shifts their gene expression pattern to me- tabolically adjust to the change in oxygen tension. Little is known of the effects of chronic hypoxia on these cells as experienced during many pathological conditions, because only short-term hypoxia (up to 24 hours) has been investigated extensi- vely. Therefore, we studied human umbilical vein and microvascular endothelial cells under culture conditions that allowed continuous culture, inspection and medium renewal under defined oxygen tensions. Endothelial cells remained healthy and maintained their proliferation properties at 1% and 5% oxygen atmosphere com- parably or better than at 21% oxygen for up to 2 weeks. Microarray analysis of gene expressions revealed a significant regulation of 435 defined genes, grouped into 8 panels after these culture conditions. Pathway analysis showed hypoxia/HIF- dependent genes specifically regulated at 1% but not 5% oxygen, while the glyco- lysis pathway was well represented in the genes that displayed a gradual increase at lowering (5% and 1%) oxygen concentration. Surprisingly, comparison with five previously published microarray studies on short-term (24 hours) hypoxia-exposed endothelial cells showed substantial overlap in identity of the genes.