The Metabolic Syndrome Modifies the Mrna Expression Profile Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Pentraxin-2 Suppresses C-Jun/AP-1 Signaling to Inhibit Progressive Fibrotic Disease
Pentraxin-2 suppresses c-Jun/AP-1 signaling to inhibit progressive fibrotic disease Naoki Nakagawa, … , Sina A. Gharib, Jeremy S. Duffield JCI Insight. 2016;1(20):e87446. https://doi.org/10.1172/jci.insight.87446. Research Article Immunology Inflammation Pentraxin-2 (PTX-2), also known as serum amyloid P component (SAP/APCS), is a constitutive, antiinflammatory, innate immune plasma protein whose circulating level is decreased in chronic human fibrotic diseases. Here we show that recombinant human PTX-2 (rhPTX-2) retards progression of chronic kidney disease in Col4a3 mutant mice with Alport syndrome, reducing blood markers of kidney failure, enhancing lifespan by 20%, and improving histological signs of disease. Exogenously delivered rhPTX-2 was detected in macrophages but also in tubular epithelial cells, where it counteracted macrophage activation and was cytoprotective for the epithelium. Computational analysis of genes regulated by rhPTX-2 identified the transcriptional regulator c-Jun along with its activator protein–1 (AP-1) binding partners as a central target for the function of rhPTX-2. Accordingly, PTX-2 attenuates c-Jun and AP-1 activity, and reduces expression of AP-1–dependent inflammatory genes in both monocytes and epithelium. Our studies therefore identify rhPTX-2 as a potential therapy for chronic fibrotic disease of the kidney and an important inhibitor of pathological c-Jun signaling in this setting. Find the latest version: https://jci.me/87446/pdf RESEARCH ARTICLE Pentraxin-2 suppresses c-Jun/AP-1 signaling to inhibit progressive fibrotic disease Naoki Nakagawa,1,2,3 Luke Barron,4 Ivan G. Gomez,1,2,4 Bryce G. -
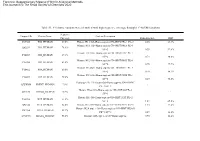
Table S1. 49 Histone Variants Were Identified with High Sequence Coverage Through LC-MS/MS Analysis Electronic Supplementary
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is © The Royal Society of Chemistry 2020 Table S1. 49 histone variants were identified with high sequence coverage through LC-MS/MS analysis Sequence Uniprot IDs Protein Name Protein Description Coverage Ratio E2+/E2- RSD P07305 H10_HUMAN 67.5% Histone H1.0 OS=Homo sapiens GN=H1F0 PE=1 SV=3 4.85 23.3% Histone H1.1 OS=Homo sapiens GN=HIST1H1A PE=1 Q02539 H11_HUMAN 74.4% SV=3 0.35 92.6% Histone H1.2 OS=Homo sapiens GN=HIST1H1C PE=1 P16403 H12_HUMAN 67.1% SV=2 0.73 80.6% Histone H1.3 OS=Homo sapiens GN=HIST1H1D PE=1 P16402 H13_HUMAN 63.8% SV=2 0.75 77.7% Histone H1.4 OS=Homo sapiens GN=HIST1H1E PE=1 P10412 H14_HUMAN 69.0% SV=2 0.70 80.3% Histone H1.5 OS=Homo sapiens GN=HIST1H1B PE=1 P16401 H15_HUMAN 79.6% SV=3 0.29 98.3% Testis-specific H1 histone OS=Homo sapiens GN=H1FNT Q75WM6 H1FNT_HUMAN 7.8% \ \ PE=2 SV=3 Histone H1oo OS=Homo sapiens GN=H1FOO PE=2 Q8IZA3 H1FOO_HUMAN 5.2% \ \ SV=1 Histone H1t OS=Homo sapiens GN=HIST1H1T PE=2 P22492 H1T_HUMAN 31.4% SV=4 1.42 65.0% Q92522 H1X_HUMAN 82.6% Histone H1x OS=Homo sapiens GN=H1FX PE=1 SV=1 1.15 33.2% Histone H2A type 1 OS=Homo sapiens GN=HIST1H2AG P0C0S8 H2A1_HUMAN 99.2% PE=1 SV=2 0.57 26.8% Q96QV6 H2A1A_HUMAN 58.0% Histone H2A type 1-A OS=Homo sapiens 0.90 11.2% GN=HIST1H2AA PE=1 SV=3 Histone H2A type 1-B/E OS=Homo sapiens P04908 H2A1B_HUMAN 99.2% GN=HIST1H2AB PE=1 SV=2 0.92 30.2% Histone H2A type 1-C OS=Homo sapiens Q93077 H2A1C_HUMAN 100.0% GN=HIST1H2AC PE=1 SV=3 0.76 27.6% Histone H2A type 1-D OS=Homo sapiens P20671 -

UNIVERSITY of CALIFORNIA, SAN DIEGO Functional Analysis of Sall4
UNIVERSITY OF CALIFORNIA, SAN DIEGO Functional analysis of Sall4 in modulating embryonic stem cell fate A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Molecular Pathology by Pei Jen A. Lee Committee in charge: Professor Steven Briggs, Chair Professor Geoff Rosenfeld, Co-Chair Professor Alexander Hoffmann Professor Randall Johnson Professor Mark Mercola 2009 Copyright Pei Jen A. Lee, 2009 All rights reserved. The dissertation of Pei Jen A. Lee is approved, and it is acceptable in quality and form for publication on microfilm and electronically: ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ Co-Chair ______________________________________________________________ Chair University of California, San Diego 2009 iii Dedicated to my parents, my brother ,and my husband for their love and support iv Table of Contents Signature Page……………………………………………………………………….…iii Dedication…...…………………………………………………………………………..iv Table of Contents……………………………………………………………………….v List of Figures…………………………………………………………………………...vi List of Tables………………………………………………….………………………...ix Curriculum vitae…………………………………………………………………………x Acknowledgement………………………………………………….……….……..…...xi Abstract………………………………………………………………..…………….....xiii Chapter 1 Introduction ..…………………………………………………………………………….1 Chapter 2 Materials and Methods……………………………………………………………..…12 -

Pentraxin-3 to Better Delineate Necrotizing Soft Tissue Infection: Not Really! Patrick M
Honore and Spapen Critical Care (2016) 20:173 DOI 10.1186/s13054-016-1319-0 LETTER Open Access Pentraxin-3 to better delineate necrotizing soft tissue infection: not really! Patrick M. Honore* and Herbert D. Spapen See related research by Hansen et al., http://ccforum.biomedcentral.com/articles/10.1186/s13054-016-1210-z Necrotizing soft tissue infection (NSTI) is a devastating assessment and persistently elevated levels during evolv- condition with high morbidity and a dismal prognosis. ing sepsis may contribute to tissue damage and predict Timely and adequate surgery and early aggressive treat- poor patient outcome [2]. Second, when looking closely ment of associated sepsis are imperative to improve at the study results, PTX3 levels appeared to be fairly in survival [1–3]. Pentraxin-3 (PTX3) is a glycoprotein line with PCT concentrations for assessing NSTI- released by endothelial and inflammatory cells upon associated morbidity and mortality. PCT also outper- stimulation by cytokines and endotoxins. In contrast with forms CRP in predicting severity of infection and organ C-reactive protein (CRP) which is produced in the liver in dysfunction in sepsis [4] but is more readily available response to systemic inflammation, PTX3 is thought to than PTX3. Finally, many patients with NSTI develop better reflect local vascular inflammation and bacterial acute kidney injury, necessitating renal replacement load [2]. PTX3 assessment might thus be a more appropri- therapy (RRT) (25 % of the patients studied by Hansen ate marker of severity and prognosis of NSTI. This was et al.!). PTX3 has a molecular weight of approximately corroborated by Hansen et al. -

Deubiquitinases in Cancer: New Functions and Therapeutic Options
Oncogene (2012) 31, 2373–2388 & 2012 Macmillan Publishers Limited All rights reserved 0950-9232/12 www.nature.com/onc REVIEW Deubiquitinases in cancer: new functions and therapeutic options JM Fraile1, V Quesada1, D Rodrı´guez, JMP Freije and C Lo´pez-Otı´n Departamento de Bioquı´mica y Biologı´a Molecular, Facultad de Medicina, Instituto Universitario de Oncologı´a, Universidad de Oviedo, Oviedo, Spain Deubiquitinases (DUBs) have fundamental roles in the Hunter, 2010). Consistent with the functional relevance ubiquitin system through their ability to specifically of proteases in these processes, alterations in their deconjugate ubiquitin from targeted proteins. The human structure or in the mechanisms controlling their genome encodes at least 98 DUBs, which can be grouped spatiotemporal expression patterns and activities cause into 6 families, reflecting the need for specificity in diverse pathologies such as arthritis, neurodegenerative their function. The activity of these enzymes affects the alterations, cardiovascular diseases and cancer. Accord- turnover rate, activation, recycling and localization ingly, many proteases are an important focus of of multiple proteins, which in turn is essential for attention for the pharmaceutical industry either as drug cell homeostasis, protein stability and a wide range of targets or as diagnostic and prognostic biomarkers signaling pathways. Consistent with this, altered DUB (Turk, 2006; Drag and Salvesen, 2010). function has been related to several diseases, including The recent availability of the genome sequence cancer. Thus, multiple DUBs have been classified as of different organisms has facilitated the identification oncogenes or tumor suppressors because of their regula- of their entire protease repertoire, which has been tory functions on the activity of other proteins involved in defined as degradome (Lopez-Otin and Overall, 2002). -

PSMD14 Is Over-Expressed in Human Endometrial Cancer
Over-expression of proteasome 26S subunit, non-ATPase 14 in human endometrial cancer. Shahan Mamoor, MS1 [email protected] East Islip, NY USA Gynecologic cancers including cancers of the endometrium are a clinical problem1-4. We mined published microarray data5,6 to discover genes associated with endometrial cancers by comparing transcriptomes of the normal and hyperplastic endometrium to endometrial tumors from humans. We identified proteasome 26S subunit, non-ATPase 14, encoded by PSMD14, as among the most differentially expressed genes, transcriptome-wide, in cancers of the endometrium. PSMD14 was expressed at significantly higher levels in endometrial tumor tissues as compared to the endometrium. Importantly, in human endometrial cancer, primary tumor expression of PSMD14 was correlated with overall survival in white patients with low mutational burden. PSMD14 may be a molecule of interest in understanding the etiology or progression of human endometrial cancer. Keywords: endometrial cancer, gynecologic cancers, endometrium, PSMD14, proteasome 26S subunit, non-ATPase 14, systems biology of endometrial cancer, targeted therapeutics in endometrial cancer. 1 Endometrial cancer is the most common gynecologic cancer in the developed world1. Over the last three decades, the incidence of endometrial cancer has increased 21%4 and the death rate has increased 100%3. We harnessed the power of independently published microarray datasets5,6 to determine in an unbiased fashion and at the systems-level genes most differentially expressed in endometrial tumors. We report here the differential and increased expression of the proteasome 26S subunit, non-ATPase 14 (PSMD14) in human endometrial cancer. Methods We utilized datasets GSE636785 and GSE1061916 for this global differential gene expression analysis of human endometrial cancer in conjunction with GEO2R. -

Genome-Wide Screen of Cell-Cycle Regulators in Normal and Tumor Cells
bioRxiv preprint doi: https://doi.org/10.1101/060350; this version posted June 23, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Genome-wide screen of cell-cycle regulators in normal and tumor cells identifies a differential response to nucleosome depletion Maria Sokolova1, Mikko Turunen1, Oliver Mortusewicz3, Teemu Kivioja1, Patrick Herr3, Anna Vähärautio1, Mikael Björklund1, Minna Taipale2, Thomas Helleday3 and Jussi Taipale1,2,* 1Genome-Scale Biology Program, P.O. Box 63, FI-00014 University of Helsinki, Finland. 2Science for Life laboratory, Department of Biosciences and Nutrition, Karolinska Institutet, SE- 141 83 Stockholm, Sweden. 3Science for Life laboratory, Division of Translational Medicine and Chemical Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 21 Stockholm, Sweden To identify cell cycle regulators that enable cancer cells to replicate DNA and divide in an unrestricted manner, we performed a parallel genome-wide RNAi screen in normal and cancer cell lines. In addition to many shared regulators, we found that tumor and normal cells are differentially sensitive to loss of the histone genes transcriptional regulator CASP8AP2. In cancer cells, loss of CASP8AP2 leads to a failure to synthesize sufficient amount of histones in the S-phase of the cell cycle, resulting in slowing of individual replication forks. Despite this, DNA replication fails to arrest, and tumor cells progress in an elongated S-phase that lasts several days, finally resulting in death of most of the affected cells. -

The Proteasomal Deubiquitinating Enzyme PSMD14 Regulates 2 Macroautophagy by Controlling Golgi-To-ER Retrograde Transport
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.29.925503; this version posted January 31, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 The proteasomal deubiquitinating enzyme PSMD14 regulates 2 macroautophagy by controlling Golgi-to-ER retrograde transport 3 Bustamante HA1,2, Cereceda K3, González AE1,4, Valenzuela GE5,6, 4 Cheuquemilla Y6, Hernández S3, Arias-Muñoz E3, Cerda-Troncoso C3, 5 Bandau S8, Soza A3, Kausel G5, Kerr B3, Mardones GA1,7, Cancino J3, Hay 6 RT8, Rojas-Fernandez A6,8¥, Burgos PV3,9¥ 7 1Instituto de Fisiología, Facultad de Medicina, Universidad Austral de Chile, 5110566, Valdivia, Chile 8 2Instituto de Microbiología Clínica, Facultad de Medicina, Universidad Austral de Chile, 5110566, Valdivia, 9 Chile 10 3Centro de Biología Celular y Biomedicina (CEBICEM), Facultad de Medicina y Ciencia, Universidad San 11 Sebastián, Lota 2465, 7510157, Santiago, Chile 12 4Institute of Biochemistry II, School of Medicine, Goethe University Frankfurt, Theoder-Stern-Kai 7, 60590, 13 Frankfurt am Main, Germany 14 5Instituto de Bioquímica y Microbiología, Facultad de Ciencias, Universidad Austral de Chile, 5110566, 15 Valdivia, Chile. 16 6Instituto de Medicina & Centro Interdisciplinario de Estudios del Sistema Nervioso (CISNe), Universidad 17 Austral de Chile, 5110566, Valdivia, Chile. 18 7Centro Interdisciplinario de Estudios del Sistema Nervioso (CISNe), Universidad Austral de Chile, 5110566, 19 Valdivia, Chile. 20 8Centre for Gene Regulation and Expression, College of Life Sciences, University of Dundee, DD1 4HN, 21 Dundee, United Kingdom. -

Histone-Related Genes Are Hypermethylated in Lung Cancer
Published OnlineFirst October 1, 2019; DOI: 10.1158/0008-5472.CAN-19-1019 Cancer Genome and Epigenome Research Histone-Related Genes Are Hypermethylated in Lung Cancer and Hypermethylated HIST1H4F Could Serve as a Pan-Cancer Biomarker Shihua Dong1,Wei Li1, Lin Wang2, Jie Hu3,Yuanlin Song3, Baolong Zhang1, Xiaoguang Ren1, Shimeng Ji3, Jin Li1, Peng Xu1, Ying Liang1, Gang Chen4, Jia-Tao Lou2, and Wenqiang Yu1 Abstract Lung cancer is the leading cause of cancer-related deaths lated in all 17 tumor types from TCGA datasets (n ¼ 7,344), worldwide. Cytologic examination is the current "gold stan- which was further validated in nine different types of cancer dard" for lung cancer diagnosis, however, this has low sensi- (n ¼ 243). These results demonstrate that HIST1H4F can tivity. Here, we identified a typical methylation signature of function as a universal-cancer-only methylation (UCOM) histone genes in lung cancer by whole-genome DNA methyl- marker, which may aid in understanding general tumorigen- ation analysis, which was validated by The Cancer Genome esis and improve screening for early cancer diagnosis. Atlas (TCGA) lung cancer cohort (n ¼ 907) and was further confirmed in 265 bronchoalveolar lavage fluid samples with Significance: These findings identify a new biomarker for specificity and sensitivity of 96.7% and 87.0%, respectively. cancer detection and show that hypermethylation of histone- More importantly, HIST1H4F was universally hypermethy- related genes seems to persist across cancers. Introduction to its low specificity, LDCT is far from satisfactory as a screening tool for clinical application, similar to other currently used cancer Lung cancer is one of the most common malignant tumors and biomarkers, such as carcinoembryonic antigen (CEA), neuron- the leading cause of cancer-related deaths worldwide (1, 2). -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -

Comparative Transcriptomics Identifies Potential Stemness-Related Markers for Mesenchymal Stromal/Stem Cells
bioRxiv preprint doi: https://doi.org/10.1101/2021.05.25.445659; this version posted May 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Comparative Transcriptomics Identifies Potential Stemness-Related Markers for Mesenchymal Stromal/Stem Cells Authors Myret Ghabriel 1, Ahmed El Hosseiny 1, 2, Ahmed Moustafa*1, 2 and Asma Amleh*1, 2 Affiliations 1Biotechnology Program, American University in Cairo, New Cairo 11835, Egypt 2Department of Biology, American University in Cairo, New Cairo 11835, Egypt *Corresponding authors: Ahmed Moustafa [email protected] Asma Amleh [email protected]. Abstract Mesenchymal stromal/stem cells (MSCs) are multipotent cells residing in multiple tissues with the capacity for self-renewal and differentiation into various cell types. These properties make them promising candidates for regenerative therapies. MSC identification is critical in yielding pure populations for successful therapeutic applications; however, the criteria for MSC identification proposed by the International Society for Cellular Therapy (ISCT) is inconsistent across different tissue sources. In this study, we aimed to identify potential markers to be used together with the ISCT’s criteria to provide a more accurate means of MSC identification. Thus, we carried out a comparative analysis of the expression of human and mouse MSCs derived from multiple tissues to identify the common differentially expressed genes. We show that six members of the proteasome degradation system are similarly expressed across MSCs derived from bone marrow, adipose tissue, amnion, and umbilical cord. -

Theranostics the PSMD14 Inhibitor Thiolutin As a Novel Therapeutic
Theranostics 2021, Vol. 11, Issue 12 5847 Ivyspring International Publisher Theranostics 2021; 11(12): 5847-5862. doi: 10.7150/thno.46109 Research Paper The PSMD14 inhibitor Thiolutin as a novel therapeutic approach for esophageal squamous cell carcinoma through facilitating SNAIL degradation Chao Jing1*, Xingchen Li1*, Mengqian Zhou1*, Shengchi Zhang1,2*, Qingchuan Lai1, Dandan Liu1, Beibei Ye1, Linqi Li1, Yue Wu1, Hong Li1, Kai Yue1, Peng Chen1, Xiaofeng Yao1, Yansheng Wu1, Yuansheng Duan1, Xudong Wang1 1. Department of Maxillofacial and Otorhinolaryngological Oncology, Tianjin Medical University Cancer Institute and Hospital, Key Laboratory of Cancer Prevention and Therapy, Tianjin Cancer Institute, National Clinical Research Center of Cancer, Tianjin 300060, China. 2. Department of Ear, Nose and Throat, Tianjin 1st Centre Hospital, Tianjin 300192, China. * These authors contributed equally to this work. Corresponding authors: Dr. Xudong Wang ([email protected]), Ms Yuansheng Duan ([email protected]) and Dr. Yansheng Wu ([email protected]). Department of Maxillofacial and Otorhinolaryngological Oncology, Tianjin Medical University Cancer Institute and Hospital, Key Laboratory of Cancer Prevention and Therapy, Tianjin Cancer Institute, National Clinical Research Center of Cancer, 18 Huanhuxi Road, Tianjin 300060, China. Tel: +86-22-23340123. © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2020.03.17; Accepted: 2021.03.11; Published: 2021.04.03 Abstract Metastasis and chemoresistance are major causes of poor prognosis in patients with esophageal squamous cell carcinoma (ESCC), manipulated by multiple factors including deubiquitinating enzyme (DUB).