Whole Exome and Transcriptome Analyses Integrated With
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’S Biological Network Based on Systematic Pharmacology
ORIGINAL RESEARCH published: 25 June 2021 doi: 10.3389/fphar.2021.601846 Exploring the Regulatory Mechanism of Hedysarum Multijugum Maxim.-Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’s Biological Network Based on Systematic Pharmacology Kailin Yang 1†, Liuting Zeng 1†, Anqi Ge 2†, Yi Chen 1†, Shanshan Wang 1†, Xiaofei Zhu 1,3† and Jinwen Ge 1,4* Edited by: 1 Takashi Sato, Key Laboratory of Hunan Province for Integrated Traditional Chinese and Western Medicine on Prevention and Treatment of 2 Tokyo University of Pharmacy and Life Cardio-Cerebral Diseases, Hunan University of Chinese Medicine, Changsha, China, Galactophore Department, The First 3 Sciences, Japan Hospital of Hunan University of Chinese Medicine, Changsha, China, School of Graduate, Central South University, Changsha, China, 4Shaoyang University, Shaoyang, China Reviewed by: Hui Zhao, Capital Medical University, China Background: Clinical research found that Hedysarum Multijugum Maxim.-Chuanxiong Maria Luisa Del Moral, fi University of Jaén, Spain Rhizoma Compound (HCC) has de nite curative effect on cerebral ischemic diseases, *Correspondence: such as ischemic stroke and cerebral ischemia-reperfusion injury (CIR). However, its Jinwen Ge mechanism for treating cerebral ischemia is still not fully explained. [email protected] †These authors share first authorship Methods: The traditional Chinese medicine related database were utilized to obtain the components of HCC. The Pharmmapper were used to predict HCC’s potential targets. Specialty section: The CIR genes were obtained from Genecards and OMIM and the protein-protein This article was submitted to interaction (PPI) data of HCC’s targets and IS genes were obtained from String Ethnopharmacology, a section of the journal database. -

Supreme Court of the United States
No. 12-398 IN THE Supreme Court of the United States THE ASSOCIATION FOR MOLECULAR PATHOLOGY, ET AL., Petitioners, v. MYRIAD GENETICS, INC., ET AL., Respondents. __________ On Writ of Certiorari to the United States Court of Appeals for the Federal Circuit __________ BRIEF OF AMICI CURIAE AMERICAN MEDICAL ASSOCIATION, AMERICAN SOCIETY OF HUMAN GENETICS, AMERICAN COLLEGE OF OBSTETRICIANS AND GYNECOLOGISTS, AMERICAN OSTEOPATHIC ASSOCIATION, AMERICAN COLLEGE OF LEGAL MEDICINE, AND THE MEDICAL SOCIETY OF THE STATE OF NEW YORK IN SUPPORT OF PETITIONERS __________ Professor Lori B. Andrews Counsel of Record Chicago-Kent College of Law Illinois Institute of Technology 565 West Adams Street Chicago, IL 60661 [email protected] 312-906-5359 TABLE OF CONTENTS TABLE OF AUTHORITIES ......................................... iii STATEMENT OF INTEREST OF AMICI CURIAE .... 1 SUMMARY OF THE ARGUMENT .............................. 4 ARGUMENT .................................................................. 6 I. PHYSICIANS’ AND RESEARCHERS’ ACCESS TO HUMAN GENE SEQUENCES IS VITAL TO HEALTH CARE AND RESEARCH ........................................................ 6 A. Patents on Human Genes Interfere with Diagnosis and Treatment of Patients ..... 7 B. Patents on Human Genes Increase the Cost of Genetic Testing .......................... 11 C. Patents on Human Genes Impede Innovation .............................................. 13 D. Existing Non-Patent Incentives Are Sufficient to Encourage Innovation in Genetics ................................................. -

Motifs in Saccharomyces Cerevisiae Master's Thesis
UNIVERSITY OF TARTU Faculty of Biology and Geography Institute of Molecular and Cell Biology Hedi Peterson The discovery and characterization of regulatory motifs in Saccharomyces cerevisiae Master's Thesis Supervisor: Jaak Vilo, PhD Tartu 2006 "Computers are useless. They can only give you answers." Pablo Picasso Contents 1 Introduction 1 1.1 Background . 1 1.2 Objective . 3 2 Background 4 2.1 Biological background . 4 2.1.1 Gene Ontology . 5 2.1.2 Regulatory motifs . 8 2.1.3 Protein-protein interactions . 8 2.2 Bioinformatics approaches . 10 2.2.1 Pattern Discovery . 10 2.2.2 Phylogenetic footprinting . 10 3 Material and methods 12 3.1 Data . 12 3.1.1 Sequences . 13 3.1.2 Previously known transcription factors and regulatory motifs . 15 3.1.3 GO groups . 16 3.1.4 Protein-protein interaction data . 17 3.2 Pattern discovery . 18 3.3 Statistical evaluation . 19 3.4 Expansion of groups by PPI data . 20 4 Results 23 4.1 Randomization . 23 4.2 Pattern discovery for known transcription factors' target groups 26 4.3 Pattern discovery in GO dataset . 27 4.4 Expansion of GO groups by PPI . 31 4.5 From protein-protein interactions to regulatory motifs and GO annotations . 31 4.5.1 Pipeline example . 32 4.6 Web-tool Gviz-PPI . 35 4.7 Data update to BiGeR . 36 5 Discussion 40 6 Conclusions 43 Summary 44 Summary in Estonian 46 Acknowledgements 48 References 49 Appendix 55 List of Figures 2.1 Example of a GO molecular function subtree. Here GO:0003887 is a leaf and GO:0003674 is a root node. -

Jbrowse: a Next-Generation Genome Browser
Downloaded from genome.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press Resource JBrowse: A next-generation genome browser Mitchell E. Skinner,1 Andrew V. Uzilov,1 Lincoln D. Stein,2 Christopher J. Mungall,3 and Ian H. Holmes1,3,4 1Department of Bioengineering, University of California at Berkeley, Berkeley, California 94720, USA; 2Ontario Institute for Cancer Research, Toronto, Ontario M5G 0A3, Canada; 3Lawrence Berkeley National Laboratory, Berkeley, California 94720, USA We describe an open source, portable, JavaScript-based genome browser, JBrowse, that can be used to navigate genome annotations over the web. JBrowse helps preserve the user’s sense of location by avoiding discontinuous transitions, instead offering smoothly animated panning, zooming, navigation, and track selection. Unlike most existing genome browsers, where the genome is rendered into images on the webserver and the role of the client is restricted to displaying those images, JBrowse distributes work between the server and client and therefore uses significantly less server overhead than previous genome browsers. We report benchmark results empirically comparing server- and client-side rendering strategies, review the architecture and design considerations of JBrowse, and describe a simple wiki plug-in that allows users to upload and share annotation tracks. [The JBrowse source code (freely licensed), live demonstrations, mailing list, documentation, bug-tracking, and virtual machine images are available at http://jbrowse.org/.] In a genome, spatial relationships often indicate functional rela- the process repeats itself. This use of CGI imposes a page-based tionships. A genome browser (Stein et al. 2002; Kent et al. 2003; model of viewing the data; that is, every action (such as moving to Stalker et al. -

Cancer Informatics 2014:13 13–20 Doi: 10.4137/CIN.S13495
Open Access: Full open access to Cancer this and thousands of other papers at http://www.la-press.com. Informatics OmicCircos: A Simple-to-Use R Package for the Circular Visualization of Multidimensional Omics Data Ying Hu, Chunhua Yan, Chih-Hao Hsu, Qing-Rong Chen, Kelvin Niu, George A. Komatsoulis and Daoud Meerzaman Center for Biomedical Informatics and Information Technology, National Cancer Institute, Rockville, MD 20850, USA. ABSTRACT SUMMarY: OmicCircos is an R software package used to generate high-quality circular plots for visualizing genomic variations, including mutation patterns, copy number variations (CNVs), expression patterns, and methylation patterns. Such variations can be displayed as scatterplot, line, or text-label figures. Relationships among genomic features in different chromosome positions can be represented in the forms of polygons or curves. Utilizing the statistical and graphic functions in an R/Bioconductor environment, OmicCircos performs statistical analyses and displays results using cluster, boxplot, histogram, and heatmap formats. In addition, OmicCircos offers a number of unique capabilities, including independent track drawing for easy modifica- tion and integration, zoom functions, link-polygons, and position-independent heatmaps supporting detailed visualization. AVaiLabiLitY AND IMPLEMENtatiON: OmicCircos is available through Bioconductor at http://www.bioconductor.org/packages/devel/bioc/ html/OmicCircos.html. An extensive vignette in the package describes installation, data formatting, and workflow procedures. The software is open source under the Artistic−2.0 license. KEYWORDS: R package, circular plot, genomic variation CITATION: Hu et al. OmicCircos: A Simple-to-Use R Package for the Circular Visualization of Multidimensional Omics Data. Cancer Informatics 2014:13 13–20 doi: 10.4137/CIN.S13495. -

Single-Cell Transcriptome Analysis Reveals Gene Signatures Associated with T-Cell Persistence
Author Manuscript Published OnlineFirst on September 4, 2019; DOI: 10.1158/2326-6066.CIR-19-0299 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Single-cell transcriptome analysis reveals gene signatures associated with T-cell persistence following adoptive cell therapy Yong-Chen Lu1, Li Jia1, Zhili Zheng1, Eric Tran1,2, Paul F. Robbins1, Steven A. Rosenberg1 1Surgery Branch, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA. 2Current address: Earle A. Chiles Research Institute, Providence Cancer Institute, Portland, OR 97213, USA Corresponding author: Yong-Chen Lu, Surgery Branch, National Cancer Institute, National Institutes of Health, Building 10-CRC, Rm 3-5930, 10 Center Dr, Bethesda, MD, 20892. Phone: 240-858-3818, E-mail: [email protected] Corresponding author: Steven A. Rosenberg, Surgery Branch, National Cancer Institute, National Institutes of Health, Building 10-CRC, Rm 3-3940, 10 Center Dr, Bethesda, MD, 20892. Phone: 240-858-3080, E-mail: [email protected] Running title: Gene signatures associated with T-cell persistence Financial Support: This work was supported by the Intramural Research Program of National Cancer Institute. Disclosure of Potential Conflicts of Interest: No potential conflicts of interest were disclosed. Keywords: single-cell, adoptive cell therapy, persistence, tumor-infiltrating lymphocytes, cancer immunotherapy 1 Downloaded from cancerimmunolres.aacrjournals.org on September 27, 2021. © 2019 American Association for Cancer Research. Author Manuscript Published OnlineFirst on September 4, 2019; DOI: 10.1158/2326-6066.CIR-19-0299 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

Blood Biomarkers for Memory: Toward Early Detection of Risk for Alzheimer Disease, Pharmacogenomics, and Repurposed Drugs
Molecular Psychiatry (2020) 25:1651–1672 https://doi.org/10.1038/s41380-019-0602-2 IMMEDIATE COMMUNICATION Blood biomarkers for memory: toward early detection of risk for Alzheimer disease, pharmacogenomics, and repurposed drugs 1,2,3 1 1 1,3,4 1 1 1 3 A. B. Niculescu ● H. Le-Niculescu ● K. Roseberry ● S. Wang ● J. Hart ● A. Kaur ● H. Robertson ● T. Jones ● 3 3,5 5 2 1 1,4 4 A. Strasburger ● A. Williams ● S. M. Kurian ● B. Lamb ● A. Shekhar ● D. K. Lahiri ● A. J. Saykin Received: 25 March 2019 / Revised: 25 September 2019 / Accepted: 11 November 2019 / Published online: 2 December 2019 © The Author(s) 2019. This article is published with open access Abstract Short-term memory dysfunction is a key early feature of Alzheimer’s disease (AD). Psychiatric patients may be at higher risk for memory dysfunction and subsequent AD due to the negative effects of stress and depression on the brain. We carried out longitudinal within-subject studies in male and female psychiatric patients to discover blood gene expression biomarkers that track short term memory as measured by the retention measure in the Hopkins Verbal Learning Test. These biomarkers were subsequently prioritized with a convergent functional genomics approach using previous evidence in the field implicating them in AD. The top candidate biomarkers were then tested in an independent cohort for ability to predict state short-term memory, 1234567890();,: 1234567890();,: and trait future positive neuropsychological testing for cognitive impairment. The best overall evidence was for a series of new, as well as some previously known genes, which are now newly shown to have functional evidence in humans as blood biomarkers: RAB7A, NPC2, TGFB1, GAP43, ARSB, PER1, GUSB, and MAPT. -

Fine Mapping of the Circling (<I>Cir</I>) Gene on the Distal
Comparative Medicine Vol 53, No 6 Copyright 2003 December 2003 by the American Association for Laboratory Animal Science Pages 642-648 Fine Mapping of the Circling (cir) Gene on the Distal Portion of Mouse Chromosome 9 Kyoung In Cho, DVM,1, 2,* Jeong Woong Lee, PhD,1,* Kil Soo Kim, PhD,3 Eun Ju Lee,1 Jun-Gyo Suh, PhD,2 Ho-Joon Lee, PhD,4 Hyun Taek Kim, PhD,5 Sung Hwa Hong, MD,6 Won Ho Chung, MD,6 Kyu Tae Chang, PhD,7 Byung Hwa Hyun, PhD,7 Yang-Seok Oh, PhD,2,† and Zae Young Ryoo, PhD,1,† Circling mice manifest profound deafness, head-tossing, and bi-directional circling behavior, which they inherit in autosomal recessive manner. Histologic examination of the inner ear reveals abnormalities of the region around the organ of Corti, spiral ganglion neurons, and outer hair cells. A genetic linkage map was constructed for an intraspecific backcross between cir and C57BL/6J mice. The cir gene was mapped to a region between D9Mit116/ D9Mit15 and D9Mit38 on mouse chromosome (Chr) 9. Estimated distances between cir and D9Mit116, and between cir and D9Mit38 were 0.70 ± 0.40 and 0.23 ± 0.23 cM, respectively. Order of the markers was defined as follows: centromere - D9Mit182 - D9Mit51/D9Mit79/ D9Mit310 - D9Mit212/D9Mit184 - D9Mit116/D9Mit15 - cir - D9Mit38 - D9Mit20 - D9Mit243 - D9Mit16 - D9Mit55/D9Mit125 - D9Mit281. On the basis of genetic mapping, we constructed a yeast artifi- cial chromosome (YAC) contig across the cir region. The cir gene is located between the lactotransferrin (ltf) and microtubule-associated protein (map4) genes. -

Genetic Technologies for Building a Better Mouse
Downloaded from genesdev.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW A most formidable arsenal: genetic technologies for building a better mouse James F. Clark,1 Colin J. Dinsmore,1 and Philippe Soriano Department of Cell, Developmental, and Regenerative Biology, Icahn School of Medicine at Mt. Sinai, New York, New York 10029, USA The mouse is one of the most widely used model organ- scribing Mendelian inheritance for mouse coat color char- isms for genetic study. The tools available to alter the acteristics (Cuénot 1902). Mendel himself had started mouse genome have developed over the preceding decades down this same road fifty years earlier, only for his bishop from forward screens to gene targeting in stem cells to the to snuff out the nascent breeding program because a monk recent influx of CRISPR approaches. In this review, we had no business, frankly, breeding (Paigen 2003). first consider the history of mice in genetic study, the Following Cuénot’s papers (Cuénot 1902), genetic re- development of classic approaches to genome modifica- search using the mouse began in earnest, often through tion, and how such approaches have been used and im- the study of spontaneous mouse mutants. One line of re- proved in recent years. We then turn to the recent surge search further explored the complicated genetics of pig- of nuclease-mediated techniques and how they are chang- mentation and the sometimes-surprising coincident ing the field of mouse genetics. Finally, we survey com- phenotypes. The Dominant white spotting allele W (an al- mon classes of alleles used in mice and discuss how lele of the receptor tyrosine kinase Kit) was found to affect they might be engineered using different methods. -

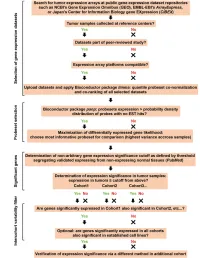
Bioinformatics Analysis: the Regulatory Network of Hsa Circ 0007843 and Hsa Circ 0007331 in Colon Cancer
Hindawi BioMed Research International Volume 2021, Article ID 6662897, 9 pages https://doi.org/10.1155/2021/6662897 Research Article Bioinformatics Analysis: The Regulatory Network of hsa_circ_ 0007843 and hsa_circ_0007331 in Colon Cancer Zeping Han ,1 Huafang Chen ,2 Zhonghui Guo,1 Jianxia Zhu,1 Xingyi Xie,1 Yuguang Li ,1 and Jinhua He 1 1Department of Laboratory Medicine, Central Hospital of Panyu District, Guangzhou, Guangdong 511400, China 2Leizhou Center for Disease Control and Prevention, Leizhou, Guangdong 524200, China Correspondence should be addressed to Yuguang Li; [email protected] and Jinhua He; [email protected] Received 5 October 2020; Revised 8 June 2021; Accepted 5 July 2021; Published 23 July 2021 Academic Editor: Federico Zambelli Copyright © 2021 Zeping Han et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Objective. To analyze the molecular regulation network of circular RNA (circRNA) in colon cancer (CC) by bioinformatics method. Methods. hsa_circ_0007843 and hsa_circ_0007331 proved to be associated with CC in previous studies were chosen as the research object. ConSite database was used to predict the transcription factors associated with circRNA, and the CC-associated transcription factors were screened out after intersection. The CircInteractome database was used to predict the RNA-binding proteins (RBPs) interacting with circRNAs and screen out the CC-associated RBPs after an intersection. Furthermore, the CircInteractome database was used to predict the miRNAs interrelated with circRNAs, and the HMDD v3.2 database was used to search for miRNAs associated with CC. -
Supplementary Data
SUPPLEMENTAL INFORMATION A study restricted to chemokine receptors as well as a genome-wide transcript analysis uncovered CXCR4 as preferentially expressed in Ewing's sarcoma (Ewing's sarcoma) cells of metastatic origin (Figure 4). Transcriptome analyses showed that in addition to CXCR4, genes known to support cell motility and invasion topped the list of genes preferentially expressed in metastasis-derived cells (Figure 4D). These included kynurenine 3-monooxygenase (KMO), galectin-1 (LGALS1), gastrin-releasing peptide (GRP), procollagen C-endopeptidase enhancer (PCOLCE), and ephrin receptor B (EPHB3). KMO, a key enzyme of tryptophan catabolism, has not been linked to metastasis. Tryptophan and its catabolites, however, are involved in immune evasion by tumors, a process that can assist in tumor progression and metastasis (1). LGALS1, GRP, PCOLCE and EPHB3 have been linked to tumor progression and metastasis of several cancers (2-4). Top genes preferentially expressed in L-EDCL included genes that suppress cell motility and/or potentiate cell adhesion such as plakophilin 1 (PKP1), neuropeptide Y (NPY), or the metastasis suppressor TXNIP (5-7) (Figure 4D). Overall, L-EDCL were enriched in gene sets geared at optimizing nutrient transport and usage (Figure 4D; Supplementary Table 3), a state that may support the early stages of tumor growth. Once tumor growth outpaces nutrient and oxygen supplies, gene expression programs are usually switched to hypoxic response and neoangiogenesis, which ultimately lead to tumor egress and metastasis. Accordingly, gene sets involved in extracellular matrix remodeling, MAPK signaling, and response to hypoxia were up-regulated in M-EDCL (Figure 4D; Supplementary Table 4), consistent with their association to metastasis in other cancers (8, 9). -

Abstracts from the 53Rd European Society of Human Genetics (ESHG) Conference: Oral Presentations
European Journal of Human Genetics (2020) 28:1–140 https://doi.org/10.1038/s41431-020-00740-6 ABSTRACTS COLLECTION Abstracts from the 53rd European Society of Human Genetics (ESHG) Conference: Oral Presentations © European Society of Human Genetics 2020. Modified from the conference website and published with permission 2020 Volume 28 | Supplement 1 Virtual Conference June 6–9, 2020 Sponsorship: Publication of this supplement was sponsored by the European Society of Human Genetics. All content was reviewed and approved by the ESHG Scientific Programme Committee, which held full responsibility for the abstract selections. 1234567890();,: 1234567890();,: Disclosure Information: In order to help readers form their own judgments of potential bias in published abstracts, authors are asked to declare any competing financial interests. Contributions of up to EUR 10 000.- (Ten thousand Euros, or equivalent value in kind) per year per company are considered “Modest”. Contributions above EUR 10 000.- per year are considered “Significant”. Presenting authors are indicated with asterisks in the contributor lists. Plenary Sessions somatic mutations affecting RNA splicing factors and tumorigenesis, the promise of correct mis-splicing for can- PL1 Opening Plenary cer therapy, and the recent creation of CRISPR/Cas-based technologies for conducting mRNA isoform-level genetic PL1.2 screens. RNA splicing defects in cancer R. K. Bradley: None. R. K. Bradley* PL2 What’s New? Highlight Session Fred Hutchinson Cancer Research Center, Seattle, WA, PL2.1 United States Evaluation of DNA methylation episignatures for diag- nosis and phenotype correlations in 42 Mendelian neu- Alternative RNA splicing, the process whereby a single rodevelopmental disorders gene can give rise to many different proteins, has long been known to be dysregulated in many cancers.