Roles of NRF3 in the Hallmarks of Cancer: Proteasomal Inactivation of Tumor Suppressors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expression Profiling of KLF4
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in wild type MEFs. 98 Am J Cancer Res 2011;1(1):85-97 Table S1: Functional Annotation Clustering of Genes Up-Regulated in Klf4 -Null MEFs ILLUMINA_ID Gene Symbol Gene Name (Description) P -value Fold-Change Cell Cycle 8.00E-03 ILMN_1217331 Mcm6 MINICHROMOSOME MAINTENANCE DEFICIENT 6 40.36 ILMN_2723931 E2f6 E2F TRANSCRIPTION FACTOR 6 26.8 ILMN_2724570 Mapk12 MITOGEN-ACTIVATED PROTEIN KINASE 12 22.19 ILMN_1218470 Cdk2 CYCLIN-DEPENDENT KINASE 2 9.32 ILMN_1234909 Tipin TIMELESS INTERACTING PROTEIN 5.3 ILMN_1212692 Mapk13 SAPK/ERK/KINASE 4 4.96 ILMN_2666690 Cul7 CULLIN 7 2.23 ILMN_2681776 Mapk6 MITOGEN ACTIVATED PROTEIN KINASE 4 2.11 ILMN_2652909 Ddit3 DNA-DAMAGE INDUCIBLE TRANSCRIPT 3 2.07 ILMN_2742152 Gadd45a GROWTH ARREST AND DNA-DAMAGE-INDUCIBLE 45 ALPHA 1.92 ILMN_1212787 Pttg1 PITUITARY TUMOR-TRANSFORMING 1 1.8 ILMN_1216721 Cdk5 CYCLIN-DEPENDENT KINASE 5 1.78 ILMN_1227009 Gas2l1 GROWTH ARREST-SPECIFIC 2 LIKE 1 1.74 ILMN_2663009 Rassf5 RAS ASSOCIATION (RALGDS/AF-6) DOMAIN FAMILY 5 1.64 ILMN_1220454 Anapc13 ANAPHASE PROMOTING COMPLEX SUBUNIT 13 1.61 ILMN_1216213 Incenp INNER CENTROMERE PROTEIN 1.56 ILMN_1256301 Rcc2 REGULATOR OF CHROMOSOME CONDENSATION 2 1.53 Extracellular Matrix 5.80E-06 ILMN_2735184 Col18a1 PROCOLLAGEN, TYPE XVIII, ALPHA 1 51.5 ILMN_1223997 Crtap CARTILAGE ASSOCIATED PROTEIN 32.74 ILMN_2753809 Mmp3 MATRIX METALLOPEPTIDASE -

NFE2L1 Antibody Cat
NFE2L1 Antibody Cat. No.: 31-335 NFE2L1 Antibody Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human, Mouse, Rat Antibody produced in rabbits immunized with a synthetic peptide corresponding a region IMMUNOGEN: of human NFE2L1. TESTED APPLICATIONS: ELISA, WB NFE2L1 antibody can be used for detection of NFE2L1 by ELISA at 1:7862500. NFE2L1 APPLICATIONS: antibody can be used for detection of NFE2L1 by western blot at 1 μg/mL, and HRP conjugated secondary antibody should be diluted 1:50,000 - 100,000. POSITIVE CONTROL: 1) Cat. No. 1205 - Jurkat Cell Lysate PREDICTED MOLECULAR 85 kDa WEIGHT: Properties PURIFICATION: Antibody is purified by peptide affinity chromatography method. CLONALITY: Polyclonal CONJUGATE: Unconjugated PHYSICAL STATE: Liquid September 27, 2021 1 https://www.prosci-inc.com/nfe2l1-antibody-31-335.html Purified antibody supplied in 1x PBS buffer with 0.09% (w/v) sodium azide and 2% BUFFER: sucrose. CONCENTRATION: batch dependent For short periods of storage (days) store at 4˚C. For longer periods of storage, store STORAGE CONDITIONS: NFE2L1 antibody at -20˚C. As with any antibody avoid repeat freeze-thaw cycles. Additional Info OFFICIAL SYMBOL: NFE2L1 ALTERNATE NAMES: NFE2L1, FLJ00380, LCR-F1, NRF1, TCF11 ACCESSION NO.: NP_003195 PROTEIN GI NO.: 4505379 GENE ID: 4779 USER NOTE: Optimal dilutions for each application to be determined by the researcher. Background and References NFE2L1 activates erythroid-specific, globin gene expression. This gene encodes a protein that is involved in globin gene expression in erythrocytes. Confusion has occurred in bibliographic databases due to the shared symbol of NRF1 for this gene, NFE2L1, and for BACKGROUND: 'nuclear respiratory factor 1' which has an official symbol of NRF1. -
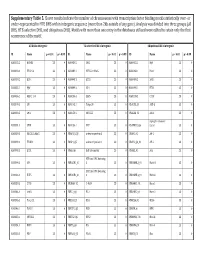
Supplementary Table 5. Clover Results Indicate the Number Of
Supplementary Table 5. Clover results indicate the number of chromosomes with transcription factor binding motifs statistically over‐ or under‐represented in HTE DHS within intergenic sequence (more than 2kb outside of any gene). Analysis was divided into three groups (all DHS, HTE‐selective DHS, and ubiquitous DHS). Motifs with more than one entry in the databases utilized were edited to retain only the first occurrence of the motif. All DHS x Intergenic TEselective DHS x Intergenic Ubiquitous DHS x Intergenic ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 MA0002.2 RUNX1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0003.1 TFAP2A 23 0 MA0089.1 NFE2L1::MafG 23 0 MA0068.1 Pax4 23 0 MA0039.2 Klf4 23 0 MA0098.1 ETS1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0099.2 AP1 23 0 MA0098.1 ETS1 23 0 MA0056.1 MZF1_1‐4 23 0 MA0136.1 ELF5 23 0 MA0139.1 CTCF 23 0 MA0079.2 SP1 23 0 MA0145.1 Tcfcp2l1 23 0 V$ALX3_01 ALX‐3 23 0 MA0080.2 SPI1 23 0 MA0150.1 NFE2L2 23 0 V$ALX4_02 Alx‐4 23 0 myocyte enhancer MA0081.1 SPIB 23 0 MA0156.1 FEV 23 0 V$AMEF2_Q6 factor 23 0 MA0089.1 NFE2L1::MafG 23 0 V$AP1FJ_Q2 activator protein 1 23 0 V$AP1_01 AP‐1 23 0 MA0090.1 TEAD1 23 0 V$AP4_Q5 activator protein 4 23 0 V$AP2_Q6_01 AP‐2 23 0 MA0098.1 ETS1 23 0 V$AR_Q6 half‐site matrix 23 0 V$ARX_01 Arx 23 0 BTB and CNC homolog MA0099.2 AP1 23 0 V$BACH1_01 1 23 0 V$BARHL1_01 Barhl‐1 23 0 BTB and CNC homolog MA0136.1 ELF5 23 0 V$BACH2_01 2 23 0 V$BARHL2_01 Barhl2 23 0 MA0139.1 CTCF 23 0 V$CMAF_02 C‐MAF 23 0 V$BARX1_01 Barx1 23 0 MA0144.1 Stat3 23 0 -

Micrornas As Modulators of Smoking-Induced Gene Expression Changes in Human Airway Epithelium
MicroRNAs as modulators of smoking-induced gene expression changes in human airway epithelium Frank Schembria,1, Sriram Sridharb,1, Catalina Perdomoc, Adam M. Gustafsond, Xiaoling Zhangd, Ayla Ergune, Jining Lua, Gang Liua, Xiaohui Zhanga, Jessica Bowersf, Cyrus Vazirib, Kristen Ottc, Kelly Sensingerf, James J. Collinse, Jerome S. Brodya, Robert Gettsf, Marc E. Lenburga,b,d, and Avrum Spiraa,b,d,2 aPulmonary Center, bDepartment of Pathology and Laboratory Medicine, and cGenetics and Genomics Graduate Program, Boston University School of Medicine, Boston, MA 02118; dBioinformatics Program, Boston University School of Engineering, Boston, MA 02215; eDepartment of Biomedical Engineering and Center for BioDynamics and Howard Hughes Medical Institute, Boston University, Boston, MA 02215; and fGenisphere, Inc., Hatfield, PA 19440 Edited by Charles R. Cantor, Sequenom Inc., San Diego, CA, and approved November 21, 2008 (received for review July 3, 2008) We have shown that smoking impacts bronchial airway gene Hundreds of human miRNAs have been cloned (11), with each of expression and that heterogeneity in this response associates with them predicted to regulate hundreds of genes by complementary smoking-related disease risk. In this study, we sought to determine binding to the 3Ј-untranslated region (UTR) of target mRNAs (12). whether microRNAs (miRNAs) play a role in regulating the airway A number of miRNAs have been shown to play important roles in gene expression response to smoking. We examined whole-ge- developmental timing, cell death, carcinogenesis, and stress responses in nome miRNA and mRNA expression in bronchial airway epithelium vitro (13–16), such as hypoxia (17). Whereas miRNA expression has and found 28 miRNAs to been examined in lung tumors (18–24), neither the expression of (20 ؍ from current and never smokers (n be differentially expressed (P < 0.05) with the majority being miRNA in bronchial epithelium nor the effect of smoking on bronchial down-regulated in smokers. -
Reprogramming of Lysosomal Gene Expression by Interleukin-4 and Stat6 Brignull Et Al
Reprogramming of lysosomal gene expression by interleukin-4 and Stat6 Brignull et al. Brignull et al. BMC Genomics 2013, 14:853 http://www.biomedcentral.com/1471-2164/14/853 Brignull et al. BMC Genomics 2013, 14:853 http://www.biomedcentral.com/1471-2164/14/853 RESEARCH ARTICLE Open Access Reprogramming of lysosomal gene expression by interleukin-4 and Stat6 Louise M Brignull1†, Zsolt Czimmerer2†, Hafida Saidi1,3, Bence Daniel2, Izabel Villela4,5, Nathan W Bartlett6, Sebastian L Johnston6, Lisiane B Meira4, Laszlo Nagy2,7 and Axel Nohturfft1* Abstract Background: Lysosomes play important roles in multiple aspects of physiology, but the problem of how the transcription of lysosomal genes is coordinated remains incompletely understood. The goal of this study was to illuminate the physiological contexts in which lysosomal genes are coordinately regulated and to identify transcription factors involved in this control. Results: As transcription factors and their target genes are often co-regulated, we performed meta-analyses of array-based expression data to identify regulators whose mRNA profiles are highly correlated with those of a core set of lysosomal genes. Among the ~50 transcription factors that rank highest by this measure, 65% are involved in differentiation or development, and 22% have been implicated in interferon signaling. The most strongly correlated candidate was Stat6, a factor commonly activated by interleukin-4 (IL-4) or IL-13. Publicly available chromatin immunoprecipitation (ChIP) data from alternatively activated mouse macrophages show that lysosomal genes are overrepresented among Stat6-bound targets. Quantification of RNA from wild-type and Stat6-deficient cells indicates that Stat6 promotes the expression of over 100 lysosomal genes, including hydrolases, subunits of the vacuolar H+ ATPase and trafficking factors. -

Identification of Dna Sequence Variants in the Estrogen
IDENTIFICATION OF DNA SEQUENCE VARIANTS IN THE ESTROGEN RECEPTOR PATHWAY IN BREAST CANCER by Seyed Amir Bahreini BSc in Cell & Mol Biology, University of Isfahan, Iran, 2010 Submitted to the Graduate Faculty of Department of Human Genetics Graduate School of Public Health in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2016 UNIVERSITY OF PITTSBURGH Graduate School of Public Health This dissertation was presented by Seyed Amir Bahreini It was defended on April 6, 2016 and approved by Dissertation Advisor: Steffi Oesterreich, PhD Professor Department of Pharmacology and Chemical Biology School of Medicine University of Pittsburgh Committee Members: Candace M. Kammerer, PhD Professor Department of Human Genetics Graduate School of Public Health University of Pittsburgh Adrian V. Lee, PhD Professor Department of Pharmacology and Chemical Biology School of Medicine University of Pittsburgh Ryan L. Minster, PhD Professor Department of Human Genetics Graduate School of Public Health University of Pittsburgh ii Copyright © by Seyed Amir Bahreini 2016 iii Steffi Oesterreich, PhD IDENTIFICATION OF DNA SEQUENCE VARIANTS IN THE ESTROGEN RECEPTOR PATHWAY IN BREAST CANCER Seyed Amir Bahreini, PhD University of Pittsburgh, 2016 ABSTRACT Breast cancer is of public health importance with an increasing incidence over the past decade. Estrogen Receptor (ER) activity is critical for promoting majority of breast cancers. Inhibiting ER is one of the most successful targeted therapies in oncology. Studies have suggested that genomic variation in ER binding sites and ESR1 gene may be responsible for endocrine treatment response and cancer progression. We investigated the role of single nucleotide variants (SNVs) in the ER pathway in breast cancer, including clinically relevant mutations in ER gene and regulatory variants in ER binding sites. -

Nfe2l1) As a Hypoxia- Inducible Factor and Its Oxygen-Dependent Regulation in Vitro and in Vivo
IDENTIFICATION OF NRFl (NFE2L1) AS A HYPOXIA- INDUCIBLE FACTOR AND ITS OXYGEN-DEPENDENT REGULATION IN VITRO AND IN VIVO by NIKOLAI L. CHEPELEV B. Sc. H. Carleton University, 2006 A thesis submitted to the Faculty of Graduate and Postdoctoral Affairs in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biology Department of Biology Ottawa-Carleton Institute of Biology Carleton University Ottawa, Ontario, Canada September 2011 © 2011 Nikolai Chepelev Library and Archives Bibliotheque et 1*1 Canada Archives Canada Published Heritage Direction du Branch Patrimoine de I'edition 395 Wellington Street 395, rue Wellington OttawaONK1A0N4 OttawaONK1A0N4 Canada Canada Your file Votre rdterence ISBN: 978-0-494-83249-3 Our file Notre reference ISBN: 978-0-494-83249-3 NOTICE: AVIS: The author has granted a non L'auteur a accorde une licence non exclusive exclusive license allowing Library and permettant a la Bibliotheque et Archives Archives Canada to reproduce, Canada de reproduire, publier, archiver, publish, archive, preserve, conserve, sauvegarder, conserver, transmettre au public communicate to the public by par telecommunication ou par I'lnternet, prefer, telecommunication or on the Internet, distribuer et vendre des theses partout dans le loan, distribute and sell theses monde, a des fins commerciales ou autres, sur worldwide, for commercial or non support microforme, papier, electronique et/ou commercial purposes, in microform, autres formats. paper, electronic and/or any other formats. The author retains copyright L'auteur conserve la propriete du droit d'auteur ownership and moral rights in this et des droits moraux qui protege cette these. Ni thesis. Neither the thesis nor la these ni des extraits substantiels de celle-ci substantial extracts from it may be ne doivent etre imprimes ou autrement printed or otherwise reproduced reproduits sans son autorisation. -

A MAFG-Lncrna Axis Links Systemic Nutrient Abundance to Hepatic Glucose Metabolism
ARTICLE https://doi.org/10.1038/s41467-020-14323-y OPEN A MAFG-lncRNA axis links systemic nutrient abundance to hepatic glucose metabolism Marta Pradas-Juni et al.# Obesity and type 2 diabetes mellitus are global emergencies and long noncoding RNAs (lncRNAs) are regulatory transcripts with elusive functions in metabolism. Here we show that a high fraction of lncRNAs, but not protein-coding mRNAs, are repressed during diet-induced 1234567890():,; obesity (DIO) and refeeding, whilst nutrient deprivation induced lncRNAs in mouse liver. Similarly, lncRNAs are lost in diabetic humans. LncRNA promoter analyses, global cistrome and gain-of-function analyses confirm that increased MAFG signaling during DIO curbs lncRNA expression. Silencing Mafg in mouse hepatocytes and obese mice elicits a fasting-like gene expression profile, improves glucose metabolism, de-represses lncRNAs and impairs mammalian target of rapamycin (mTOR) activation. We find that obesity-repressed LincIRS2 is controlled by MAFG and observe that genetic and RNAi-mediated LincIRS2 loss causes elevated blood glucose, insulin resistance and aberrant glucose output in lean mice. Taken together, we identify a MAFG-lncRNA axis controlling hepatic glucose metabolism in health and metabolic disease. #A full list of authors and their affiliations appears at the end of the paper. NATURE COMMUNICATIONS | (2020) 11:644 | https://doi.org/10.1038/s41467-020-14323-y | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-020-14323-y ellular and organism-level energy homeostasis and nutri- by MAFG and CRISPR–Cas9-mediated knockout, or antisense- Cent partitioning are instrumental for survival. In higher mediated RNA interference of LincIRS2 causes hyperglycemia, organisms, multi-organ systems evolved to react to shifts insulin resistance, likely caused by alterations in glucogenic in energy supply by storing (anabolic) or metabolizing (catabolic) gene expression in lean mice. -

Distinct Isoforms of Nrf1 Diversely Regulate Different Subsets of Its Cognate Target Genes
bioRxiv preprint doi: https://doi.org/10.1101/356071; this version posted June 28, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Distinct isoforms of Nrf1 diversely regulate different subsets of its cognate target genes Meng Wang, Lu Qiu, Xufang Ru, Yijiang Song, Yiguo Zhang* The Laboratory of Cell Biochemistry and Topogenetic Regulation, College of Bioengineering and Faculty of Sciences, Chongqing University, No. 174 Shazheng Street, Shapingba District, Chongqing 400044, China *Correspondence should be addressed to Yiguo Zhang (email: [email protected] or [email protected]). Abstract The single Nrf1 gene has capability to be differentially transcripted alongside with alternative mRNA-splicing and subsequent translation through different initiation signals so as to yield distinct lengths of polypeptide isoforms. Amongst them, three of the most representatives are Nrf1α, Nrf1β and Nrf1γ, but the putative specific contribution of each isoform to regulating ARE-driven target genes remains unknown. To address this, we have here established three cell lines on the base of the Flp-In™ T-REx™ system, which are allowed for tetracycline-inducibly stable expression of Nrf1α, Nrf1β and Nrf1γ. The RNA-Sequencing results have demonstrated that a vast majority of differentially expressed genes (i.e. 90 DEGs detected) were dominantly up-regulated by Nrf1α and/or Nrf1β following induction by tetracycline. By contrast, other DEGs regulated by Nrf1γ were far less than those regulated by Nrf1α/β (i.e. ~11 of Nrf1α and 7 of Nrf1β). Further transcriptomic analysis revealed that tetracycline-induced expression of Nrf1γ significantly increased the percentage of down-regulated genes in total DEGs. -

Is Regulated by O-Glcnac Transferase
UC Irvine UC Irvine Previously Published Works Title Nuclear factor-erythroid-2 related transcription factor-1 (Nrf1) is regulated by O-GlcNAc transferase. Permalink https://escholarship.org/uc/item/5mp8q65s Journal Free radical biology & medicine, 110 ISSN 0891-5849 Authors Han, Jeong Woo Valdez, Joshua L Ho, Daniel V et al. Publication Date 2017-09-01 DOI 10.1016/j.freeradbiomed.2017.06.008 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Free Radical Biology and Medicine 110 (2017) 196–205 Contents lists available at ScienceDirect Free Radical Biology and Medicine journal homepage: www.elsevier.com/locate/freeradbiomed Original article Nuclear factor-erythroid-2 related transcription factor-1 (Nrf1) is regulated MARK by O-GlcNAc transferase Jeong Woo Hana,1, Joshua L. Valdeza,1, Daniel V. Hoa, Candy S. Leea, Hyun Min Kima, ⁎ Xiaorong Wangb, Lan Huangb,Jefferson Y. Chana, a Department of Laboratory Medicine and Pathology, University of California, Irvine, D440 Medical Sciences, Irvine, CA 92697, USA b Departments of Physiology and Biophysics, University of California, Irvine, D440 Medical Sciences, Irvine, CA 92697, USA ARTICLE INFO ABSTRACT Keywords: The Nrf1 (Nuclear factor E2-related factor 1) transcription factor performs a critical role in regulating cellular Oxidative stress homeostasis. Using a proteomic approach, we identified Host Cell Factor-1 (HCF1), a co-regulator of tran- O-GlcNAcylation scription, and O-GlcNAc transferase (OGT), the enzyme that mediates protein O-GlcNAcylation, as cellular OGT partners of Nrf1a, an isoform of Nrf1. Nrf1a directly interacts with HCF1 through the HCF1 binding motif Transcription factor (HBM), while interaction with OGT is mediated through HCF1. -

Oxidized Phospholipids Regulate Amino Acid Metabolism Through MTHFD2 to Facilitate Nucleotide Release in Endothelial Cells
ARTICLE DOI: 10.1038/s41467-018-04602-0 OPEN Oxidized phospholipids regulate amino acid metabolism through MTHFD2 to facilitate nucleotide release in endothelial cells Juliane Hitzel1,2, Eunjee Lee3,4, Yi Zhang 3,5,Sofia Iris Bibli2,6, Xiaogang Li7, Sven Zukunft 2,6, Beatrice Pflüger1,2, Jiong Hu2,6, Christoph Schürmann1,2, Andrea Estefania Vasconez1,2, James A. Oo1,2, Adelheid Kratzer8,9, Sandeep Kumar 10, Flávia Rezende1,2, Ivana Josipovic1,2, Dominique Thomas11, Hector Giral8,9, Yannick Schreiber12, Gerd Geisslinger11,12, Christian Fork1,2, Xia Yang13, Fragiska Sigala14, Casey E. Romanoski15, Jens Kroll7, Hanjoong Jo 10, Ulf Landmesser8,9,16, Aldons J. Lusis17, 1234567890():,; Dmitry Namgaladze18, Ingrid Fleming2,6, Matthias S. Leisegang1,2, Jun Zhu 3,4 & Ralf P. Brandes1,2 Oxidized phospholipids (oxPAPC) induce endothelial dysfunction and atherosclerosis. Here we show that oxPAPC induce a gene network regulating serine-glycine metabolism with the mitochondrial methylenetetrahydrofolate dehydrogenase/cyclohydrolase (MTHFD2) as a cau- sal regulator using integrative network modeling and Bayesian network analysis in human aortic endothelial cells. The cluster is activated in human plaque material and by atherogenic lipo- proteins isolated from plasma of patients with coronary artery disease (CAD). Single nucleotide polymorphisms (SNPs) within the MTHFD2-controlled cluster associate with CAD. The MTHFD2-controlled cluster redirects metabolism to glycine synthesis to replenish purine nucleotides. Since endothelial cells secrete purines in response to oxPAPC, the MTHFD2- controlled response maintains endothelial ATP. Accordingly, MTHFD2-dependent glycine synthesis is a prerequisite for angiogenesis. Thus, we propose that endothelial cells undergo MTHFD2-mediated reprogramming toward serine-glycine and mitochondrial one-carbon metabolism to compensate for the loss of ATP in response to oxPAPC during atherosclerosis. -

HIV Protease Inhibitor Nelfinavir Targets Human DDI2 and Potentiates Proteasome Inhibitor-Based Chemotherapy
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.03.075572; this version posted May 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. HIV Protease Inhibitor Nelfinavir Targets Human DDI2 and Poten- tiates Proteasome Inhibitor-based Chemotherapy Yuan Gu1,#, Xin Wang1,#, Yu Wang1,#, Jie Li2, Fa-Xing Yu1,* 1 Institute of Pediatrics, Children’s Hospital of Fudan University, and the Shanghai Key Laboratory of Medical Epigenetics, Institutes of Biomedical Sciences, Fudan University, Shanghai, China 2 Large-scale Protein Preparation System, National Facility for Protein Sciences, Shanghai, China #, These authors contributed equally * Corresponding author: Fa-Xing Yu Keywords: protease inhibitor, HIV‐1 protease, anticancer drug, proteasome, multiple myeloma, Nelfinavir, DNA dam- age inducible 1 homolog 2 Abstract: Proteasome inhibitors (PIs) are currently used in the clinic to treat cancers such as multiple myeloma (MM). However, cancer cells often rapidly develop drug resistance towards PIs due to a compensatory mechanism mediated by nuclear factor erythroid 2 like 1 (NFE2L1) and aspartic protease DNA damage inducible 1 homolog 2 (DDI2). Following DDI2-mediated cleavage, NFE2L1 is able to induce transcription of virtually all proteasome subunit genes. Under normal condition, cleaved NFE2L1 is constantly de- graded by proteasome, whereas in the presence of PIs, it accumu- lates and induces proteasome synthesis which in turn promotes the development of drug resistance towards PIs. Here, we report that Nelfinavir (NFV), an HIV protease inhibitor, can inhibit DDI2 ac- tivity directly. Inhibition of DDI2 by NFV effectively blocks NFE2L1 proteolysis and potentiates cytotoxicity of PIs in cancer cells.