Orbiviruses: a North American Perspective
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Guide for Common Viral Diseases of Animals in Louisiana
Sampling and Testing Guide for Common Viral Diseases of Animals in Louisiana Please click on the species of interest: Cattle Deer and Small Ruminants The Louisiana Animal Swine Disease Diagnostic Horses Laboratory Dogs A service unit of the LSU School of Veterinary Medicine Adapted from Murphy, F.A., et al, Veterinary Virology, 3rd ed. Cats Academic Press, 1999. Compiled by Rob Poston Multi-species: Rabiesvirus DCN LADDL Guide for Common Viral Diseases v. B2 1 Cattle Please click on the principle system involvement Generalized viral diseases Respiratory viral diseases Enteric viral diseases Reproductive/neonatal viral diseases Viral infections affecting the skin Back to the Beginning DCN LADDL Guide for Common Viral Diseases v. B2 2 Deer and Small Ruminants Please click on the principle system involvement Generalized viral disease Respiratory viral disease Enteric viral diseases Reproductive/neonatal viral diseases Viral infections affecting the skin Back to the Beginning DCN LADDL Guide for Common Viral Diseases v. B2 3 Swine Please click on the principle system involvement Generalized viral diseases Respiratory viral diseases Enteric viral diseases Reproductive/neonatal viral diseases Viral infections affecting the skin Back to the Beginning DCN LADDL Guide for Common Viral Diseases v. B2 4 Horses Please click on the principle system involvement Generalized viral diseases Neurological viral diseases Respiratory viral diseases Enteric viral diseases Abortifacient/neonatal viral diseases Viral infections affecting the skin Back to the Beginning DCN LADDL Guide for Common Viral Diseases v. B2 5 Dogs Please click on the principle system involvement Generalized viral diseases Respiratory viral diseases Enteric viral diseases Reproductive/neonatal viral diseases Back to the Beginning DCN LADDL Guide for Common Viral Diseases v. -

Rapid Identification of Known and New RNA Viruses from Animal Tissues
Rapid Identification of Known and New RNA Viruses from Animal Tissues Joseph G. Victoria1,2*, Amit Kapoor1,2, Kent Dupuis3, David P. Schnurr3, Eric L. Delwart1,2 1 Department of Molecular Virology, Blood Systems Research Institute, San Francisco, California, United States of America, 2 Department of Laboratory Medicine, University of California, San Francisco, California, United States of America, 3 Viral and Rickettsial Disease Laboratory, Division of Communicable Disease Control, California State Department of Public Health, Richmond, California, United States of America Abstract Viral surveillance programs or diagnostic labs occasionally obtain infectious samples that fail to be typed by available cell culture, serological, or nucleic acid tests. Five such samples, originating from insect pools, skunk brain, human feces and sewer effluent, collected between 1955 and 1980, resulted in pathology when inoculated into suckling mice. In this study, sequence-independent amplification of partially purified viral nucleic acids and small scale shotgun sequencing was used on mouse brain and muscle tissues. A single viral agent was identified in each sample. For each virus, between 16% to 57% of the viral genome was acquired by sequencing only 42–108 plasmid inserts. Viruses derived from human feces or sewer effluent belonged to the Picornaviridae family and showed between 80% to 91% amino acid identities to known picornaviruses. The complete polyprotein sequence of one virus showed strong similarity to a simian picornavirus sequence in the provisional Sapelovirus genus. Insects and skunk derived viral sequences exhibited amino acid identities ranging from 25% to 98% to the segmented genomes of viruses within the Reoviridae family. Two isolates were highly divergent: one is potentially a new species within the orthoreovirus genus, and the other is a new species within the orbivirus genus. -

Peruvian Horse Sickness Virus and Yunnan Orbivirus, Isolated from Vertebrates and Mosquitoes in Peru and Australia
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Virology 394 (2009) 298–310 Contents lists available at ScienceDirect Virology journal homepage: www.elsevier.com/locate/yviro Peruvian horse sickness virus and Yunnan orbivirus, isolated from vertebrates and mosquitoes in Peru and Australia Houssam Attoui a,⁎,1, Maria Rosario Mendez-lopez b,⁎,1, Shujing Rao c,1, Ana Hurtado-Alendes b,1, Frank Lizaraso-Caparo b, Fauziah Mohd Jaafar a, Alan R. Samuel a, Mourad Belhouchet a, Lindsay I. Pritchard d, Lorna Melville e, Richard P. Weir e, Alex D. Hyatt d, Steven S. Davis e, Ross Lunt d, Charles H. Calisher f, Robert B. Tesh g, Ricardo Fujita b, Peter P.C. Mertens a a Department of Vector Borne Diseases, Institute for Animal Health, Pirbright, Woking, Surrey, GU24 0NF, UK b Research Institute and Institute of Genetics and Molecular Biology, Universidad San Martín de Porres Medical School, Lima, Perú c Clemson University, 114 Long Hall, Clemson, SC 29634-0315, USA d Australian Animal Health Laboratory, CSIRO, Geelong, Victoria, Australia e Northern Territory Department of Primary Industries, Fisheries and Mines, Berrimah Veterinary Laboratories, Berrimah, Northern Territory 0801, Australia f Department of Microbiology, Immunology and Pathology, College of Veterinary Medicine and Biomedical Sciences, Colorado State University, Fort Collins, CO 80523, USA g Department of Pathology, University of Texas Medical Branch, 301 University Boulevard, Galveston, TX 77555-0609, USA article info abstract Article history: During 1997, two new viruses were isolated from outbreaks of disease that occurred in horses, donkeys, Received 11 June 2009 cattle and sheep in Peru. -

Adenovirus Infections in Humans
CHAPTER 11 Adenovirus Infections in Humans STEPHEN E. STRAUS I. INTRODUCTION Adenoviruses are ubiquitous agents that infect humans of all ages. The discovery of the first adenovirus types three decades ago paved the way for countless studies that continue to uncover additional viral strains and ever expand our comprehension of their significance to man. Whereas some human adenoviruses are being examined at the increasingly so phisticated molecular and cellular levels, the means by which they infect man, provoke illness, or are handled by the body's immune system remain very poorly understood. This chapter represents an attempt to catalogue the known human adenovirus agents, to summarize the data pertaining to their acquisition and transmission, to review the range of illness with which they are associated, and to highlight those aspects of adenovirus infection that are in need of further investigation. II. ADENOVIRUSES RECOVERED FROM HUMANS The 41 distinct adenovirus types that have been recovered from hu mans thus far are listed in Table I. Most of these agents were isolated during an extremely fruitful decade of studies that followed the initial discovery of adenoviruses. A wide range of illnesses has been associated with the better-defined, lower-numbered virus types, but the predilection STEPHEN E. STRAUS • Medical Virology Section, Laboratory of Clinical Investigation, National Institutes of Health, Bethesda, Maryland 20205. 451 H. S. Ginsberg (ed.), The Adenoviruses © Plenum Press, New York 1984 452 STEPHEN E. STRAUS TABLE I. Adenovirus ImmunotypesQ Major Prototype associated Type strain Source Patient diagnosis diseasesb Ad71 Adenoid Hypertrophied tonsils and Respiratory adenoids 2 Ad6 Adenoid Hypertrophied tonsils and Respiratory adenoids 3 G.B. -
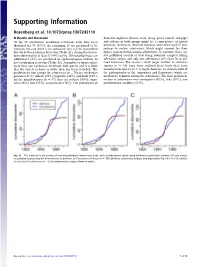
Supporting Information
Supporting Information Rosenberg et al. 10.1073/pnas.1307243110 SI Results and Discussion domestic ungulates (horses, cows, sheep, goats, camels, and pigs) Of the 83 arboviruses, nonhuman vertebrate hosts have been and rodents in both groups might be a consequence of spatial identified for 70 (84%); the remaining 13 are presumed to be proximity to humans. Sentinel monkeys were often used in pro- zoonoses because there is no indication they can be transmitted cedures to isolate arboviruses, which might account for their directly between humans by vectors (Table S1). Animal hosts have higher representation among arboviruses. In contrast, there are been identified for at least 57 (44%) of the 130 nonarboviruses; an few published records of bats being routinely sampled during additional 5 (8%) are presumed on epidemiological evidence to arbovirus studies, and only two arboviruses (3%) have been iso- have nonhuman reservoirs (Table S1). A number of viruses infect lated from bats. The reason a much larger number of arbovirus more than one nonhuman vertebrate host species and it is likely species (n = 16) have been isolated from birds than have that the variety of hosts is wider than has been recorded. The nonarbovirus species (n = 1) might, however, be characteristic of predominant host groups for arboviruses (n = 70) are nonhuman the pathogenicity of the togaviruses and flaviviruses, which are primates (31%), rodents (29%), ungulates (26%), and birds (23%); much more common among the arboviruses. The most prominent for the nonarboviruses (n = 57), they are rodents (30%), ungu- vectors of arboviruses were mosquitoes (67%), ticks (19%), and lates (26%), bats (23%), and primates (16%). -

Protein Composition of the Hepatitis a Virus Quasi-Envelope
Protein composition of the hepatitis A virus quasi-envelope Kevin L. McKnighta,b,c, Ling Xiec,d, Olga González-Lópeza,b,c, Efraín E. Rivera-Serranoa,b,c, Xian Chenc,d, and Stanley M. Lemona,b,c,1 aDepartment of Medicine, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599-7292; bDepartment of Microbiology and Immunology, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599-7292; cLineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599-7295; and dDepartment of Biochemistry and Biophysics, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599-7260 Edited by Mary K. Estes, Baylor College of Medicine, Houston, TX, and approved April 12, 2017 (received for review November 27, 2016) The Picornaviridae are a diverse family of RNA viruses including many within extracellular vesicles before cell lysis, often with many pathogens of medical and veterinary importance. Classically consid- capsids packaged within a single vesicle (4–6). The cellular egress ered “nonenveloped,” recent studies show that some picornaviruses, of these classically nonenveloped viruses in membrane-bound notably hepatitis A virus (HAV; genus Hepatovirus) and some mem- vesicles has blurred the distinction between enveloped and bers of the Enterovirus genus, are released from cells nonlytically in nonenveloped viruses and has important implications for path- membranous vesicles. To better understand the biogenesis of quasi- ogenesis (2). enveloped HAV (eHAV) virions, we conducted a quantitative proteo- Several lines of evidence indicate that the biogenesis of quasi- mics analysis of eHAV purified from cell-culture supernatant fluids by enveloped eHAV virions is dependent upon components of the isopycnic ultracentrifugation. -

Taxonomy and Comparative Virology of the Influenza Viruses
2 Taxonomy and Comparative Virology of the Influenza Viruses INTRODUCTION Viral taxonomy has evolved slowly (and often contentiously) from a time when viruses were identified at the whim of the investigator by place names, names of persons (investigator or patient), sigla, Greco-Latin hybrid names, host of origin, or name of associated disease. Originally studied by pathologists and physicians, viruses were first named for the diseases they caused or the lesions they induced. Yellow fever virus turned its victims yellow with jaundice, and the virus now known as poliovirus destroyed the anterior horn cells or gray (polio) matter of the spinal cord. But the close kinship of polioviruses with coxsackievirus B1, the cause of the epidemic pleurodynia, is not apparent from names derived variously from site of pathogenic lesion and place of original virus isolation (Cox sackie, New York). In practice, many of the older names of viruses remain in common use, but the importance of a formal, more regularized approach to viral classification has been increasingly recognized by students and practitioners of virology as more basic information has become available about the nature of viruses. Accordingly, the International Committee on Taxonomy of Viruses has devised a generally ac cepted system for the classification and nomenclature of viruses that serves the needs of comparative virology in formulating regular guidelines for viral nomen clature. Although this nomenclature remains highly diversified and at times in consistent, retaining many older names, it affirms "an effort ... toward a latinized nomenclature" (Matthews, 1981) and places primary emphasis on virus structure and replication in viral classification. -

Genetic and Epidemiological Characterization of Stretch Lagoon Orbivirus, a Novel Orbivirus Isolated from Culex and Aedes Mosquitoes in Northern Australia
Journal of General Virology (2009), 90, 1433–1439 DOI 10.1099/vir.0.010074-0 Genetic and epidemiological characterization of Stretch Lagoon orbivirus, a novel orbivirus isolated from Culex and Aedes mosquitoes in northern Australia Chris Cowled,1 Gustavo Palacios,2 Lorna Melville,3 Richard Weir,3 Susan Walsh,3 Steven Davis,3 Aneta Gubala,1 W. Ian Lipkin,2 Thomas Briese2 and David Boyle1 Correspondence 1CSIRO Livestock Industries, Australian Animal Health Laboratory, East Geelong, VIC 3220, Chris Cowled Australia [email protected] 2Center for Infection and Immunity, Mailman School of Public Health, Columbia University, New York, NY, USA 3Northern Territory Department of Primary Industries, Fisheries and Mines, Berrimah Veterinary Laboratories, Berrimah, Northern Territory 0801, Australia Stretch Lagoon orbivirus (SLOV) was isolated in 2002 from pooled Culex annulirostris mosquitoes collected at Stretch Lagoon, near the Wolfe Creek national park in the Kimberley region of Western Australia. Conventional serological tests were unable to identify the isolate, and electron microscopy indicated a virus of the genus Orbivirus, family Reoviridae. Here, a cDNA subtraction method was used to obtain approximately one-third of the viral genome, and further sequencing was performed to complete the sequences of segment 1 (viral polymerase) and segment 2 (conserved inner-core protein). Phylogenetic analysis showed that SLOV should be considered a new species within the genus Orbivirus. A real-time RT-PCR test was designed to study the epidemiology of SLOV in the field. Six additional isolates of SLOV were identified, Received 4 January 2009 including isolates from four additional locations and two additional mosquito species. Horses, Accepted 8 March 2009 donkeys and goats were implicated as potential vertebrate hosts in a serological survey. -

Structure Unveils Relationships Between RNA Virus Polymerases
viruses Article Structure Unveils Relationships between RNA Virus Polymerases Heli A. M. Mönttinen † , Janne J. Ravantti * and Minna M. Poranen * Molecular and Integrative Biosciences Research Programme, Faculty of Biological and Environmental Sciences, University of Helsinki, Viikki Biocenter 1, P.O. Box 56 (Viikinkaari 9), 00014 Helsinki, Finland; heli.monttinen@helsinki.fi * Correspondence: janne.ravantti@helsinki.fi (J.J.R.); minna.poranen@helsinki.fi (M.M.P.); Tel.: +358-2941-59110 (M.M.P.) † Present address: Institute of Biotechnology, Helsinki Institute of Life Sciences (HiLIFE), University of Helsinki, Viikki Biocenter 2, P.O. Box 56 (Viikinkaari 5), 00014 Helsinki, Finland. Abstract: RNA viruses are the fastest evolving known biological entities. Consequently, the sequence similarity between homologous viral proteins disappears quickly, limiting the usability of traditional sequence-based phylogenetic methods in the reconstruction of relationships and evolutionary history among RNA viruses. Protein structures, however, typically evolve more slowly than sequences, and structural similarity can still be evident, when no sequence similarity can be detected. Here, we used an automated structural comparison method, homologous structure finder, for comprehensive comparisons of viral RNA-dependent RNA polymerases (RdRps). We identified a common structural core of 231 residues for all the structurally characterized viral RdRps, covering segmented and non-segmented negative-sense, positive-sense, and double-stranded RNA viruses infecting both prokaryotic and eukaryotic hosts. The grouping and branching of the viral RdRps in the structure- based phylogenetic tree follow their functional differentiation. The RdRps using protein primer, RNA primer, or self-priming mechanisms have evolved independently of each other, and the RdRps cluster into two large branches based on the used transcription mechanism. -

Emerging Issues in Virus Taxonomy Marc H.V
PERSPECTIVES Emerging Issues in Virus Taxonomy Marc H.V. van Regenmortel* and Brian W.J. Mahy† Viruses occupy a unique position in biology. Although are nonliving infectious entities that can be said, at best, to they possess some of the properties of living systems such lead a kind of borrowed life. as having a genome, they are actually nonliving infectious entities and should not be considered microorganisms. A Viruses versus Virus Particles or Virions clear distinction should be drawn between the terms virus, A virus is a general term which denotes any number of virion, and virus species. Species is the most fundamental taxonomic category used in all biological classification. In concrete objects that possess various relational properties 1991, the International Committee on Taxonomy of Viruses (for instance, its host, vector, and infectivity) that arise by (ICTV) decided that the category of virus species should be virtue of a relation with other objects. These relational used in virus classification together with the categories of properties, also called emergent properties, are characteris- genus and family. More than 50 ICTV study groups were tic of the viral biosystem as a whole and are not present in given the task of demarcating the 1,550 viral species that its constituent parts. When a virus undergoes its so-called were recognized in the 7th ICTV report, which was pub- life cycle, it takes on various forms and manifestations, for lished in 2000. We briefly describe the changes in virus instance, as a replicating nucleic acid in the host cell or classification that were introduced in that report. -

Inhibition of Orbivirus Replication by Fluvastatin and Identification of The
viruses Article Inhibition of Orbivirus Replication by Fluvastatin and Identification of the Key Elements of the Mevalonate Pathway Involved Fauziah Mohd Jaafar 1, Baptiste Monsion 1 , Mourad Belhouchet 2, Peter P. C. Mertens 3 and Houssam Attoui 1,* 1 UMR Virologie 1161, INRAE, Ecole Nationale Vétérinaire d’Alfort, ANSES, Université Paris-Est, 94700 Maisons-Alfort, France; [email protected] (F.M.J.); [email protected] (B.M.) 2 Division of Structural Biology, Henry Wellcome Building for Genomic Medicine, Oxford OX3 7BN, UK; [email protected] 3 Sutton Bonington Campus, School of Veterinary Medicine and Science, University of Nottingham, Leicestershire LE12 5RD, UK; [email protected] * Correspondence: [email protected] or [email protected]; Tel.: +33-143967007 Abstract: Statin derivatives can inhibit the replication of a range of viruses, including hepatitis C virus (HCV, Hepacivirus), dengue virus (Flavivirus), African swine fever virus (Asfarviridae) and poliovirus (Picornaviridae). We assess the antiviral effect of fluvastatin in cells infected with orbiviruses (bluetongue virus (BTV) and Great Island virus (GIV)). The synthesis of orbivirus outer-capsid protein VP2 (detected by confocal immunofluorescence imaging) was used to assess levels of virus replication, showing a reduction in fluvastatin-treated cells. A reduction in virus titres of ~1.7 log (98%) in fluvastatin-treated cells was detected by a plaque assay. We have previously identified a fourth Citation: Mohd Jaafar, F.; Monsion, non-structural protein (NS4) of BTV and GIV, showing that it interacts with lipid droplets in infected B.; Belhouchet, M.; Mertens, P.P.C.; cells. -

B Directive 2000/54/Ec of the European
02000L0054 — EN — 24.06.2020 — 002.001 — 1 This text is meant purely as a documentation tool and has no legal effect. The Union's institutions do not assume any liability for its contents. The authentic versions of the relevant acts, including their preambles, are those published in the Official Journal of the European Union and available in EUR-Lex. Those official texts are directly accessible through the links embedded in this document ►B DIRECTIVE 2000/54/EC OF THE EUROPEAN PARLIAMENT AND OF THE COUNCIL of 18 September 2000 on the protection of workers from risks related to exposure to biological agents at work (seventh individual directive within the meaning of Article 16(1) of Directive 89/391/EEC) (OJ L 262, 17.10.2000, p. 21) Amended by: Official Journal No page date ►M1 Commission Directive (EU) 2019/1833 of 24 October 2019 L 279 54 31.10.2019 ►M2 Commission Directive (EU) 2020/739 of 3 June 2020 L 175 11 4.6.2020 02000L0054 — EN — 24.06.2020 — 002.001 — 2 ▼B DIRECTIVE 2000/54/EC OF THE EUROPEAN PARLIAMENT AND OF THE COUNCIL of 18 September 2000 on the protection of workers from risks related to exposure to biological agents at work (seventh individual directive within the meaning of Article 16(1) of Directive 89/391/EEC) CHAPTER I GENERAL PROVISIONS Article 1 Objective 1. This Directive has as its aim the protection of workers against risks to their health and safety, including the prevention of such risks, arising or likely to arise from exposure to biological agents at work.