Growth and Adaptation of Microorganisms on the Cheese Surface Christophe Monnet, Sophie Landaud-Liautaud, Pascal Bonnarme, Dominique Swennen
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Pseudomonas Spp. and Other Psychrotrophic
Pesq. Vet. Bras. 39(10):807-815, October 2019 DOI: 10.1590/1678-5150-PVB-6037 Original Article Livestock Diseases ISSN 0100-736X (Print) ISSN 1678-5150 (Online) PVB-6037 LD Pseudomonas spp. and other psychrotrophic microorganisms in inspected and non-inspected Brazilian Minas Frescal cheese: proteolytic, lipolytic and AprX production potential1 Pedro I. Teider Junior2, José C. Ribeiro Júnior3* , Eric H. Ossugui2, Ronaldo Tamanini2, Juliane Ribeiro2, Gislaine A. Santos2 2 and Vanerli Beloti2 , Amauri A. Alfieri ABSTRACT.- Teider Junior P.I., Ribeiro Júnior J.C., Ossugui E.H., Tamanini R., Ribeiro J., Santos G.A., Pseudomonas spp. and other psychrotrophic microorganisms Pseudomonas spp. and other psychrotrophic in inspected and non-inspected Brazilian Minas Frescal cheese: proteolytic, lipolytic andAlfieri AprX A.A. production& Beloti V. 2019. potential . Pesquisa Veterinária Brasileira 39(10):807-815. Instituto microorganisms in inspected and non- Nacional de Ciência e Tecnologia para a Cadeia Produtiva do Leite, Universidade Estadual de inspected Brazilian Minas Frescal cheese: Londrina, Rodovia Celso Garcia Cid PR-445 Km 380, Cx. Postal 10.011, Campus Universitário, proteolytic, lipolytic and AprX production Londrina, PR 86057-970, Brazil. E-mail: [email protected] The most consumed cheese in Brazil, Minas Frescal cheese (MFC) is highly susceptible to potential microbial contamination and clandestine production and commercialization can pose a risk to consumer health. The storage of this fresh product under refrigeration, although more appropriate, may favor the growth of spoilage psychrotrophic bacteria. The objective of this [Pseudomonas spp. e outros micro-organismos study was to quantify and compare Pseudomonas spp. and other psychrotrophic bacteria in inspected and non-inspected MFC samples, evaluate their lipolytic and proteolytic activities and e não inspecionados: potencial proteolítico, lipolítico e their metalloprotease production potentials. -

Compositional and Functional Comparisons of the Microbiota in the Colostrum and Mature Milk of Dairy Goats
Compositional and functional comparisons of the microbiota in the colostrum and mature milk of dairy goats Zhannur Niyazbekova Northwest Agriculture and Forestry University Xiao-Ting Yao Northwest Agriculture and Forestry University Ming-Jie Liu Northwest Agriculture and Forestry University Nomin Bold Northwest Agriculture and Forestry University Juan-Zhen Tong Northwest Agriculture and Forestry University Jian-Jun Chang Qinghai University Ying Wen Qinghai University Li Li Qinghai University Yong Wang Qinghai University De-Kun Chen Northwest Agriculture and Forestry University Wen-Tao Ma ( [email protected] ) Northwest Agriculture and Forestry University https://orcid.org/0000-0002-4747-1489 Research article Keywords: Goat milk, Colostrum, Mature milk, Milk microbiota, Microbial diversity Posted Date: May 12th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-26653/v1 Page 1/20 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 2/20 Abstract Background Goat milk is essential for the initial development of kids by providing a great source of commensal bacteria. Here we analyzed the microbiota of goat colostrum which was collected daily for ve days post delivery and mature milk collected at the 7th, 10th, 20nd, 30th, 40th, 50th and 60th days, respectively, from three farms of Shaanxi province. Results The result showed that microbial alpha diversity was higher in the mature milk compared with that in the colostrum. According to taxonomy results, Proteobacteria, Firmicutes, -

Exploring the Diversity and Antimicrobial Potential of Marine Actinobacteria from the Comau Fjord in Northern Patagonia, Chile
ORIGINAL RESEARCH published: 19 July 2016 doi: 10.3389/fmicb.2016.01135 Exploring the Diversity and Antimicrobial Potential of Marine Actinobacteria from the Comau Fjord in Northern Patagonia, Chile Agustina Undabarrena 1, Fabrizio Beltrametti 2, Fernanda P. Claverías 1, Myriam González 1, Edward R. B. Moore 3, 4, Michael Seeger 1 and Beatriz Cámara 1* 1 Laboratorio de Microbiología Molecular y Biotecnología Ambiental, Departamento de Química & Centro de Biotecnología Daniel Alkalay Lowitt, Universidad Técnica Federico Santa María, Valparaíso, Chile, 2 Actygea S.r.l., Gerenzano, Italy, 3 Culture Collection University of Gothenburg (CCUG), Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden, 4 Department of Infectious Diseases, Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden Edited by: Bioprospecting natural products in marine bacteria from fjord environments are attractive Learn-Han Lee, due to their unique geographical features. Although, Actinobacteria are well known Monash University Malaysia Campus, Malaysia for producing a myriad of bioactive compounds, investigations regarding fjord-derived Reviewed by: marine Actinobacteria are scarce. In this study, the diversity and biotechnological Atte Von Wright, potential of Actinobacteria isolated from marine sediments within the Comau University of Eastern Finland, Finland fjord, in Northern Chilean Patagonia, were assessed by culture-based approaches. Polpass Arul Jose, Central Salt and Marine Chemicals The 16S rRNA gene sequences revealed that members phylogenetically related Research Institute, India to the Micrococcaceae, Dermabacteraceae, Brevibacteriaceae, Corynebacteriaceae, *Correspondence: Microbacteriaceae, Dietziaceae, Nocardiaceae, and Streptomycetaceae families were Beatriz Cámara [email protected] present at the Comau fjord. A high diversity of cultivable Actinobacteria (10 genera) was retrieved by using only five different isolation media. -

UNIVERSITY of CALIFORNIA RIVERSIDE Gustatory Receptors In
UNIVERSITY OF CALIFORNIA RIVERSIDE Gustatory Receptors in Mosquito Olfaction and Host-Seeking Behavior A Dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Entomology by Genevieve Mitchell Tauxe March 2015 Dissertation Committee: Dr. Anandasankar Ray, Chairperson Dr. Michael E. Adams Dr. Ring T. Cardé Dr. Jocelyn G. Millar Copyright by Genevieve Mitchell Tauxe 2015 The Dissertation of Genevieve Mitchell Tauxe is approved: Committee Chairperson University of California, Riverside Acknowledgements First, I thank the anonymous odor donors who participated in the experiments described here. They variously washed their feet in the bathroom sink, walked around for hours with beads in their socks (which, no, is not terribly comfortable), and continually showed up no matter how many times we kept asking for more socks. Thank you for your generosity, and for your stinky feet. Now go find yourself on page 73. I thank my advisor, Anand Ray, for all of his help and support over my graduate career. He has taught me so much about how to do research, and also how to compose manuscripts, give presentations, and talk about science. I also thank my committee members, Ring Cardé, Jocelyn Millar, and Mike Adams, for offering so much helpful advice and support over the years, both during official committee meetings and on many other occasions. I thank all the Ray lab and Dahanukar lab members for making the laboratory a fun and productive place to work. In particular, Paulina Ngo performed most of the behavior tests reported in Chapter 4, Tom Guda helped with many behavioral assays, and Greg Pask helped with cloning. -

Genomic and Transcriptomic Insights Into Calcium Carbonate Biomineralization by Marine Actinobacterium Brevibacterium Linens BS258
fmicb-08-00602 April 4, 2017 Time: 15:2 # 1 ORIGINAL RESEARCH published: 06 April 2017 doi: 10.3389/fmicb.2017.00602 Genomic and Transcriptomic Insights into Calcium Carbonate Biomineralization by Marine Actinobacterium Brevibacterium linens BS258 Yuying Zhu1,2,3†, Ning Ma1,2,3†, Weihua Jin4, Shimei Wu5 and Chaomin Sun1,2* 1 Key Laboratory of Experimental Marine Biology, Institute of Oceanology, Chinese Academy of Sciences, Qingdao, China, 2 Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory for Marine Science and Technology, Qingdao, China, 3 College of Earth Science, University of Chinese Academy of Sciences, Beijing, China, 4 College of Biotechnology and Bioengineering, Zhejiang University of Technology, Hangzhou, China, 5 College of Life Sciences, Qingdao University, Qingdao, China Calcium carbonate (CaCO3) biomineralization has been investigated due to its wide range of scientific and technological implications, however, the molecular mechanisms of this important geomicrobiological process are largely unknown. Here, a urease- Edited by: positive marine actinobacterium Brevibacterium linens BS258 was demonstrated to Qiang Wang, Institute of Hydrobiology (CAS), China effectively form CaCO3 precipitates. Surprisingly, this bacterium could also dissolve 2C Reviewed by: the formed CaCO3 with the increase of the Ca concentration. To disclose the Xiaoke Hu, mechanisms of biomineralization, the genome of B. linens BS258 was further completely Yantai Institute of Coastal Zone Research (CAS), China sequenced. Interestingly, the expression of three carbonic anhydrases was significantly C Bruno A. Martinez, up-regulated along with the increase of Ca2 concentration and the extent of calcite Lawrence Berkeley National dissolution. Moreover, transcriptome analyses revealed that increasing concentration Laboratory, USA of Ca2C induced KEGG pathways including quorum sensing (QS) in B. -

Psychrobacter Arenosus Bacteremia After Blood Transfusion, France
DISPATCHES rapidly increased to 40°C) and had chills and headache. Psychrobacter The transfusion was stopped and the patient transferred to the Department of Internal Medicine. At examination, there arenosus was no hypotension, jaundice, or red urine. Standard laboratory testing showed no ABO Bacteremia incompatibility, hemoglobinemia, hemoglobinuria, and coagulation disorders. According to recommendations after Blood of the Agence Nationale de Sécurité du Médicament Transfusion, (Saint-Denis, France), 3 sets of aerobic and anaerobic blood cultures (Bactec; Becton Dickinson, Pont de Clay, France France) for the recipient (1 immediately and 2 others 4 hours later) and the remaining part of the third erythrocyte Yvan Caspar, Christine Recule, Patricia Pouzol, unit were sent to the bacteriology laboratory for culture. Bruno Lafeuillade, Marie-Reine Mallaret, Gram staining of a blood smear prepared from the third Max Maurin, and Jacques Croize erythrocyte unit showed a large number (≈106 CFU/mL) of gram-variable coccobacilli. We report a case of transfusion-associated bactere- Samples were placed on Columbia blood agar mia caused by Psychrobacter arenosus. This psychrotoler- ant bacterium was previously isolated in 2004 from coastal (bioMérieux, Marcy L’Etoile, France) and incubated at sea ice and sediments in the Sea of Japan, but not from 37°C in anaerobic or 5% CO2–enriched atmospheres. humans. P. arenosus should be considered a psychrotoler- Sample inoculated into blood culture bottles were ant bacterial species that can cause transfusion-transmitted incubated at 37°C under aerobic and anaerobic conditions bacterial infections. (Figure). The aerobic blood culture bottle of the first sample obtained from the recipient and aerobic cultures of the third erythrocyte unit enabled isolation of the same acteria are the leading cause of transfusion-transmitted gram-variable coccobacilli after incubation for 48 hours infections (1). -

Rind Austrian Hard Cheese Surfaces and Its Production Environment
Animal Science Publications Animal Science 2-2018 Autochthonous facility-specific microbiota dominates washed- rind Austrian hard cheese surfaces and its production environment Narciso M. Quijada University of Veterinary Medicine Vienna Evelyne Mann University of Veterinary Medicine Vienna Martin Wagner University of Veterinary Medicine Vienna David Rodríguez-Lázaro Universidad de Burgos Marta Hernández Instituto Tecnológico Agrario de Castilla y León See next page for additional authors Follow this and additional works at: https://lib.dr.iastate.edu/ans_pubs Part of the Bacteriology Commons, Environmental Microbiology and Microbial Ecology Commons, Food Processing Commons, and the Genetics and Genomics Commons The complete bibliographic information for this item can be found at https://lib.dr.iastate.edu/ ans_pubs/525. For information on how to cite this item, please visit http://lib.dr.iastate.edu/ howtocite.html. This Article is brought to you for free and open access by the Animal Science at Iowa State University Digital Repository. It has been accepted for inclusion in Animal Science Publications by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Autochthonous facility-specific microbiota dominates washed-rind Austrian hard cheese surfaces and its production environment Abstract Cheese ripening involves the succession of complex microbial communities that are responsible for the organoleptic properties of the final products. The food processing environment can act as a source of natural microbial inoculation, especially in traditionally manufactured products. Austrian Vorarlberger Bergkäse (VB) is an artisanal washed-rind hard cheese produced in the western part of Austria without the addition of external ripening cultures. -

Brevibacterium from Austrian Hard Cheese Harbor a Putative Histamine Catabolism Pathway and a Plasmid for Adaptation to the Cheese Environment
Animal Science Publications Animal Science 2019 Brevibacterium from Austrian hard cheese harbor a putative histamine catabolism pathway and a plasmid for adaptation to the cheese environment Justin M. Anast Iowa State University, [email protected] Monika Dzieciol University of Veterinary Medicine Vienna Dylan L. Schultz Iowa State University, [email protected] Martin Wagner University of Veterinary Medicine Vienna Evelyne Mann University of Veterinary Medicine Vienna See next page for additional authors Follow this and additional works at: https://lib.dr.iastate.edu/ans_pubs Part of the Agriculture Commons, Animal Sciences Commons, Food Microbiology Commons, and the Genetics and Genomics Commons The complete bibliographic information for this item can be found at https://lib.dr.iastate.edu/ ans_pubs/520. For information on how to cite this item, please visit http://lib.dr.iastate.edu/ howtocite.html. This Article is brought to you for free and open access by the Animal Science at Iowa State University Digital Repository. It has been accepted for inclusion in Animal Science Publications by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Brevibacterium from Austrian hard cheese harbor a putative histamine catabolism pathway and a plasmid for adaptation to the cheese environment Abstract The genus Brevibacterium harbors many members important for cheese ripening. We performed real-time quantitative PCR (qPCR) to determine the abundance of Brevibacterium on rinds of Vorarlberger Bergkäse, an Austrian artisanal washed-rind hard cheese, over 160 days of ripening. Our results show that Brevibacterium are abundant on Vorarlberger Bergkäse rinds throughout the ripening time. -
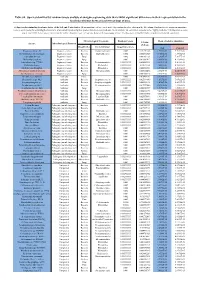
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp. -

Active Microorganisms Thrive Among Extremely Diverse Communities in Cloud Water Pierre Amato, Muriel Joly, Ludovic Besaury, Anne Oudart, Najwa Taïb, Anne I
Active microorganisms thrive among extremely diverse communities in cloud water Pierre Amato, Muriel Joly, Ludovic Besaury, Anne Oudart, Najwa Taïb, Anne I. Moné, Laurent Deguillaume, Anne-Marie Delort, Didier Debroas To cite this version: Pierre Amato, Muriel Joly, Ludovic Besaury, Anne Oudart, Najwa Taïb, et al.. Active microorganisms thrive among extremely diverse communities in cloud water. PLoS ONE, Public Library of Science, 2017, 12 (8), 10.1371/journal.pone.0182869. hal-01598571 HAL Id: hal-01598571 https://hal.archives-ouvertes.fr/hal-01598571 Submitted on 6 Nov 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. RESEARCH ARTICLE Active microorganisms thrive among extremely diverse communities in cloud water Pierre Amato1*, Muriel Joly1, Ludovic Besaury1, Anne Oudart1,2, Najwa Taib2, Anne I. MoneÂ2, Laurent Deguillaume3, Anne-Marie Delort1, Didier Debroas2 1 Universite Clermont Auvergne, CNRS, Institut de Chimie de Clermont-Ferrand, Clermont-Ferrand, France, 2 Universite Clermont Auvergne, CNRS, Laboratoire Microorganismes: GeÂnome et Environnement, Clermont-Ferrand, France, 3 Universite Clermont Auvergne, CNRS, Observatoire de Physique du Globe, Clermont-Ferrand, France a1111111111 a1111111111 * [email protected] a1111111111 a1111111111 a1111111111 Abstract Clouds are key components in Earth's functioning. -

Marine Drugs
Mar. Drugs 2015, 13, 4539-4555; doi:10.3390/md13074539 OPEN ACCESS marine drugs ISSN 1660-3397 www.mdpi.com/journal/marinedrugs Article Structural Investigation of the Oligosaccharide Portion Isolated from the Lipooligosaccharide of the Permafrost Psychrophile Psychrobacter arcticus 273-4 Angela Casillo 1, Ermenegilda Parrilli 1, Sannino Filomena 1,2, Buko Lindner 3, Rosa Lanzetta 1, Michelangelo Parrilli 4, Maria Luisa Tutino 1 and Maria Michela Corsaro 1,* 1 Dipartimento di Scienze Chimiche, Università degli Studi di Napoli Federico II, Complesso Universitario Monte S. Angelo, Via Cintia 4, Napoli 80126, Italy; E-Mails: [email protected] (A.C.); [email protected] (E.P.); [email protected] (S.F.); [email protected] (R.L.); [email protected] (M.L.T.) 2 Institute of Protein Biochemistry, CNR, Via Pietro Castellino 111, Napoli 80131, Italy 3 Division of Bioanalytical Chemistry, Research Center Borstel, Leibniz-Center for Medicine and Biosciences, Parkallee 10, BorstelD-23845, Germany; E-Mail: [email protected] 4 Dipartimento di Biologia, Università degli Studi di Napoli Federico II, Complesso Universitario Monte S. Angelo, Via Cintia 4, Napoli 80126, Italy; E-Mail: [email protected] * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +39-081-674149; Fax: +39-081-674393. Academic Editor: Antonio Trincone Received: 22 June 2015 / Accepted: 14 July 2015 /Published: 22 July 2015 Abstract: Psychrophilic microorganisms have successfully colonized all permanently cold environments from the deep sea to mountain and polar regions. The ability of an organism to survive and grow in cryoenviroments depends on a number of adaptive strategies aimed at maintaining vital cellular functions at subzero temperatures, which include the structural modifications of the membrane. -

Evolutionary Strategies of Highly Functional Catalases for Adaptation to High H2O2 Environments Isao Yumoto, Yoshiko Hanaoka and Isao Hara
Chapter Evolutionary Strategies of Highly Functional Catalases for Adaptation to High H2O2 Environments Isao Yumoto, Yoshiko Hanaoka and Isao Hara Abstract Enzymatic evolutionary strategies for adaptation to a high H2O2 environment have been evaluated using catalases with high catalytic efficiency isolated from two H2O2-tolerant bacteria, Exiguobacterium oxidotolerans and Psychrobacter piscatori. The entrance size of the narrow main channel in catalase has been estimated by deter- mining the formation rate of the intermediate state of peracetic acid (b), which is a larger substrate than H2O2 versus that of catalase activity with H2O2 (a) (calculated as b/a). The ratio of b/a in E. oxidotolerans catalase (EKTA) is much higher than that of P. piscatori catalase (PKTA). To elucidate the structural differences between the catalases, the amino acids present in the main channel have been compared between the two catalases and other catalases in the database. The combination of amino acid residues, which contribute high catalytic efficiency in the narrow main chan- nel of EKTA were different from those in PKTA. In this review, we discuss strategic differences in the elimination of high concentration of H2O2 owing to differences in the phylogenetic positions of catalases. In addition, we describe the relationships between the environmental distributions of genera involved in H2O2-resistant bacte- ria and their catalase functions based on the main channel structure of catalase. Keywords: H2O2-tolerant bacteria, Exiguobacterium, Psychrobacter, Vibrio, catalase, narrow main channel, bottleneck size 1. Introduction Oxygen is important for metabolism, acting as a terminal electron acceptor in aerobic bacteria, and these bacteria produce intracellular reactive oxygen species 2•− (ROS), such as hydrogen peroxide (H2O2), superoxide (O ), and hydroxyl radical • (OH ) as by-products of oxygen metabolism [1–4].